This is the multi-page printable view of this section. Click here to print.
Notes
- 1: Machine Learning
- 1.1: Introduction
- 1.1.1: What is ML ?
- 1.1.2: Types of ML
- 1.2: Supervised Learning
- 1.2.1: Linear Regression
- 1.2.1.1: Meaning of 'Linear'
- 1.2.1.2: Meaning of 'Regression'
- 1.2.1.3: Linear Regression
- 1.2.1.4: Probabilistic Interpretation
- 1.2.1.5: Convex Function
- 1.2.1.6: Gradient Descent
- 1.2.1.7: Polynomial Regression
- 1.2.1.8: Data Splitting
- 1.2.1.9: Cross Validation
- 1.2.1.10: Bias Variance Tradeoff
- 1.2.1.11: Regularization
- 1.2.1.12: Regression Metrics
- 1.2.1.13: Assumptions
- 1.2.2: Logistic Regression
- 1.2.2.1: Binary Classification
- 1.2.2.2: Log Loss
- 1.2.2.3: Regularization
- 1.2.2.4: Log Odds
- 1.2.2.5: Probabilistic Interpretation
- 1.2.3: K Nearest Neighbors
- 1.2.3.1: KNN Introduction
- 1.2.3.2: KNN Optimizations
- 1.2.3.3: Curse Of Dimensionality
- 1.2.3.4: Bias Variance Tradeoff
- 1.2.4: Decision Tree
- 1.2.4.1: Decision Trees Introduction
- 1.2.4.2: Purity Metrics
- 1.2.4.3: Decision Trees For Regression
- 1.2.4.4: Regularization
- 1.2.4.5: Bagging
- 1.2.4.6: Random Forest
- 1.2.4.7: Extra Trees
- 1.2.4.8: Boosting
- 1.2.4.9: AdaBoost
- 1.2.4.10: Gradient Boosting Machine
- 1.2.4.11: GBDT Algorithm
- 1.2.4.12: GBDT Example
- 1.2.4.13: Advanced GBDT Algorithms
- 1.2.4.14: XgBoost
- 1.2.4.15: LightGBM
- 1.2.4.16: CatBoost
- 1.2.5: Support Vector Machine
- 1.2.5.1: SVM Intro
- 1.2.5.2: Hard Margin SVM
- 1.2.5.3: Soft Margin SVM
- 1.2.5.4: Primal Dual Equivalence
- 1.2.5.5: Kernel Trick
- 1.2.5.6: RBF Kernel
- 1.2.5.7: Support Vector Regression
- 1.2.6: Naive Bayes'
- 1.2.6.1: Naive Bayes Intro
- 1.2.6.2: Naive Bayes Issues
- 1.2.6.3: Naive Bayes Example
- 1.3: Unsupervised Learning
- 1.3.1: K Means
- 1.3.1.1: K Means
- 1.3.1.2: Lloyds Algorithm
- 1.3.1.3: K Means++
- 1.3.1.4: K Medoid
- 1.3.1.5: Clustering Quality Metrics
- 1.3.1.6: Silhouette Score
- 1.3.2: Hierarchical Clustering
- 1.3.2.1: Hierarchical Clustering
- 1.3.3: DBSCAN
- 1.3.3.1: DBSCAN
- 1.3.4: Gaussian Mixture Model
- 1.3.4.1: Gaussian Mixture Models
- 1.3.4.2: Latent Variable Model
- 1.3.4.3: Expectation Maximization
- 1.3.5: Anomaly Detection
- 1.3.5.1: Anomaly Detection
- 1.3.5.2: Elliptic Envelope
- 1.3.5.3: One Class SVM
- 1.3.5.4: Local Outlier Factor
- 1.3.5.5: Isolation Forest
- 1.3.5.6: RANSAC
- 1.3.6: Dimensionality Reduction
- 1.4: Feature Engineering
- 1.4.1: Data Pre Processing
- 1.4.2: Categorical Variables
- 1.4.3: Feature Engineering
- 1.4.4: Data Leakage
- 1.4.5: Model Interpretability
- 1.5: ML System
- 1.5.1: Data Distribution Shift
- 1.5.2: Retraining Strategies
- 1.5.3: Deployment Patterns
- 1.6: Interview Questions
- 2: Maths
- 2.1: Probability
- 2.1.1: Introduction to Probability
- 2.1.2: Conditional Probability
- 2.1.3: Independence of Events
- 2.1.4: Cumulative Distribution Function
- 2.1.5: Probability Mass Function
- 2.1.6: Probability Density Function
- 2.1.7: Expectation
- 2.1.8: Moment Generating Function
- 2.1.9: Joint & Marginal
- 2.1.10: Independent & Identically Distributed
- 2.1.11: Convergence
- 2.1.12: Law of Large Numbers
- 2.1.13: Markov's Inequality
- 2.1.14: Cross Entropy & KL Divergence
- 2.1.15: Parametric Model Estimation
- 2.2: Statistics
- 2.2.1: Data Distribution
- 2.2.2: Correlation
- 2.2.3: Central Limit Theorem
- 2.2.4: Confidence Interval
- 2.2.5: Hypothesis Testing
- 2.2.6: T-Test
- 2.2.7: Z-Test
- 2.2.8: Chi-Square Test
- 2.2.9: Performance Metrics
- 2.3: Linear Algebra
- 2.3.1: Vector Fundamentals
- 2.3.2: Matrix Operations
- 2.3.3: Eigen Value Decomposition
- 2.3.4: Principal Component Analysis
- 2.3.5: Singular Value Decomposition
- 2.3.6: Vector & Matrix Calculus
- 2.3.7: Vector Norms
- 2.3.8: Hyperplane
- 2.4: Calculus
- 2.4.1: Calculus Fundamentals
- 2.4.2: Optimization
- 2.4.3: Gradient Descent
- 2.4.4: Newton's Method
1 - Machine Learning
End of Section
1.1 - Introduction
End of Section
1.1.1 - What is ML ?
The machine learning lifecycle ♼ is generally divided into two main stages:
- Training Phase
- Runtime (Inference) Phase
Training Phase:
Where the machine learning model is developed and taught to understand a specific task using a large volume
of historical data.

Runtime (Inference) Phase:
Where the fully trained and deployed model is put to practical use in a real-world 🌎 environment, i.e.,
to make predictions on new, unseen data.

End of Section
1.1.2 - Types of ML
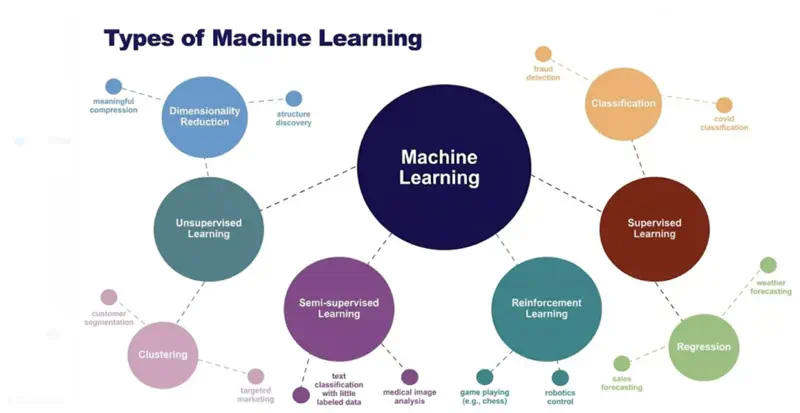
Supervised Learning uses labelled data (input-output pairs) to predict outcomes, such as, spam filters.
- Regression
- Classification
Unsupervised Learning finds hidden patterns in unlabelled data (like customer segmentation).
- Clustering (k-means, hierarchical)
- Dimensionality Reduction and Data Visualization (PCA, t-SNE, UMAP)
Semi-Supervised Learning uses a mix of both, leveraging a small amount of labelled data with a large amount of
unlabelled data to improve accuracy.
- Pseudo-labeling
- Graph-based methods
1. Pseudo-labelling:
- A model is initially trained on the available, limited labelled dataset.
- This trained model is then used to predict labels for the unlabelled data.These predictions are called ‘pseudo-labels’.
- The model is then retrained using both the original labelled data and the newly pseudo-labelled data.
Benefit:
It effectively expands the training data by assigning labels to previously unlabelled examples,
allowing the model to learn from a larger dataset.
2. Graph-based methods:
- Data points (both labelled and unlabelled) are represented as nodes in a graph.
- Edges are established between nodes based on their similarity or proximity in the feature space.The weight of an edge often reflects the degree of similarity.
- The core principle is that similar data points should have similar labels.This assumption is propagated through the graph, effectively ‘spreading’ the labels from the labelled nodes to the unlabelled nodes based on the graph structure.
- Various algorithms, such as label propagation or graph neural networks (GNNs), can be employed to infer the labels of unlabelled nodes.
Benefit:
These methods are particularly useful when the data naturally exhibits a graph-like structure or when local
neighborhood information is crucial for classification.
Agent learns to make optimal decisions by interacting with an environment, receiving rewards (positive feedback)
or penalties (negative feedback) for its actions.
- Mimic human trial-and-error learning to achieve a goal 🎯.
- Agent: The learning entity that makes decisions and takes actions within the environment.
- Environment: The external system with which the agent interacts.It defines the rules, states, and the consequences of the agent’s actions.
- State: A specific configuration or situation of the environment at a given point in time.
- Action: A move or decision made by the agent in a particular state.
- Reward: A numerical signal received by the agent from the environment, indicating the desirability of an action taken
in a specific state.
Positive rewards encourage certain behaviors, while negative rewards (penalties) discourage them. - Policy: The strategy or mapping that defines which action the agent should take in each state to maximize long-term rewards 💰.
- Exploration: The agent tries out new actions to discover their effects and potentially find better strategies.
- Exploitation: The agent utilizes its learned knowledge to choose actions that have yielded high rewards in the past.
Note: The agent continuously balances exploration and exploitation to refine its policy and achieve the optimal behavior.
Large Language Models (LLMs) are deep learning models that often employ unsupervised learning techniques during their pre-training phase.
LLMs are trained on massive amounts of raw, unlabelled text data (e.g., books, articles, web pages) to predict the next
word in a sequence or fill in masked words.
This process, often called self-supervised learning, allows the model to learn grammar, syntax, semantics,
and general world knowledge by identifying statistical relationships within the text.
LLMs generally also undergo supervised fine-tuning (SFT) for specific tasks, where they are trained on labeled datasets to improve performance on those tasks.
Reinforcement Learning from Human Feedback (RLHF) allows LLMs to learn from human judgment, enabling them to generate more nuanced, context-aware, and ethically aligned outputs that better meet human expectations.
End of Section
1.2 - Supervised Learning
End of Section
1.2.1.1 - Meaning of 'Linear'
Equation of a line is of the form \(y = mx + c\).
To represent a line in 2D space, we need 2 things:
- m = slope or direction of the line
- c = y-intercept or distance from the origin
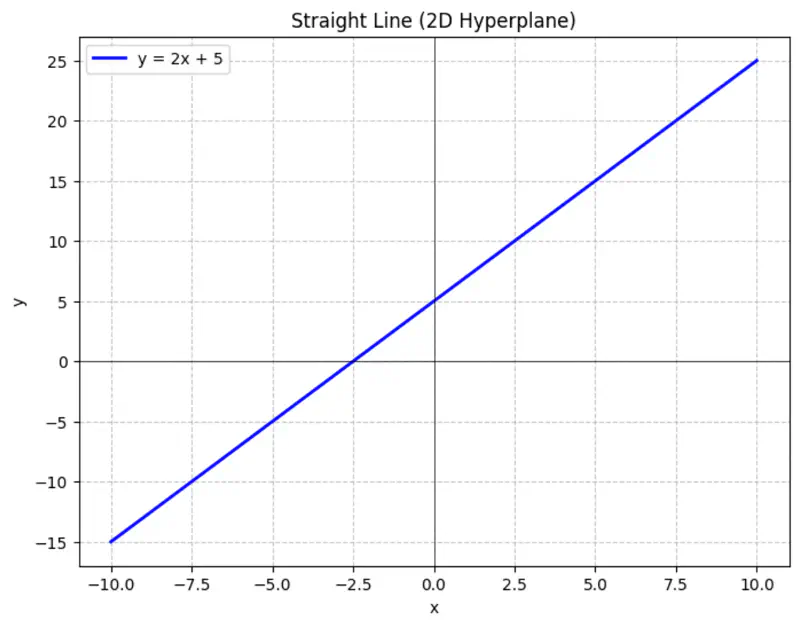
Similarly,
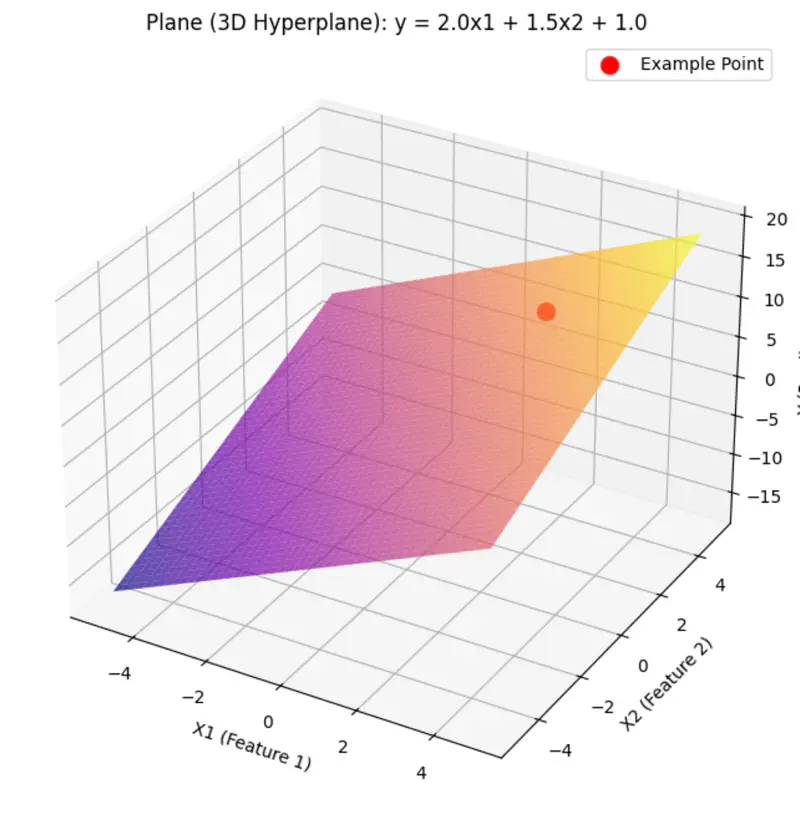
A hyperplane is a lower (d-1) dimensional sub-space that divides a d-dimensional space into 2 distinct parts.
Equation of a hyperplane:
Here, ‘y’ is expressed as a linear combination of parameters - \( w_0, w_1, w_2, \dots, w_n \)
Hence - Linear means the model is ‘linear’ with respect to its parameters NOT the variables.
Read more about Hyperplane
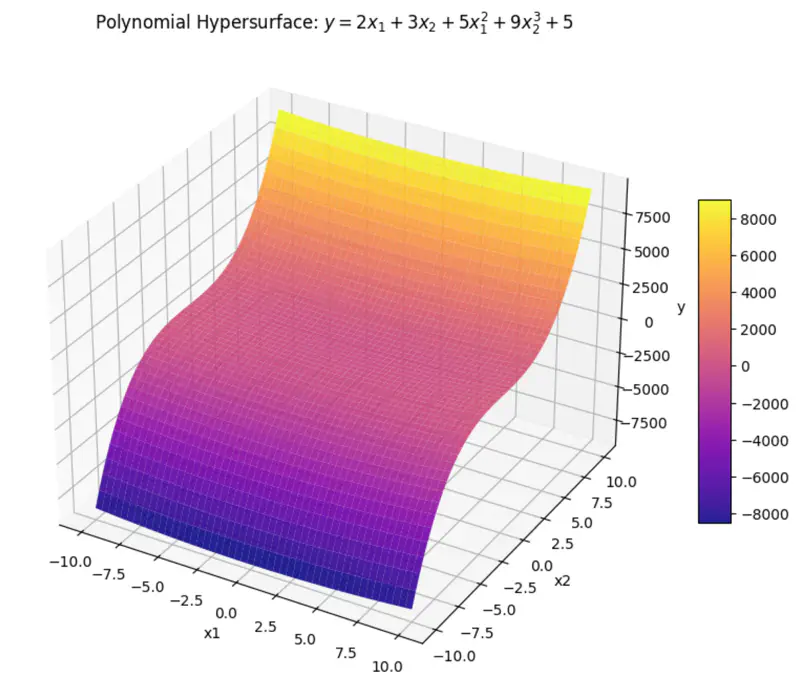
can be rewritten as:
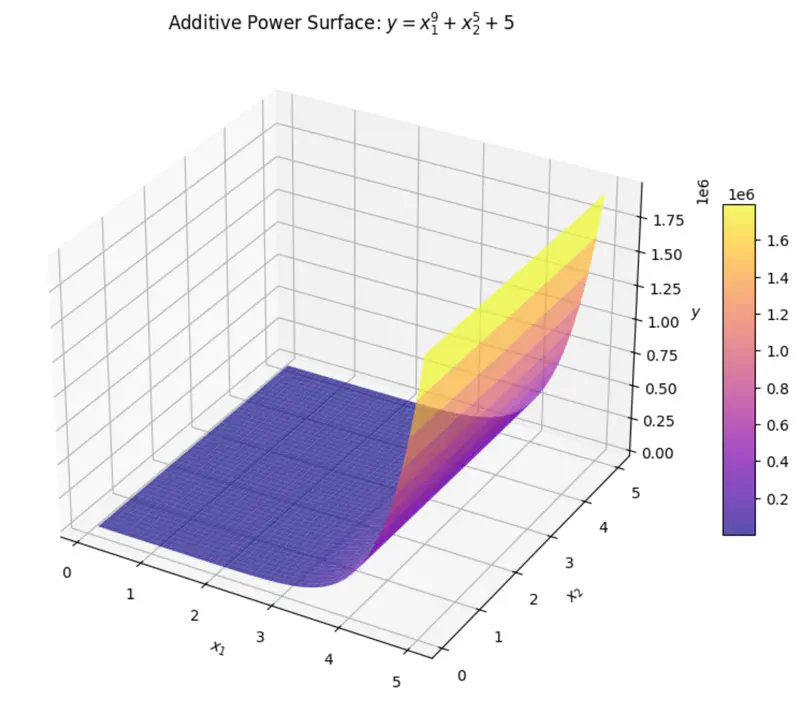
\[y = w_1x_1 + w_2x_2 + w_3x_3 + w_4x_4 + w_0\]where, \(x_3 = x_1^2 ~ and ~ x_4 = x_2^3 \)
\(x_3 ~ and ~ x_4 \) are just 2 new (polynomial) variables.
And, ‘y’ is still a linear combination of parameters: \(w_0, w_1, \dots w_4\)
can be rewritten as:
\[y = w_1x_1 + w_2x_2 + w_0\]where, \(x_1 = log(x) ~ and ~ x_2 = \sqrt{x} \)
\(x_1 ~ and ~ x_2 \) are are transformations of variable \(x\).
And, ‘y’ is still a linear combination of parameters: \(w_0, w_1, ~and~ w_2\)
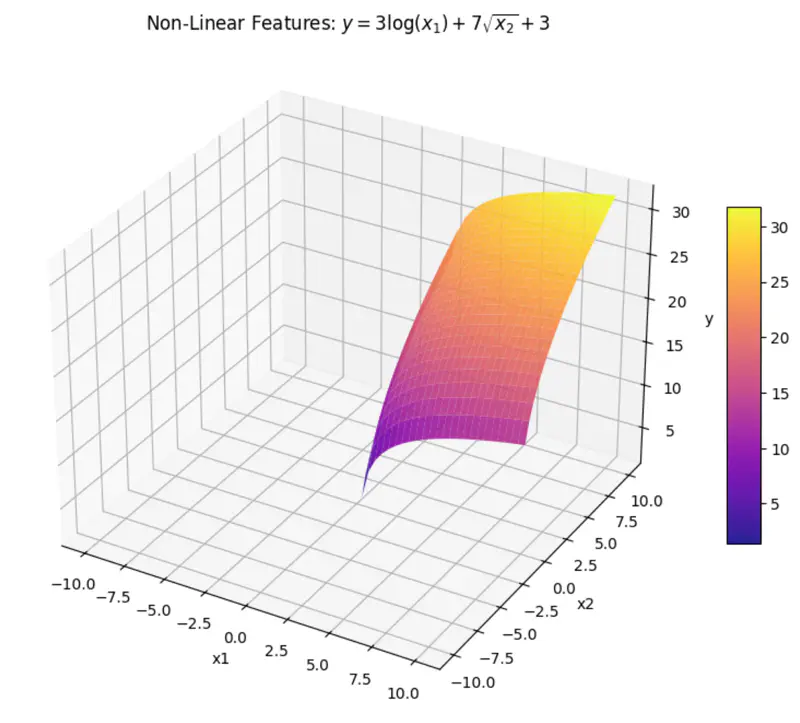
Even if we take log, we get:
\[log(y) = w_1log(x_1) + w_2log(x_2) + log(w_0)\]here, \(log(w_0) \) parameter is NOT linear.
End of Section
1.2.1.2 - Meaning of 'Regression'
Regression = Going Back
Regression has a very specific historical origin that is different from its current statistical meaning.
Sir Francis Galton (19th century), cousin of Charles Darwin, coined 🪙 this term.
Observation:
Galton observed that -
the children 👶 of unusually tall ⬆️ parents 🧑🧑🧒🧒,
tended to be shorter ⬇️ than their parents 🧑🧑🧒🧒,
and children 👶 of unusually short ⬇️ parents 🧑🧑🧒🧒,
tended to be taller ⬆️ than their parents 🧑🧑🧒🧒.
Galton named this biological tendency - ‘regression towards mediocrity/mean’.
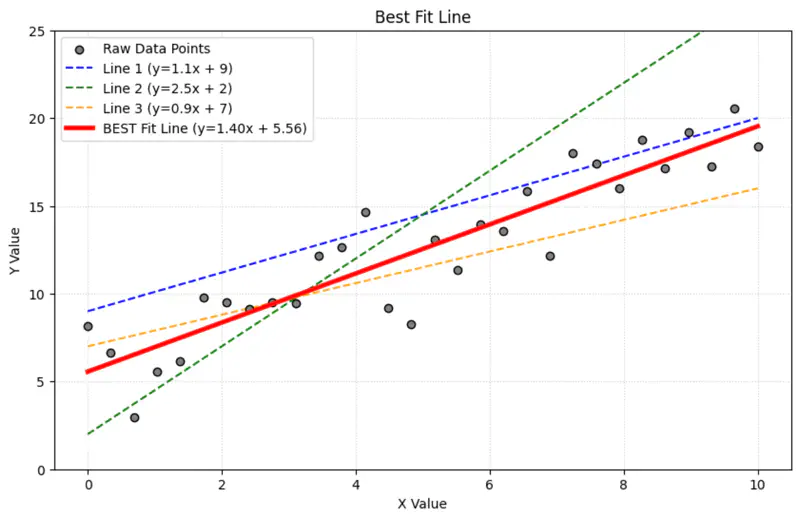
Galton used method of least squares to model this relationship, by fitting a line to the data 📊.
Regression = Fitting a Line
Over time ⏳, the name ‘regression’ got permanently attached to the method of fitting line to the data 📊.
Today in statistics and machine learning, ‘regression’ universally refers to the method of finding the
‘line of best fit’ for a set of data points, NOT the concept of ‘regressing towards the mean’.
End of Section
1.2.1.3 - Linear Regression
Let’s understand linear regression using an example to predict salary.
Predict the salary 💰 of an IT employee, based on various factors, such as, years of experience, domain, role, etc.
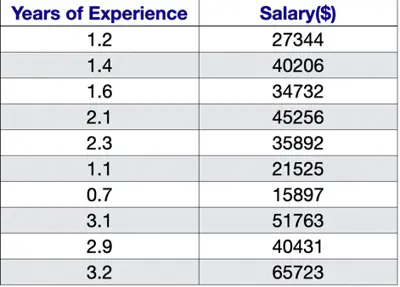
Let’s start with a simple problem and predict the salary using only one input feature.
Goal 🎯 : Find the line of best fit.
Plot: Salary vs Years of Experience
\[y = mx + c = w_1x + w_0\]Slope = \(m = w_1 \)
Intercept = \(c = w_0\)
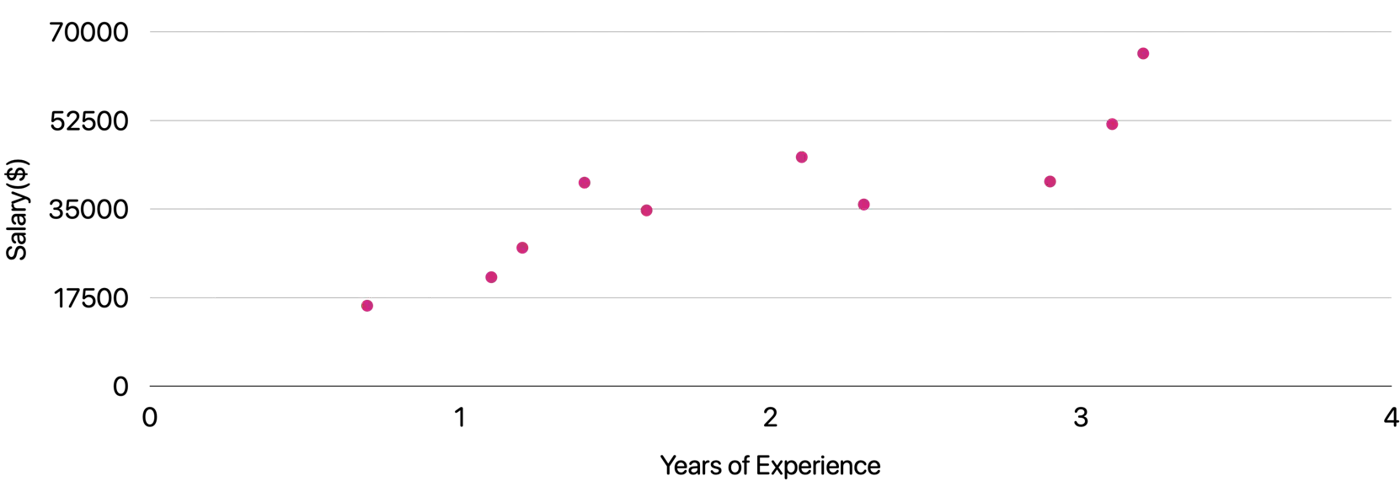
Similarly, if we include other factors/features impacting the salary 💰, such as, domain, role, etc, we get an equation of a fitting hyperplane:
\[y = w_1x_1 + w_2x_2 + \dots + w_dx_d + w_0\]where,
\[ \begin{align*} x_1 &= \text{Years of Experience} \\ x_2 &= \text{Domain (Tech, BFSI, Telecom, etc.)} \\ x_3 &= \text{Role (Dev, Tester, DevOps, ML, etc.)} \\ x_d &= d^{th} ~ feature \\ w_0 &= \text{Salary of 0 years experience} \\ \end{align*} \]Space = ’d’ features + 1 target variable = ’d+1’ dimensions
In a ’d+1’ dimensional space, we try to fit a ’d’ dimensional hyperplane.
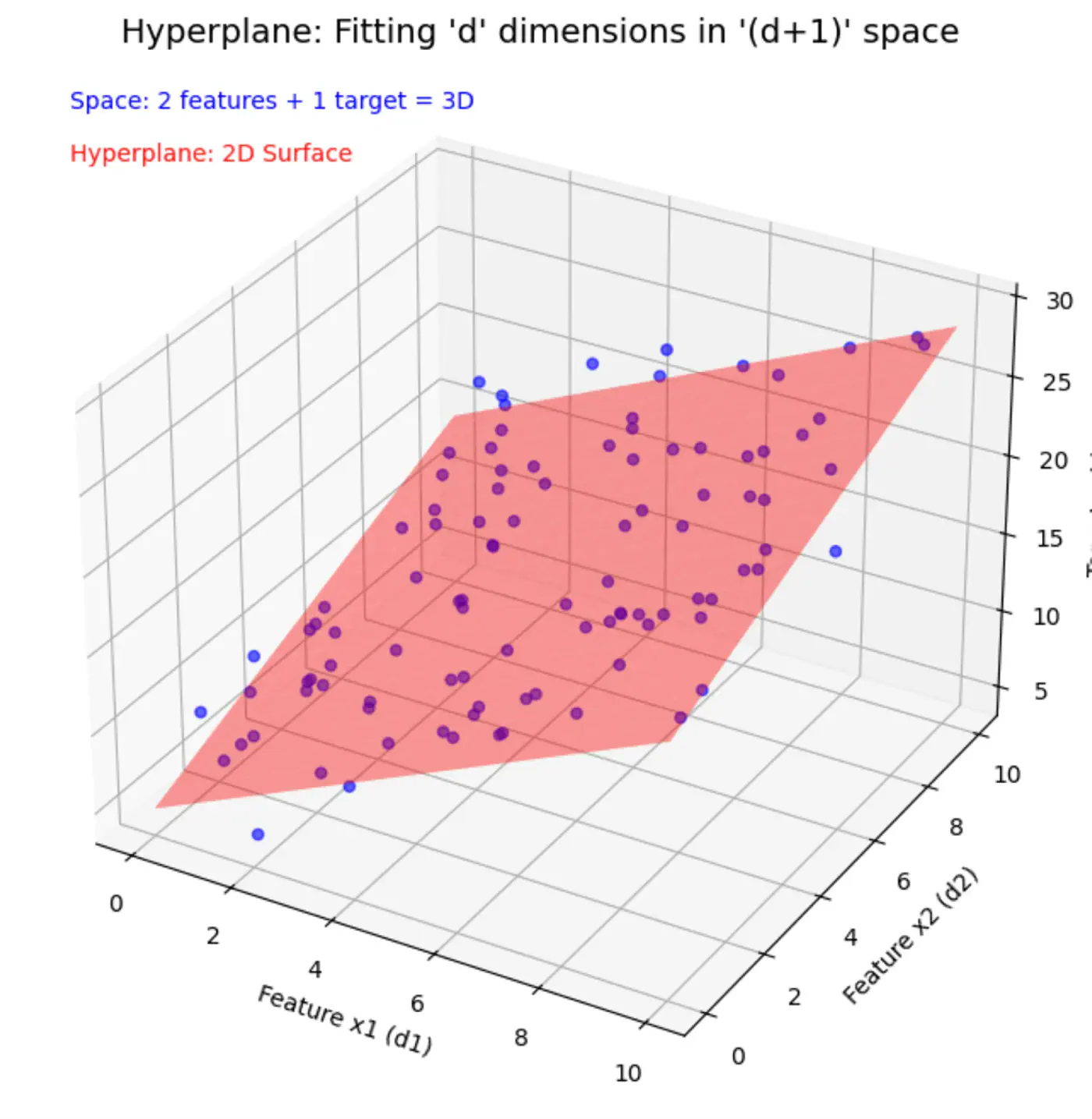
Let, data = \( {(x_i, y_i)}_{i=1}^N ; ~ x_i \in R^d , y_i \in R^d\)
where, N = number of training samples.
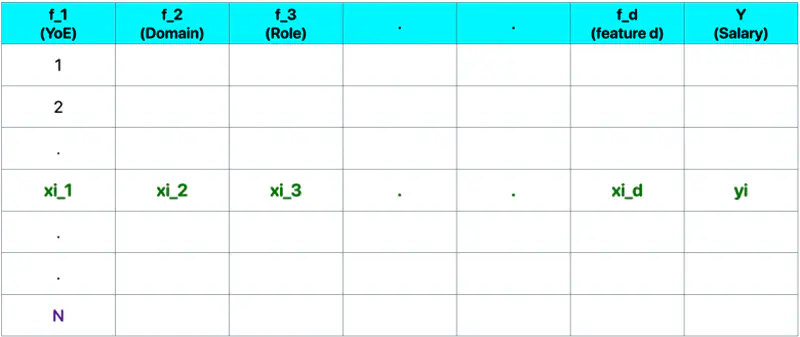
Note: Fitting hyperplane (\(y = w_1x_1 + w_2x_2 + \dots + w_dx_d + w_0\)) is the model.
Objective 🎯: find the parameters/weights (\(w_0, w_1, w_2, \dots w_d \)) of the model.
\(\mathbf{w} = \begin{bmatrix} w_1 \\ w_2 \\ \vdots \\ w_d \end{bmatrix}_{\text{d x 1}}\)
\( \mathbf{x_i} = \begin{bmatrix} x_{i_1} \\ x_{i_2} \\ \vdots \\ x_{i_d} \end{bmatrix}_{\text{d x 1}} \)
\( \mathbf{y} = \begin{bmatrix} y_1 \\ y_2 \\ \vdots \\ y_i \\ \vdots \\ y_n \end{bmatrix}_{\text{n x 1}} \)
\( X =
\begin{bmatrix}
x_{11} & x_{12} & \cdots & x_{1d} \\
x_{21} & x_{22} & \cdots & x_{2d} \\
\vdots & \vdots & \ddots & \vdots \\
x_{i1} & x_{i2} & \cdots & x_{id} \\
\vdots & \vdots & \ddots & \vdots \\
x_{n1} & x_{n2} & \cdots & x_{nd} \\
\end{bmatrix}
_{\text{n x d}}
\)
Prediction:
Error = Actual - Predicted
\[ \epsilon_i = y_i - \hat{y_i}\]Goal 🎯: Minimize error between actual and predicted.
We can quantify the error for a single data point in following ways:
- Absolute error = \(|y_i - \hat{y_i}|\)
- Squared error = \((y_i - \hat{y_i})^2\)
Not differentiable at x=0, required for gradient descent.
Constant gradient, i.e \(\pm 1\), model learns at same rate, whether the error is large or small.
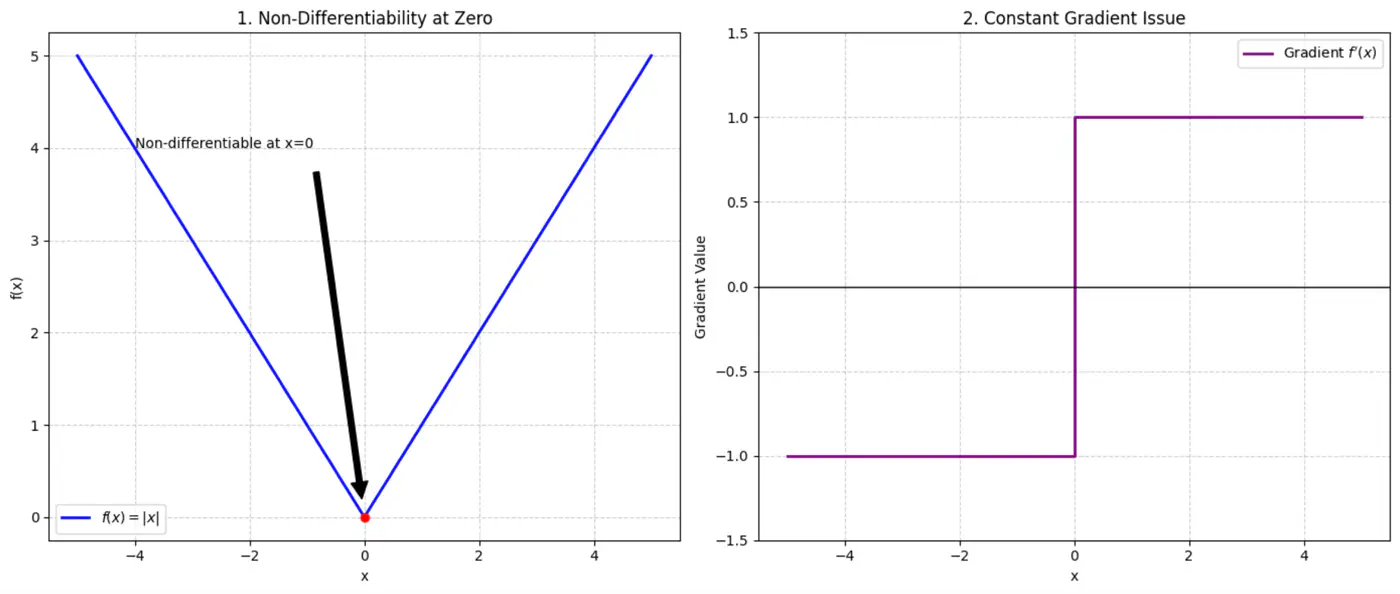
Average loss across all data points.
Mean Squared Error (MSE) =
\[ J(w) = \frac{1}{n} \sum_{i=1}^N (y_i - \hat{y_i})^2 \]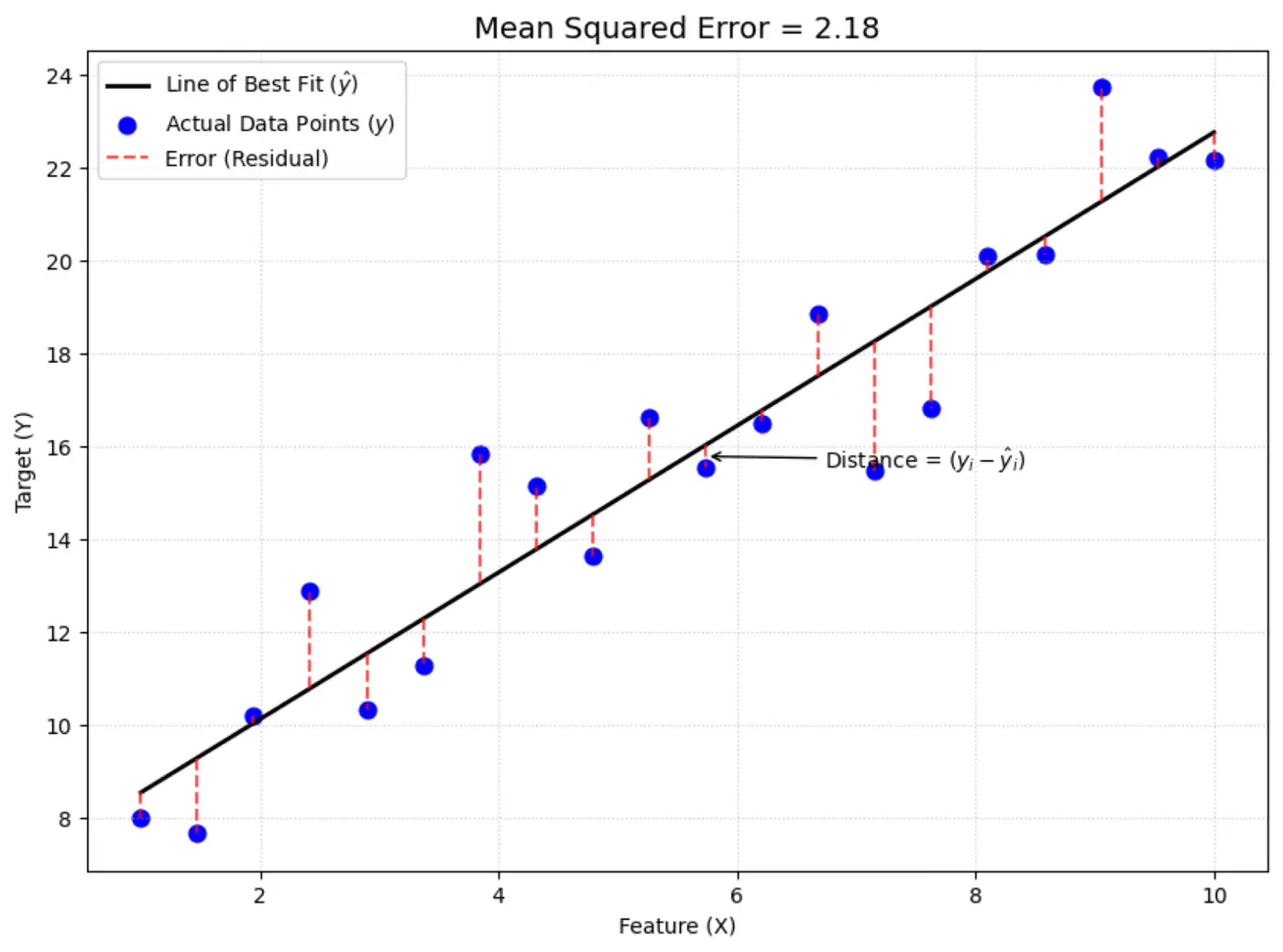
The above equation is quadratic in \(w_0, w_1, w_2, \dots w_d \).
Below is an image of a Paraboloid in 3D, similarly we will have a Paraboloid in ’d’ dimensions.
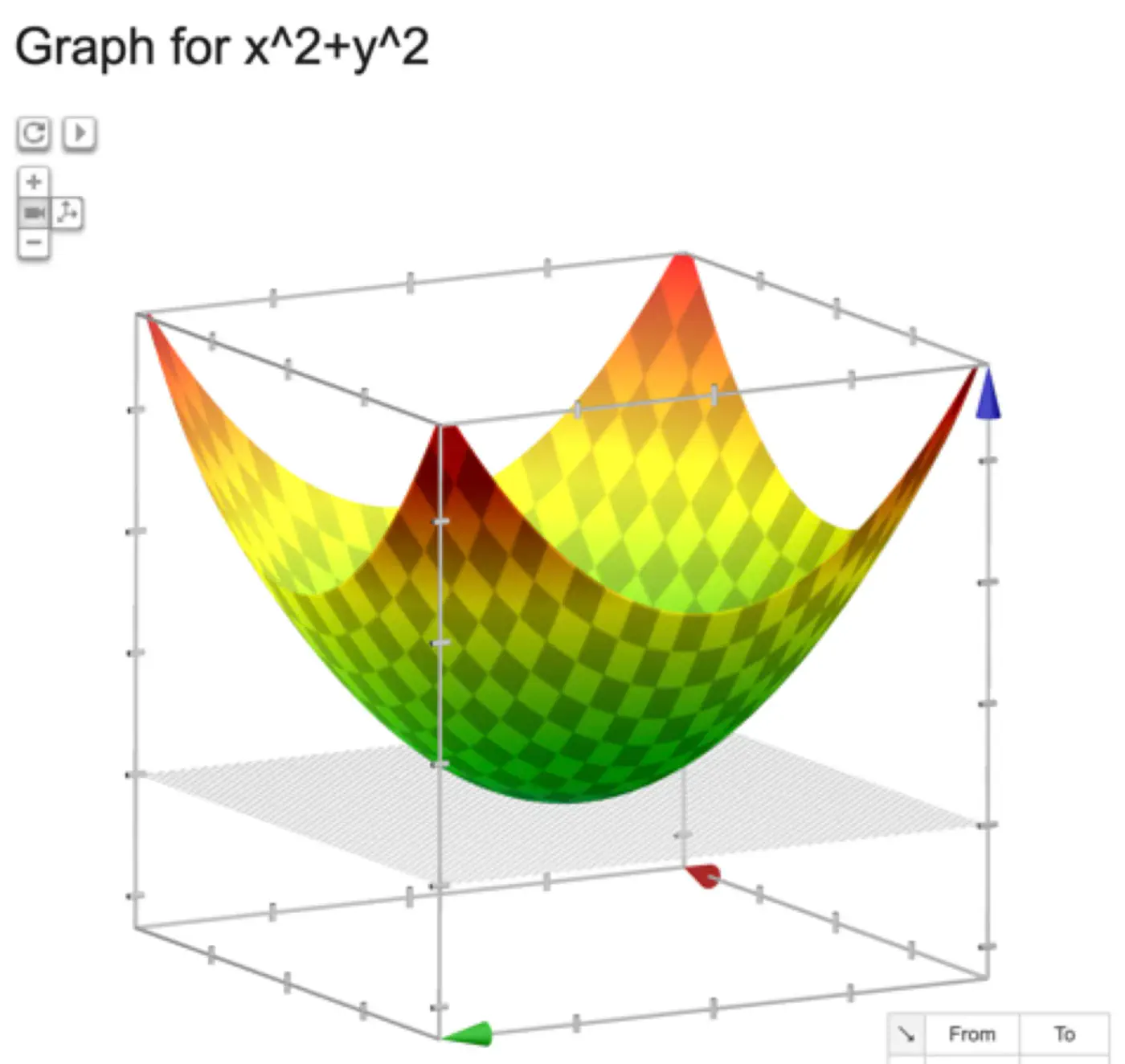
In order to find the minima of the cost function we need to take its derivative w.r.t weights and equate to 0.
\[ \begin{align*} \frac{\partial{J(w)}}{\partial{w_0}} = 0 \\ \frac{\partial{J(w)}}{\partial{w_1}} = 0 \\ \frac{\partial{J(w)}}{\partial{w_2}} = 0 \\ \vdots \\ \frac{\partial{J(w)}}{\partial{w_d}} = 0 \\ \end{align*} \]We have ‘d+1’ linear equations to solve for ‘d+1’ weights \(w_0, w_1, w_2, \dots , w_d\).
But solving ‘d+1’ system of linear equations (called the ’normal equations’) is tedious and NOT used for practical purposes.
where,
\(\mathbf{w} = \begin{bmatrix} w_1 \\ w_2 \\ \vdots \\ w_d \end{bmatrix}_{\text{d x 1}}\)
\( \mathbf{x_i} = \begin{bmatrix} x_{i_1} \\ x_{i_2} \\ \vdots \\ x_{i_d} \end{bmatrix}_{\text{d x 1}} \)
\( \mathbf{y} = \begin{bmatrix} y_1 \\ y_2 \\ \vdots \\ y_i \\ \vdots \\ y_n \end{bmatrix}_{\text{n x 1}} \)
\( X =
\begin{bmatrix}
x_{11} & x_{12} & \cdots & x_{1d} \\
x_{21} & x_{22} & \cdots & x_{2d} \\
\vdots & \vdots & \ddots & \vdots \\
x_{i1} & x_{i2} & \cdots & x_{id} \\
\vdots & \vdots & \ddots & \vdots \\
x_{n1} & x_{n2} & \cdots & x_{nd} \\
\end{bmatrix}
_{\text{n x d}}
\)
Prediction:
Let’s expand the cost function J(w):
\[ \begin{align*} J(w) &= \frac{1}{n} (y - Xw)^2 \\ &= \frac{1}{n} (y - Xw)^T(y - Xw) \\ &= \frac{1}{n} (y^T - w^TX^T)(y - Xw) \\ J(w) &= \frac{1}{n} (y^Ty - w^TX^Ty - y^TXw + w^TX^TXw) \end{align*} \]Since,\(w^TX^Ty\), is a scalar, so it is equal to its transpose.
\[ w^TX^Ty = (w^TX^Ty)^T = y^TXw\]\[ J(w) = \frac{1}{n} (y^Ty - y^TXw - y^TXw + w^TX^TXw)\]\[ J(w) = \frac{1}{n} (y^Ty - 2y^TXw + w^TX^TXw) \]Note: \(X^2 = X^TX\) and \((AB)^T = B^TA^T\)
To find the minimum, take the derivative of cost function J(w) w.r.t ‘w’, and equate to 0 vector.
\[\frac{\partial{J(w)}}{\partial{w}} = \vec{0}\]\[ \begin{align*} &\frac{\partial{[\frac{1}{n} (y^Ty - 2y^TXw + w^TX^TXw)]}}{\partial{w}} = 0\\ & \implies 0 - 2X^Ty + (X^TX + X^TX)w = 0 \\ & \implies \cancel{2}X^TXw = \cancel{2} X^Ty \\ & \therefore \mathbf{w} = (X^TX)^{-1}X^T\mathbf{y} \end{align*} \]Note: \(\frac{\partial{(a^T\mathbf{x})}}{\partial{\mathbf{x}}} = a\) and \(\frac{\partial{(\mathbf{x}^TA\mathbf{x})}}{\partial{\mathbf{x}}} = (A + A^T)\mathbf{x}\)
This is the closed-form solution of normal equations.
- Inverse may NOT exist (non-invertible).
- Time complexity of calculating the inverse is O(n^3).
If the inverse does NOT exist then we can use the approximation of the inverse, also called Pseudo Inverse or Moore Penrose Inverse (\(A^+\)).
Moore Penrose Inverse ( \(A^+\)) is calculated using Singular Value Decomposition (SVD).
SVD of \(A = U \Sigma V^T\)
Pseudo Inverse \(A^+ = V \Sigma^+ U^T\)
Where, \(\Sigma^+\) is a transpose of \(\Sigma\) with reciprocals of non-zero singular values on its diagonals.
e.g:
Note: Time Complexity = O(mn^2)
End of Section
1.2.1.4 - Probabilistic Interpretation
Error = Random Noise, Un-modeled effects
\[ \begin{align*} \epsilon_i = y_i - \hat{y_i} \\ \implies y_i = \hat{y_i} + \epsilon_i \\ \because \hat{y_i} = x_i^Tw \\ \therefore y_i = x_i^Tw + \epsilon_i \\ \end{align*} \]Actual value(\(y_i\)) = Deterministic linear predictor(\(x_i^Tw\)) + Error term(\(\epsilon_i\))
- Independent and Identically Distributed (I.I.D):
Each error term is independent of others. - Normal (Gaussian) Distributed:
Error follows a normal distribution with mean = 0 and a constant variance, .
This implies that the target variable itself is a random variable, normally distributed around the linear relationship.
\[(y_{i}|x_{i};w)∼N(x_{i}^{T}w,\sigma^{2})\]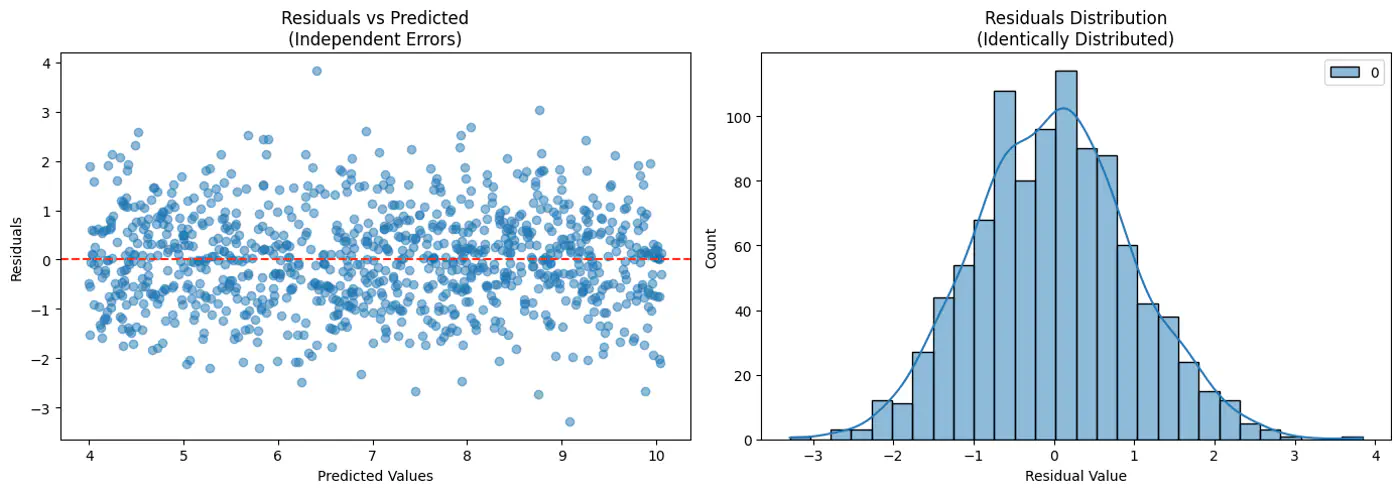
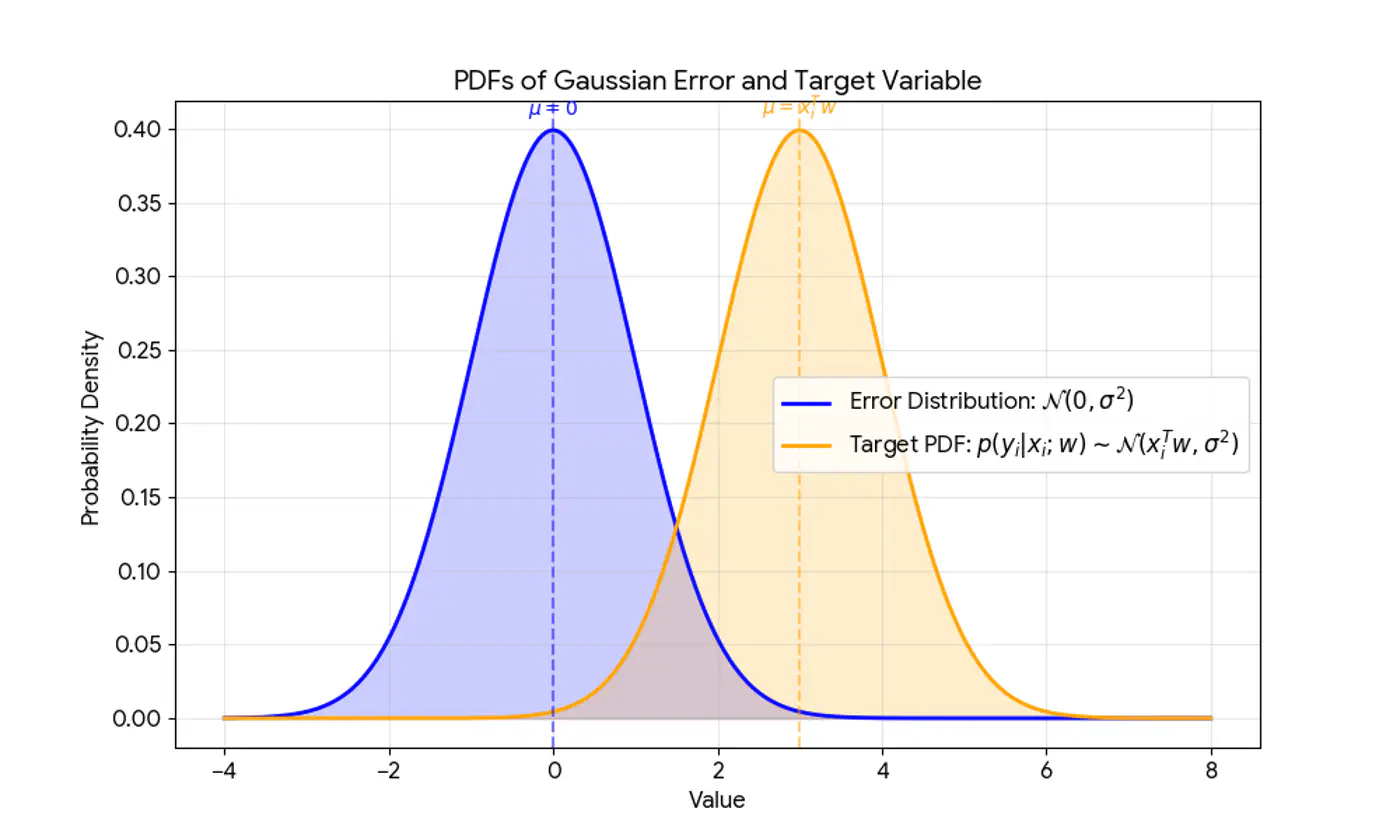
Central Limit Theorem (CLT) states that for a sequence of I.I.D random variables, the distribution of the sample mean(sum) approximates to a normal distribution, regardless of the original population distribution.
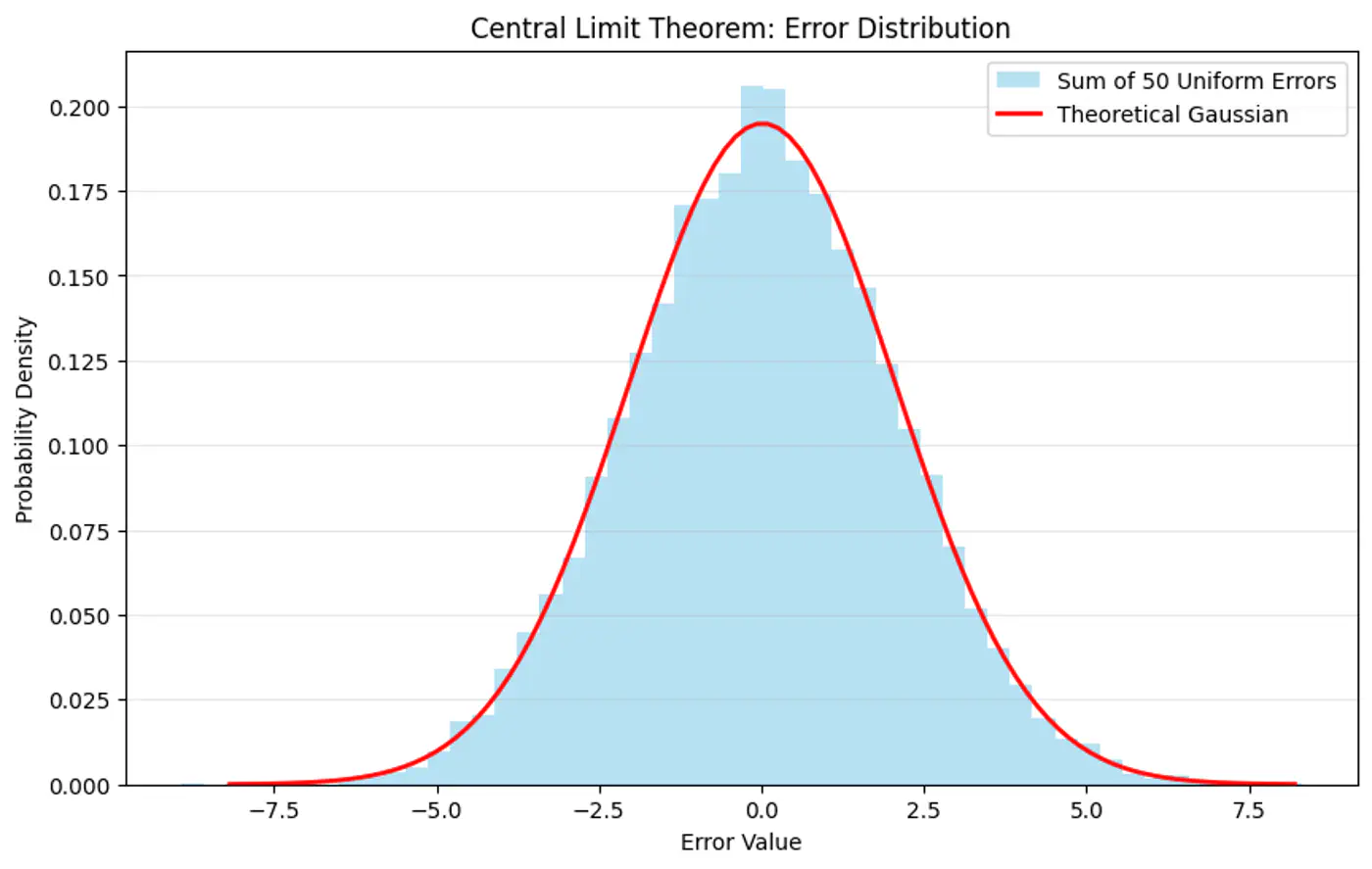
- Probability (Forward View):
Quantifies the chance of observing a specific outcome given a known, fixed model. - Likelihood (Backward/Inverse View):
Inverse concept used for inference (working backward from results to causes). It is a function of the parameters and measures how ‘likely’ a specific set of parameters makes the observed data appear.
‘Find the most plausible explanation for what I see.'
The goal of the probabilistic interpretation is to find the parameters ‘w’ that maximize the probability (likelihood) of observing the given dataset.
Assumption: Training data is I.I.D.
\[ \begin{align*} Likelihood &= \mathcal{L}(w) \\ \mathcal{L}(w) &= p(y|x;w) \\ &= \prod_{i=1}^N p(y_i| x_i; w) \\ &= \prod_{i=1}^N \frac{1}{\sigma\sqrt{2\pi}}e^{-\frac{(y_i-x_i^Tw)^2}{2\sigma^2}} \end{align*} \]Maximizing the likelihood function is mathematically complex due to the product term and the exponential function.
A common simplification is to maximize the log-likelihood function instead, which converts the product into a sum.
Note: Log is a strictly monotonically increasing function.
Note: The first term is constant w.r.t ‘w'.
So, we need to find parameters ‘w’ that maximize the log likelihood.
\[ \begin{align*} log \mathcal{L}(w) & \propto -\frac{1}{2\sigma^2} \sum_{i=1}^N (y_i-x_i^Tw)^2 \\ & \because \frac{1}{2\sigma^2} \text{ is constant} \\ log \mathcal{L}(w) & \propto -\sum_{i=1}^N (y_i-x_i^Tw)^2 \\ \end{align*} \]Maximizing the log-likelihood is equivalent to minimizing the sum of squared errors, which is the exact objective of the ordinary least squares (OLS) method.
\[ \underset{w}{\mathrm{min}}\ J(w) = \underset{w}{\mathrm{min}}\ \sum_{i=1}^N (y_i - x_i^Tw)^2 \]End of Section
1.2.1.5 - Convex Function
Refers to a property of a function where a line segment connecting any two points on its graph lies above or on the graph itself.
A convex function is curved upwards.
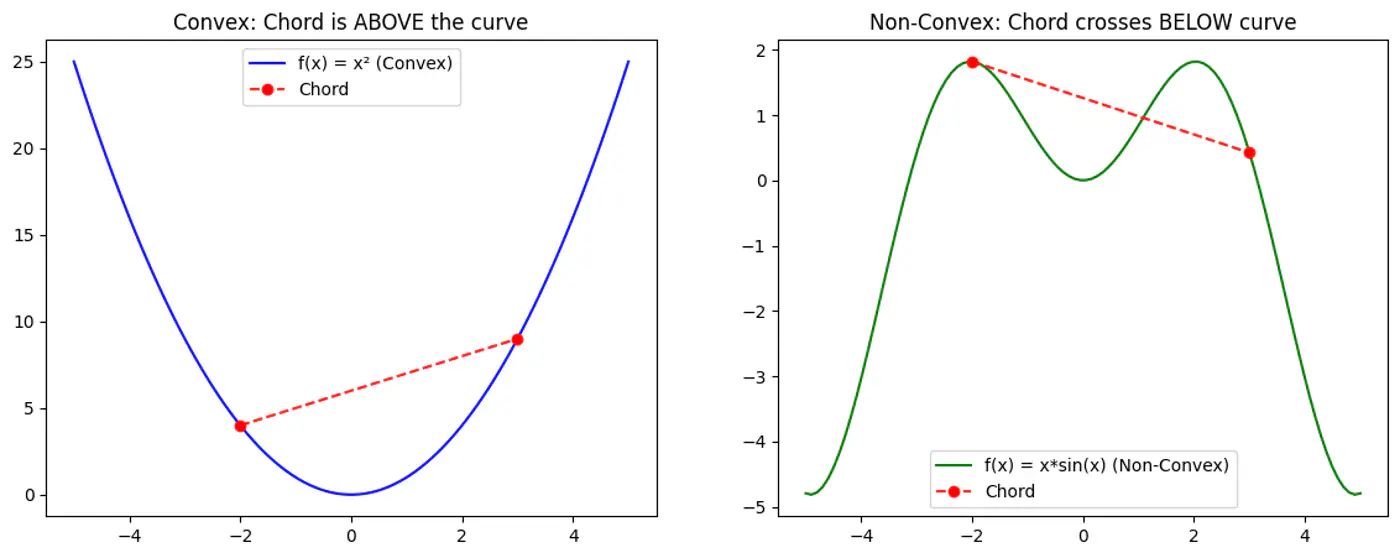
MSE cost function J(w) is convex because its Hessian (H) is always positive semi definite.
\[\nabla J(\mathbf{w})=\frac{1}{n}\mathbf{X}^{T}(\mathbf{Xw}-\mathbf{y})\]\[\mathbf{H}=\frac{\partial (\nabla J(\mathbf{w}))}{\partial \mathbf{w}^{T}}=\frac{1}{n}\mathbf{X}^{T}\mathbf{X}\]\[\therefore \mathbf{H} = \nabla^2J(w) = \frac{1}{n} \mathbf{X}^{T}\mathbf{X}\]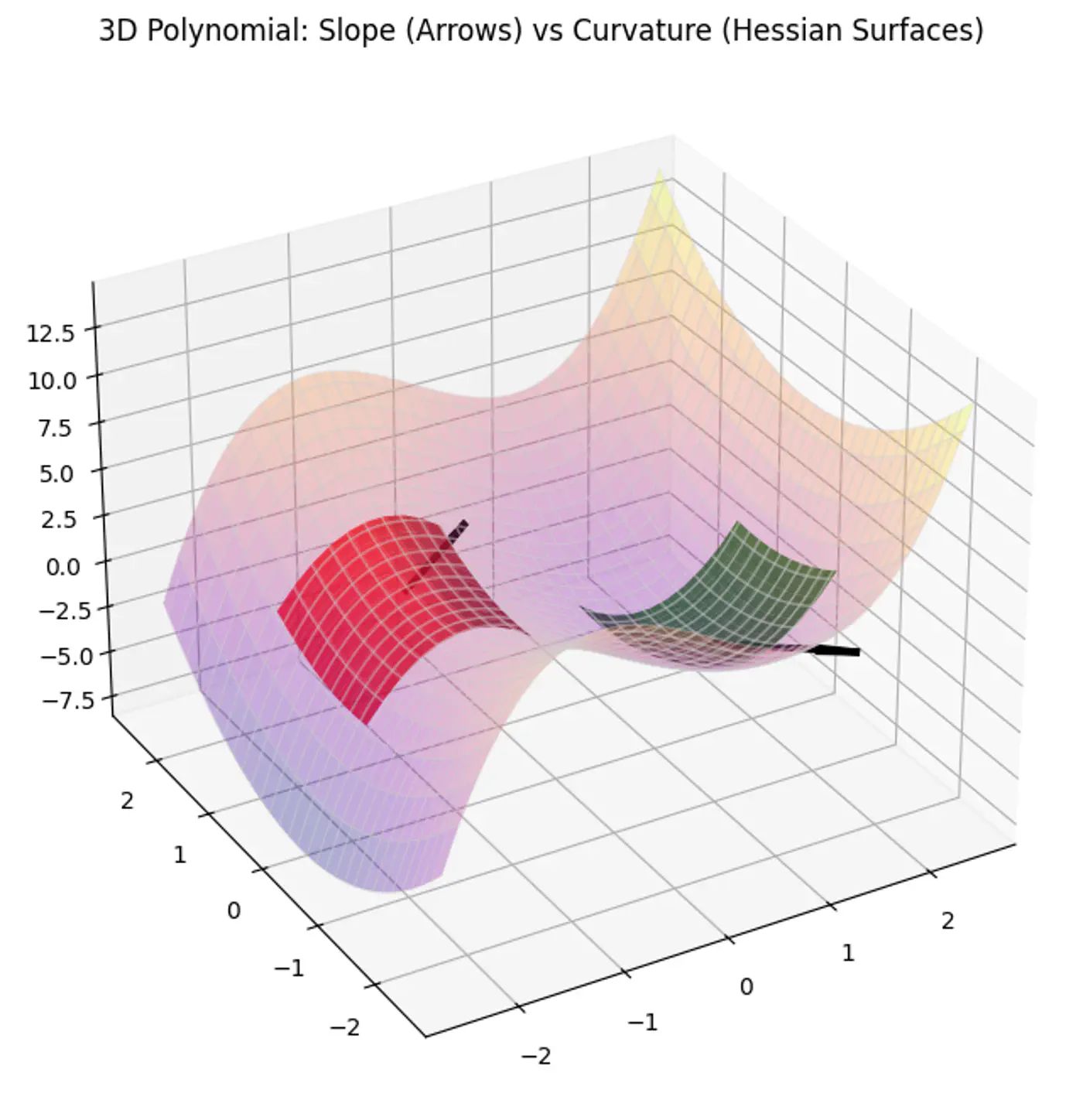
A matrix H is positive semi-definite if and only if for any non-zero vector ‘z’, the quadratic form \(z^THz \ge 0\).
For the Hessian of J(w),
\[ z^THz = z^T(\frac{1}{n}X^TX)z = \frac{1}{n}(Xz)^T(Xz) \]\((Xz)^T(Xz) = \lVert Xz \rVert^2\) = squared L2 norm (magnitude) of the vector
Note: The squared norm of any real-valued vector is always \(\ge 0\).
End of Section
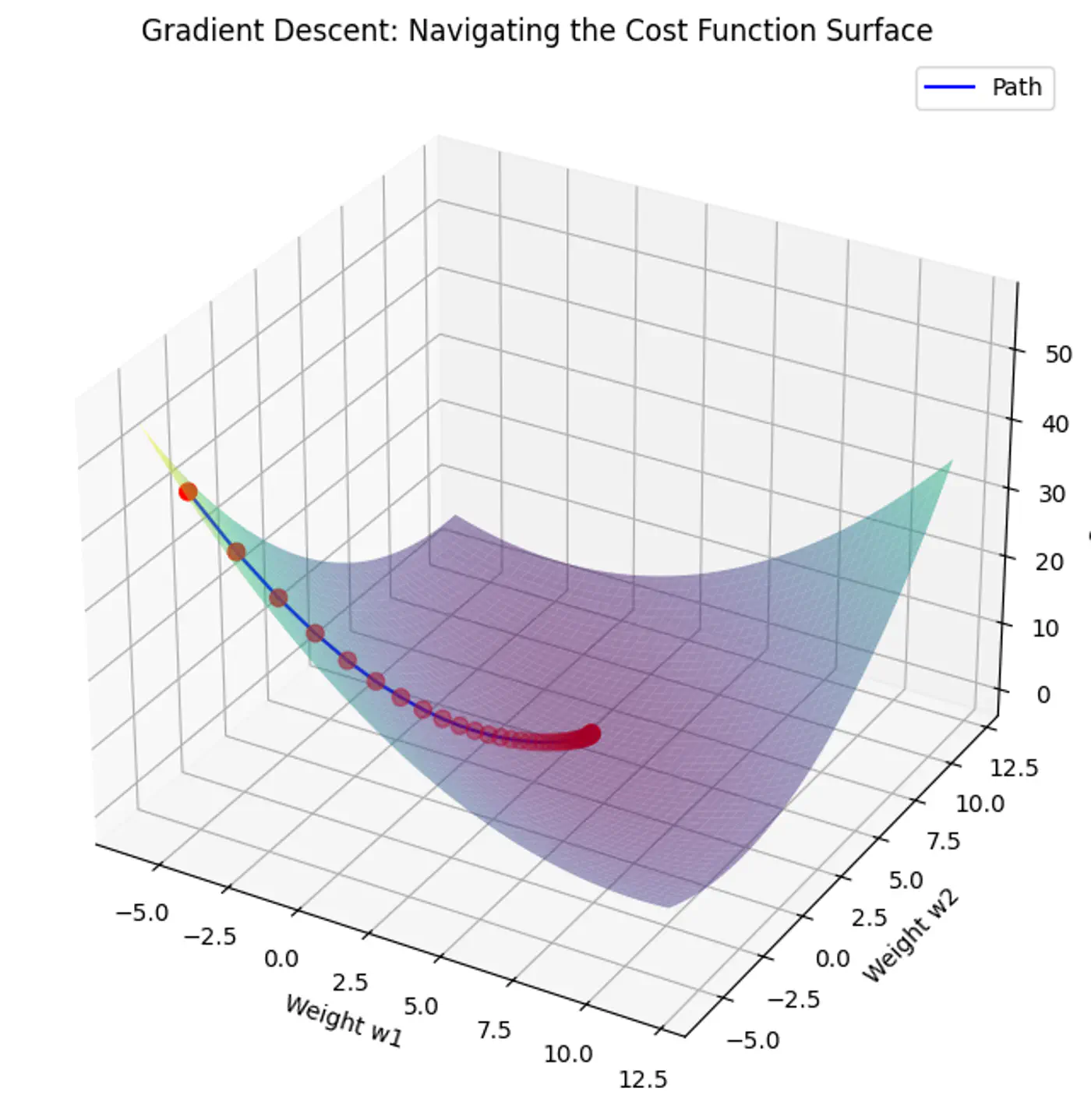
1.2.1.6 - Gradient Descent
Minimize the cost 💰function.
\[J(w)=\frac{1}{2n}(y-Xw)^{2}\]Note: The 1/2 term is included simply to make the derivative cleaner (it cancels out the 2 from the square).
Normal Equation (Closed-form solution) jumps straight to the optimal point in one step.
\[w=(X^{T}X)^{-1}X^{T}y\]But it is not always feasible and computationally expensive 💰(due to inverse calculation 🧮)
An iterative optimization algorithm slowly nudges parameters ‘w’ towards a value that minimize the cost💰 function.
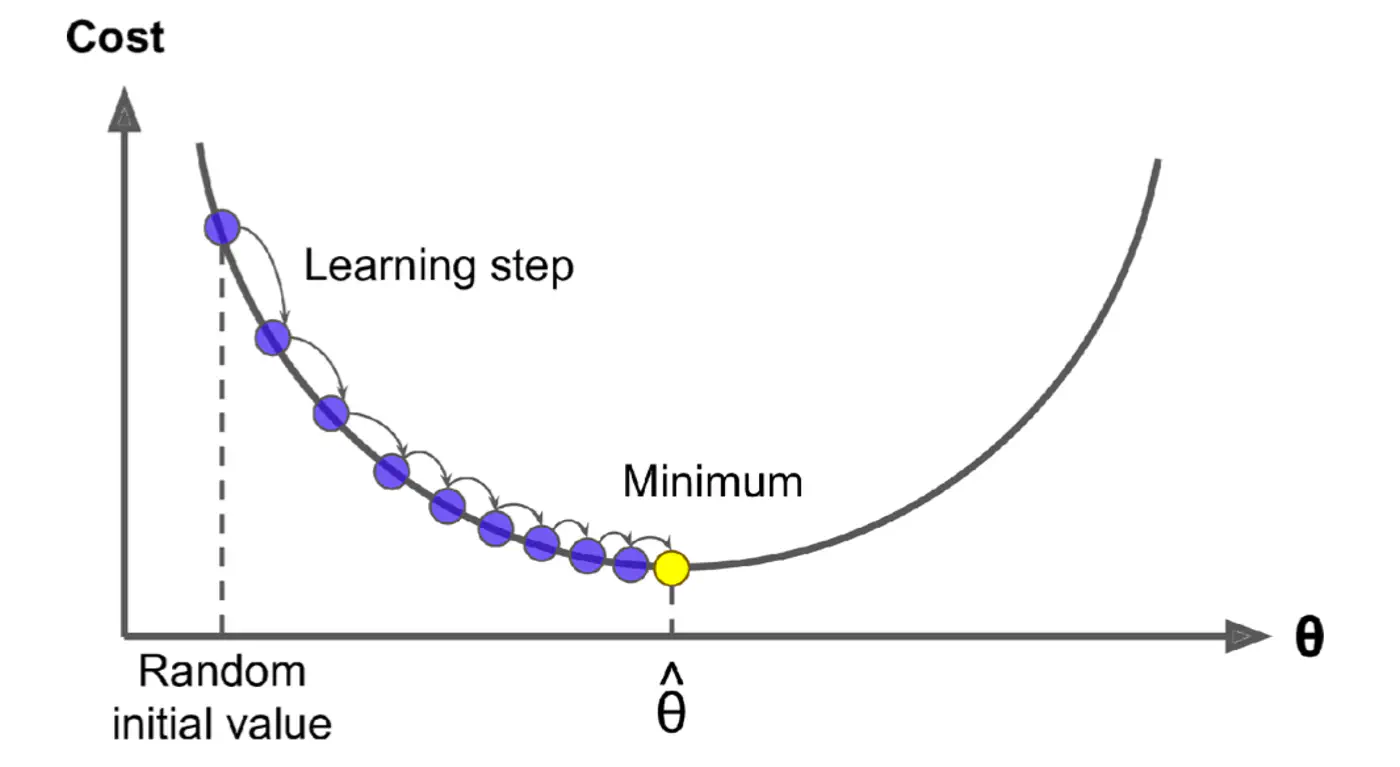
- Initialize the weights/parameters with random values.
- Calculate the gradient of the cost function at current parameter values.
- Update the parameters using the gradient. \[ w_{new} = w_{old} - \eta \frac{\partial{J(w)}}{\partial{w_{old}}} \] \( \eta \) = learning rate or step size to take for each parameter update.
- Repeat 🔁 steps 2 and 3 iteratively until convergence (to minima).
Applying chain rule:
\[ \begin{align*} &\text{Let } u = (y - Xw) \\ &\text{Derivative of } u^2 \text{ w.r.t 'w' }= 2u.\frac{\partial{u}}{\partial{w}} \\ \frac{\partial{(\frac{1}{2n} (y - Xw)^2)}}{\partial{w}} &= \frac{1}{\cancel2n}.\cancel2(y - Xw).\frac{\partial{(y - Xw)}}{\partial{w}} \\ &= \frac{1}{n}(y - Xw).X^T.(-1) \\ \therefore \frac{\partial{J(w)}}{\partial{w}} &= \frac{1}{n}X^T(Xw - y) \end{align*} \]Note: \(\frac{∂(a^{T}x)}{∂x}=a\)
Update parameter using gradient:
\[ w_{new} = w_{old} - \eta'. X^T(Xw - y) \]- Most important hyper parameter of gradient descent.
- Dictates the size of the steps taken down the cost function surface.
Small \(\eta\) ->
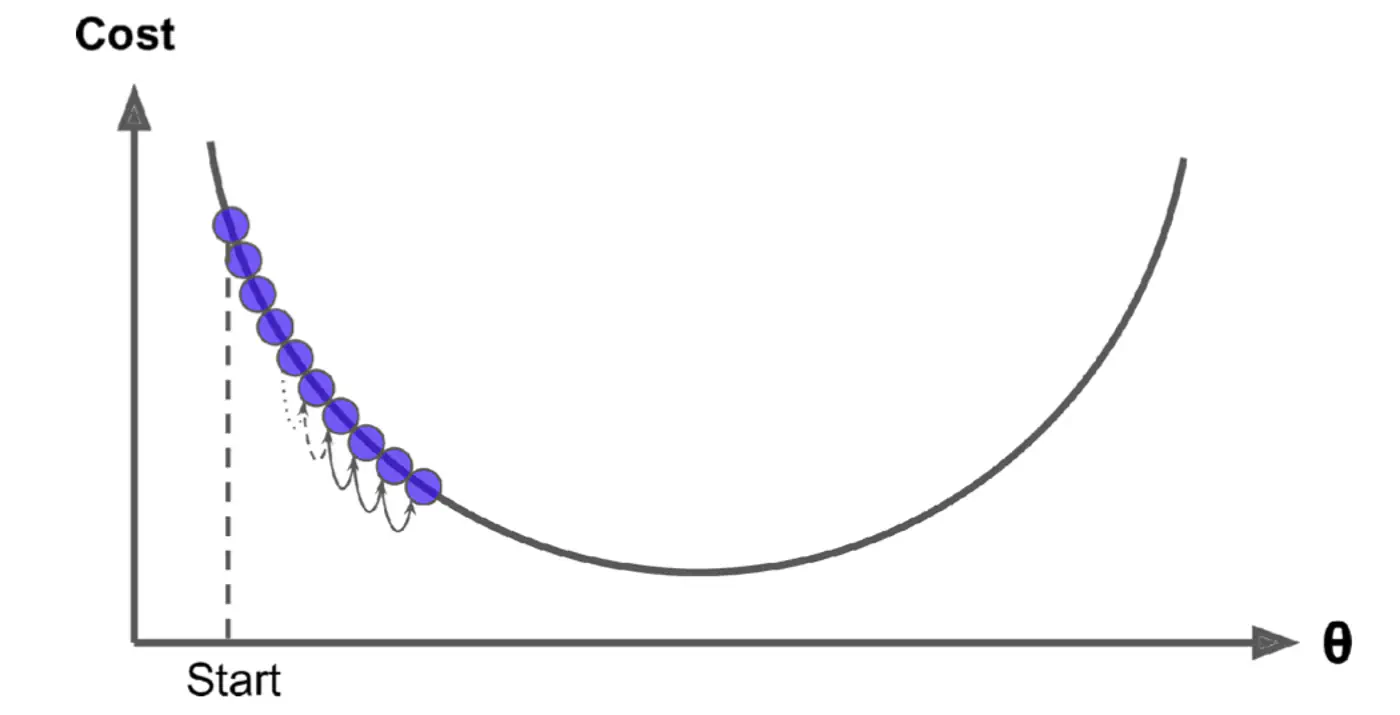
Large \(\eta\) ->
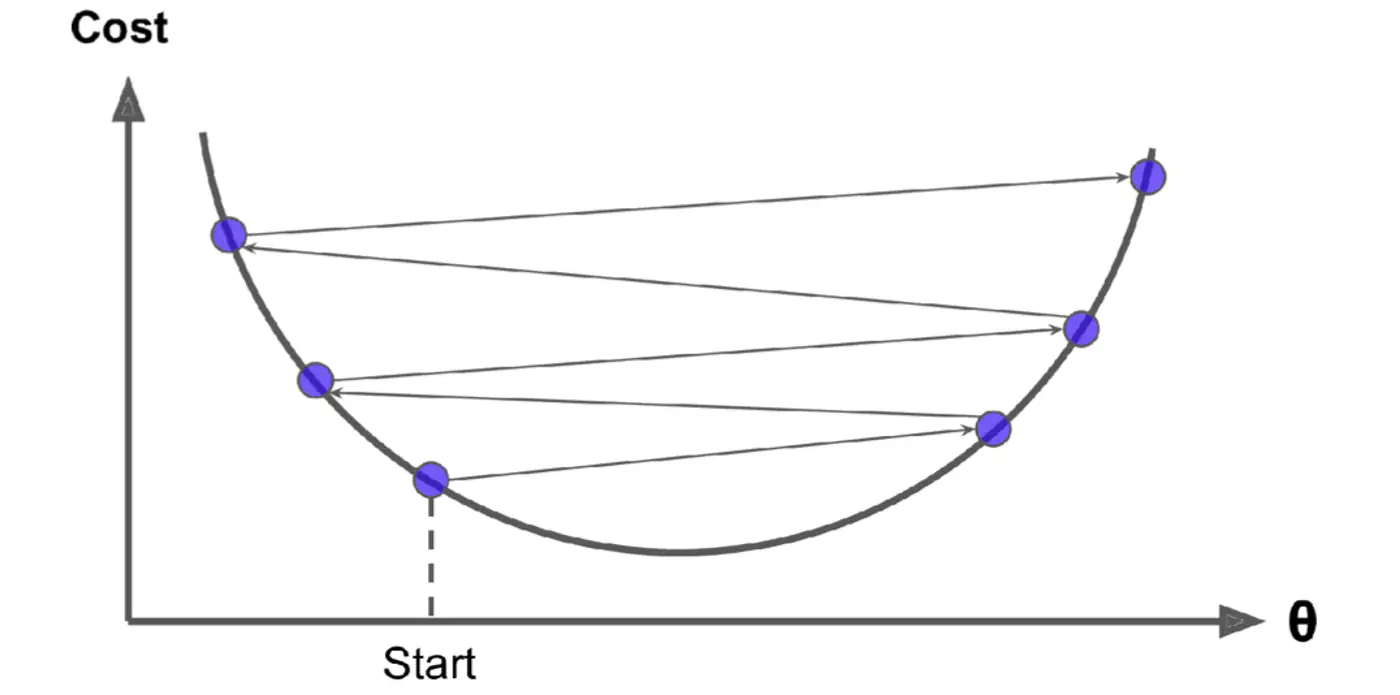
- Learning Rate Schedule:
The learning rate is decayed (reduced) over time.
Large steps initially and fine-tuning near the minimum, e.g., step decay or exponential decay. - Adaptive Learning Rate Methods:
Automatically adjust the learning rate for each parameter ‘w’ based on the history of gradients.
Preferred in modern deep learning as they require less manual tuning, e.g., Adagrad, RMSprop, and Adam.
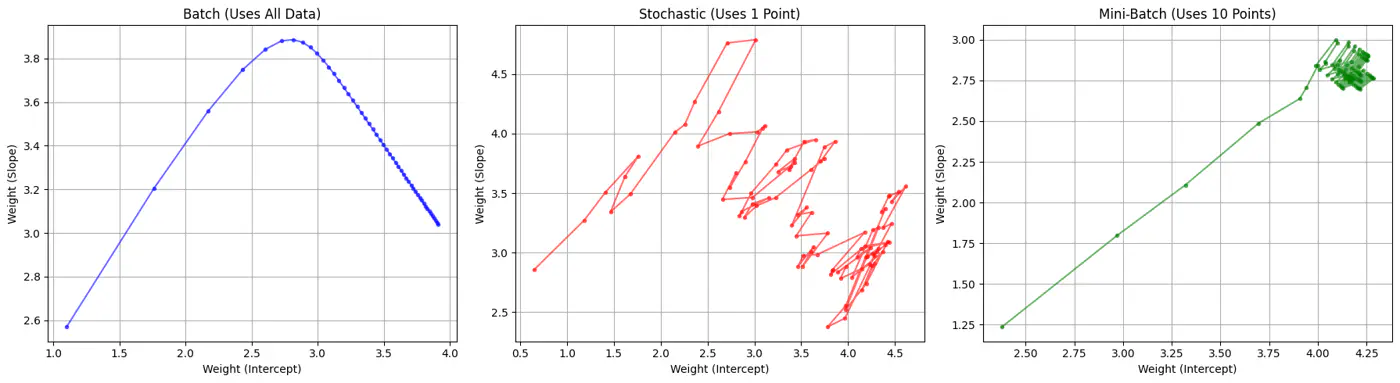
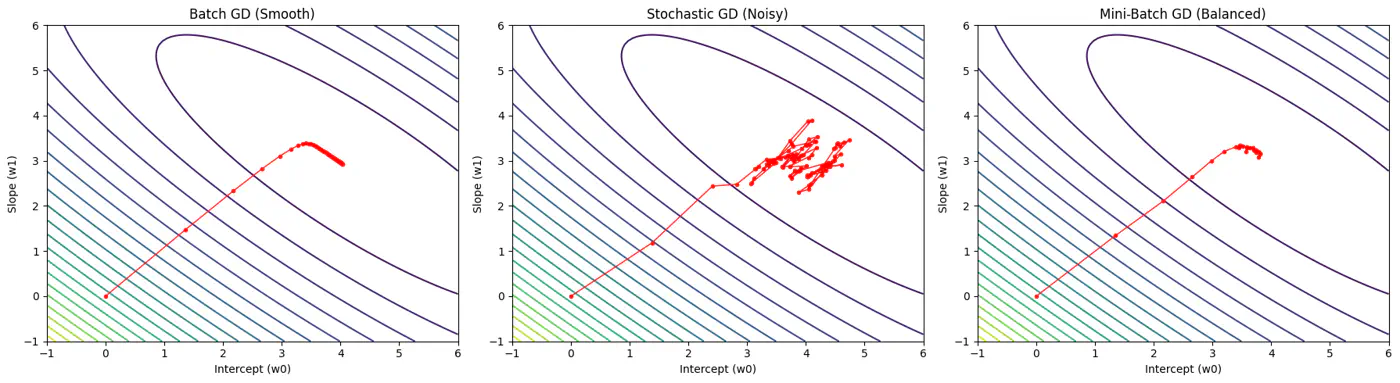
Batch, Stochastic, Mini-Batch are classified by data usage for gradient calculation in each iteration.
- Batch Gradient Descent (BGD): Entire Dataset
- Stochastic Gradient Descent (SGD): Random Point
- Mini-Batch Gradient Descent (MBGD): Subset
Computes the gradient using all the data points in the dataset for parameter update in each iteration.
\[w_{new} = w_{old} - \eta.\text{(average of all ’n’ gradients)}\]🔑Key Points:
- Slow 🐢 steps towards convergence, i.e, TC = O(n).
- Smooth, direct path towards minima.
- Minimum number of steps/iterations.
- Not suitable for large datasets; impractical for Deep Learning, as n = millions/billions.
Uses only 1 data point selected randomly from dataset to compute gradient for parameter update in each iteration.
\[w_{new} = w_{old} - \eta.\text{(gradient at any random data point)}\]🔑Key Points:
- Computationally fastest 🐇 per step; TC = O(1).
- Highly noisy, zig-zag path to minima.
- High variance in gradient estimation makes path to minima volatile, requiring a careful decay of learning rate to ensure convergence to minima.
- Uses small randomly selected subsets of dataset, called mini-batch, (1<k<n), to compute gradient for parameter update in each iteration. \[w_{new} = w_{old} - \eta.\text{(average gradient of ‘k' data points)}\]
🔑Key Points:
- Moderate time ⏰ consumption per step; TC = O(k<n).
- Less noisy, and more reliable convergence than stochastic gradient descent.
- More efficient and faster than batch gradient descent.
- Standard optimization algorithm for Deep Learning.Note: Vectorization on GPUs allows for parallel processing of mini-batches; also GPUs are the reason for the mini-batch size to be a power of 2.
End of Section
1.2.1.7 - Polynomial Regression
We can use a linear model to fit non-linear data.
Add powers of each feature as new features, then train a linear model on this extended set of features.
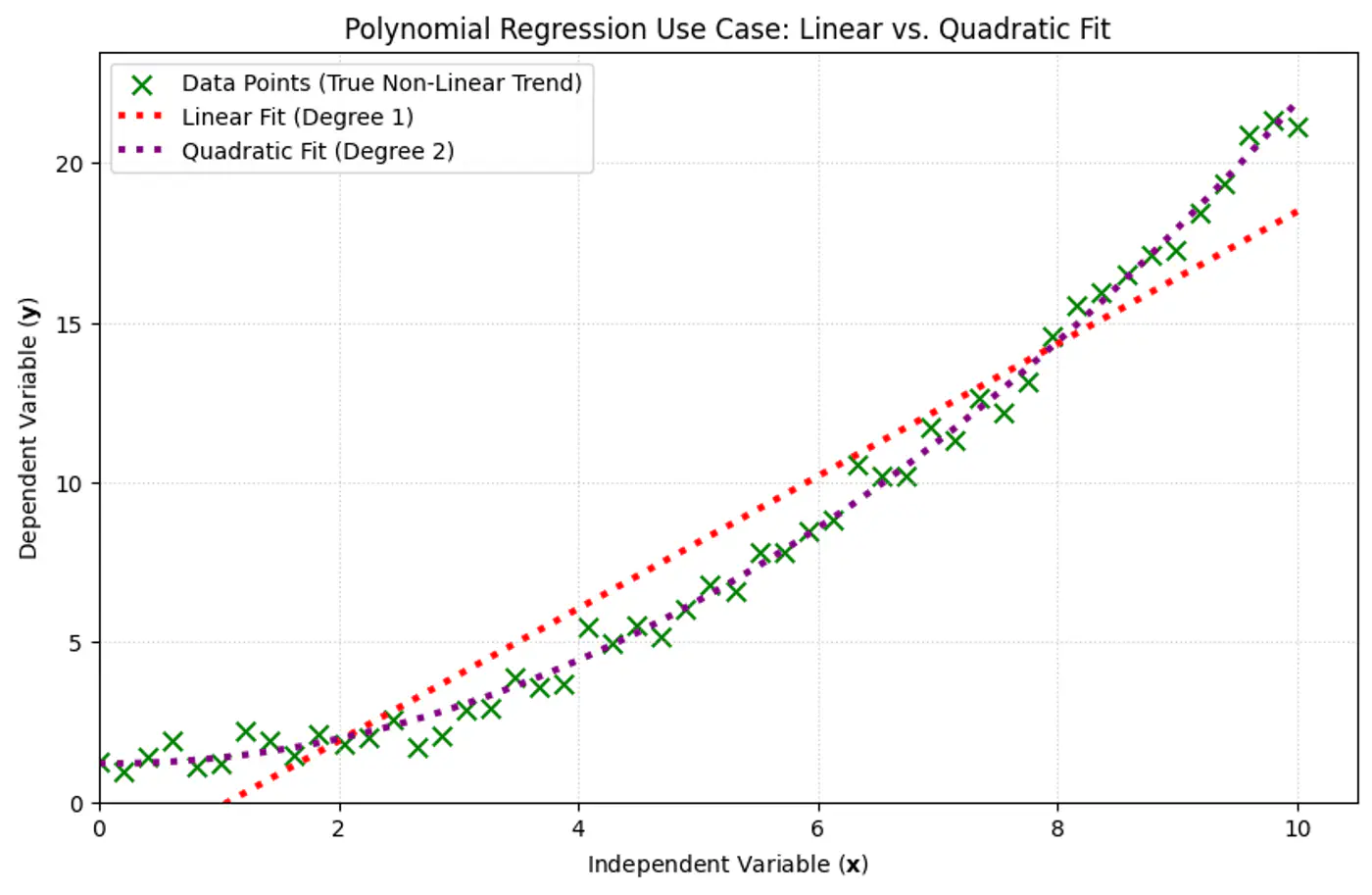
Linear: \(\hat{y_i} = w_0 + w_1x_{i_1} \)
Polynomial (quadratic): \(\hat{y_i} = w_0 + w_1x_{i_1} + w_2x_{i_1}^2\)
Polynomial (n-degree): \(\hat{y_i} = w_0 + w_1x_{i_1} + w_2x_{i_1}^2 +w_3x_{i_1}^3 + \dots + w_nx_{i_1}^n \)
Above polynomial can be re-written as linear equation:
\[\hat{y_i} = w_0 + w_1X_1 + w_2X_2 +w_3X_3 + \dots + w_nX_n \]where, \(X_1 = x_{i_1}, X_2 = x_{i_1}^2, X_3 = x_{i_1}^3, \dots, X_n = x_{i_1}^n\)
=> the model is still linear w.r.t to its parameters/weights \(w_0, w_1, w_2, \dots , w_n \).
e.g:
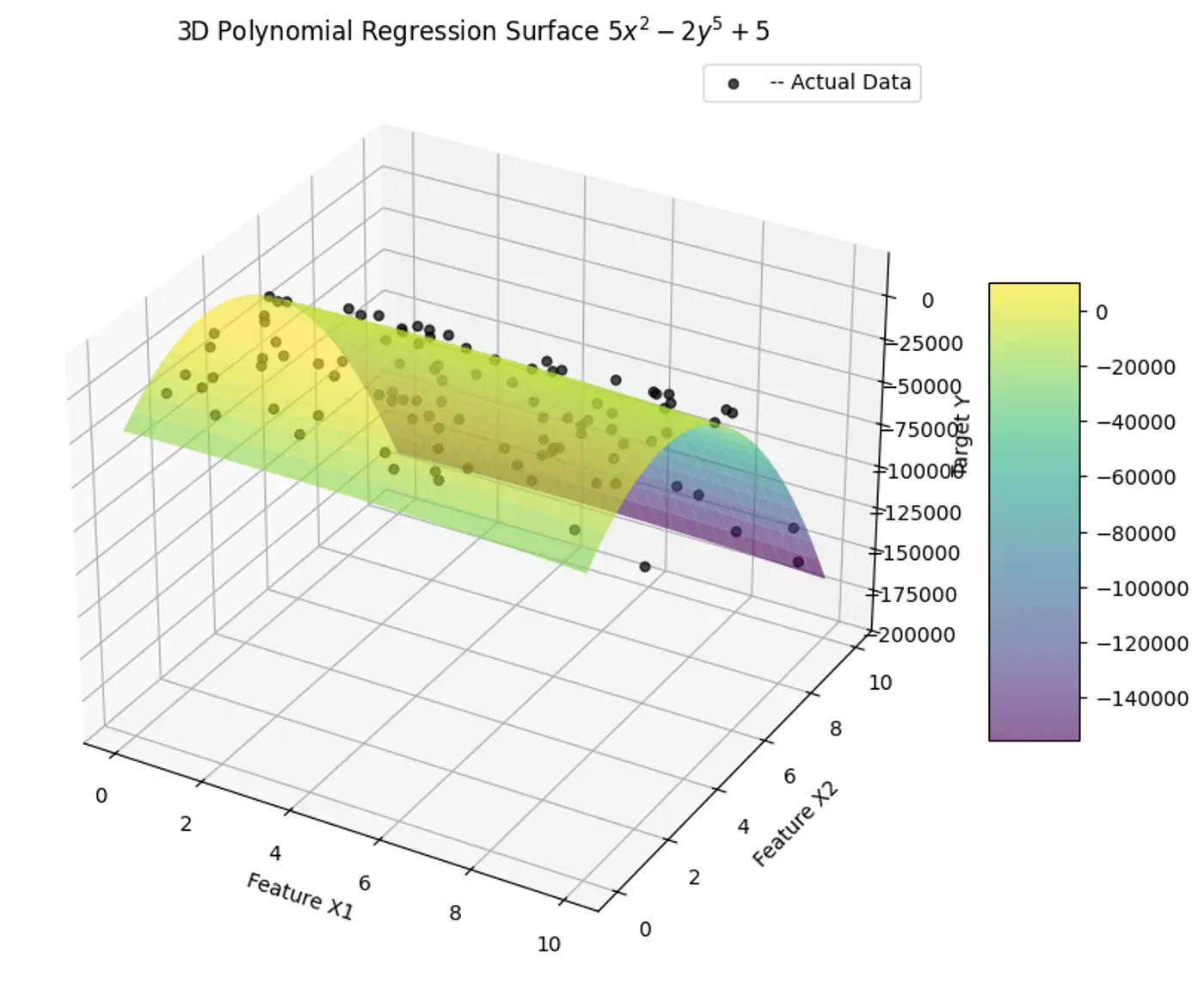
- Fit a linear model to the data points.
- Plot the errors.
- If the variance of errors is too high, then try polynomial features.
Note: Detect and remove outliers from error distribution.
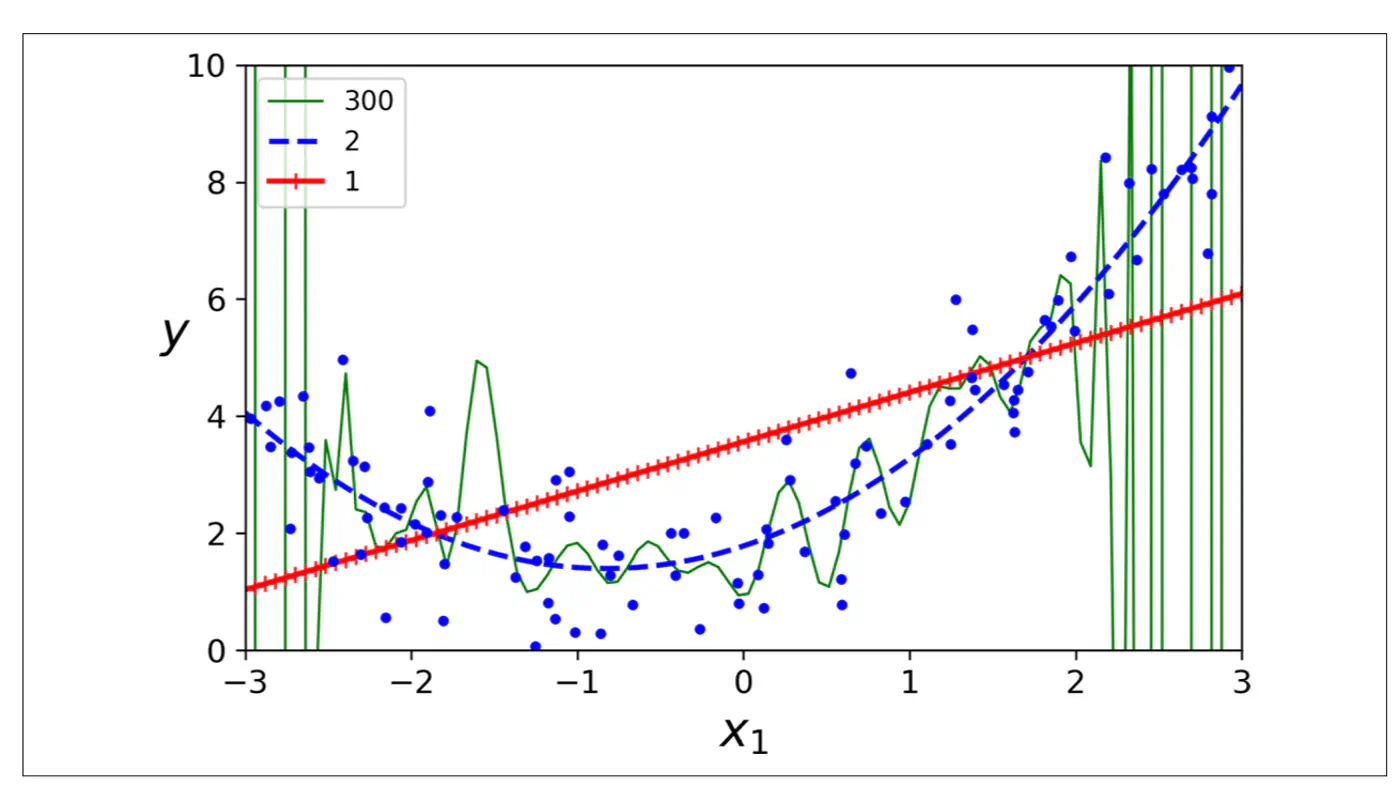
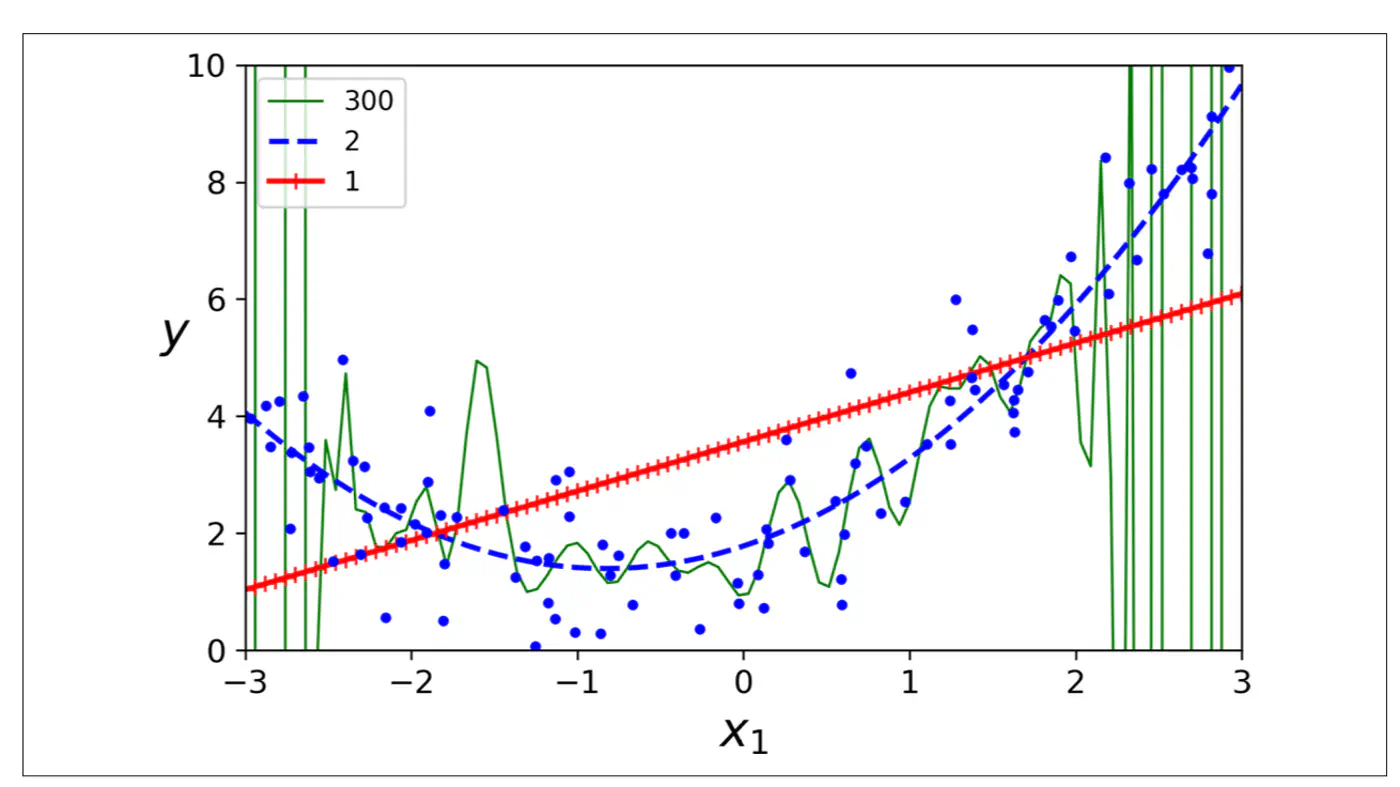
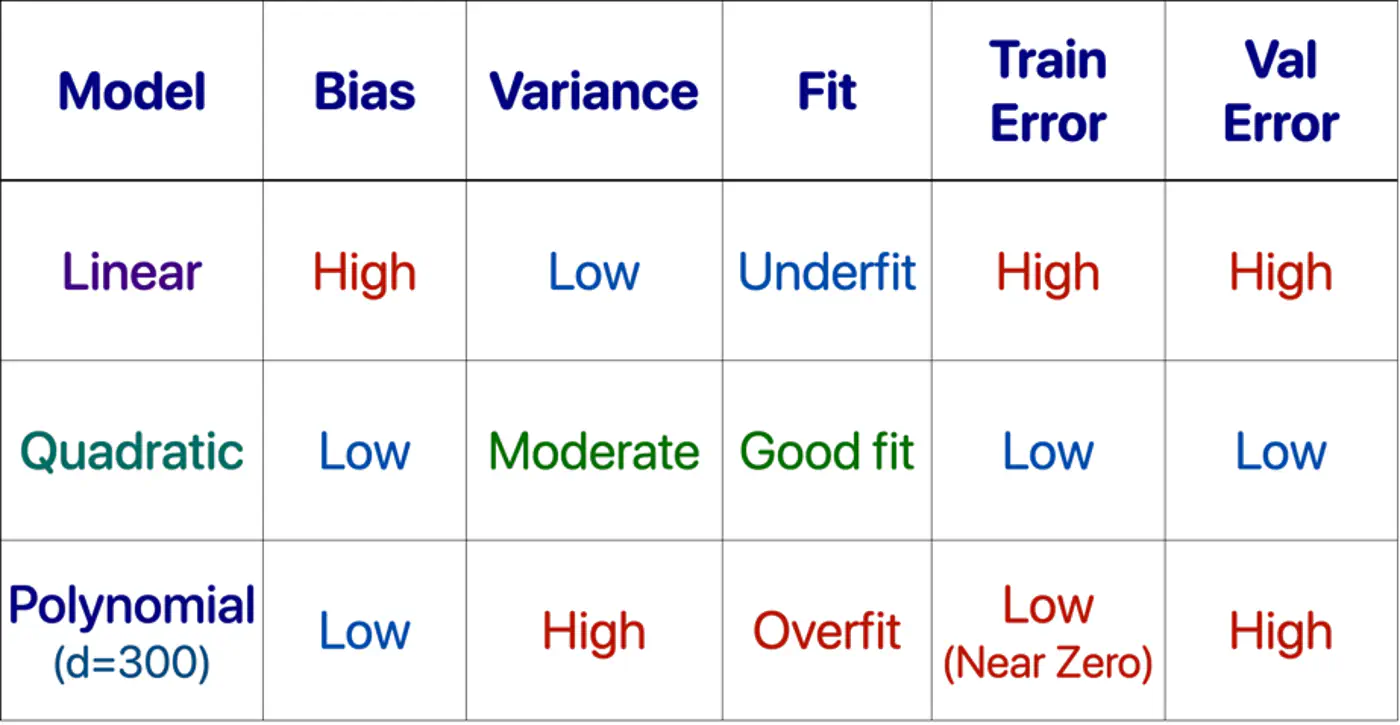
- Polynomial model : Over-fitting ❌
- Linear model : Under-fitting ❌
- Quadratic model: Generalizes best ✅
End of Section
1.2.1.8 - Data Splitting
- Training Data: Learn model parameters (Textbook + Practice problems)
- Validation Data: Tune hyper-parameters (Mock tests)
- Test Data: Evaluate model performance (Real (final) exam)
Data leakage occurs when information from the validation or test set is inadvertently used to train 🏃♂️ the model.
The model ‘cheats’ by learning to exploit information it should not have access to, resulting in artificially inflated performance metrics during testing 🧪.
- Small datasets(1K-100K): 60/20/20, 70/15/15 or 80/10/10
- Large datasets(>1M): 98/1/1 would suffice, as 1% of 1M is still 10K.
Note: There is no fixed rule, its trial and error.
If there is class imbalance in the dataset, (e.g., 95% class A , 5% class B), a random split might result in the validation set having 99% class A.
Solution: Use stratified sampling to ensure class proportions are maintained across all splits (train️/validation/test).
Note: Non-negotiable for imbalanced data.
- In time-series ⏰ data, divide the data chronologically, not randomly, i.e, training data time ⏰ should precede validation data time ⏰.
- We always train 🏃♂️ on past data to predict future data.
Golden rule: Never look 👀 into the future.
End of Section
1.2.1.9 - Cross Validation
Do not trust one split of the data; validate across many splits, and average the result to reduce randomness and bias.
Note: Two different splits of the same dataset can give very different validation scores.
Cross-validation is a statistical resampling technique used to evaluate how well a machine learning model generalizes to an independent, unseen dataset.
It works by systematically partitioning the available data into multiple subsets, or ‘folds’, and then training and testing the model on different combinations of these folds.
- K-Fold Cross-Validation
- Leave-One-Out Cross-Validation (LOOCV)
- Shuffle the dataset randomly (except time-series ⏳).
- Split data into k equal subsets(folds).
- Iterate through each unique fold, using it as the validation set.
- Use remaining k-1 fold for training 🏃♂️.
- Take an average of the results.Note: Common choice for k=5 or 10.
- Iteration 1: [V][T][T][T][T]
- Iteration 2: [T][V][T][T][T]
- Iteration 3: [T][T][V][T][T]
- Iteration 4: [T][T][T][V][T]
- Iteration 5: [T][T][T][T][V]
Model is trained 🏃♂️on all data points except one, and then tested 🧪on that remaining single observation.
LOOCV is an extreme case of k-fold cross-validation, where, k=n (number of data points).
Pros:
Useful for small (<1000) datasets.Cons:
Computationally 💻 expensive 💰.
End of Section
1.2.1.10 - Bias Variance Tradeoff
Mean Squared Error (MSE) = \(\frac{1}{n} \sum_{i=1}^n (y_i - \hat{y_i})^2\)
Total Error = Bias^2 + Variance + Irreducible Error
- Bias = Systematic Error
- Bias measures how far the average prediction of a model is from the true value.
- Variance = Sensitivity to Data
- Variance measures how much the predictions of a model vary for different training datasets.
- Irreducible Error = Sensor noise, Human randomness
- Inherent uncertainty in the data generation process itself and cannot be reduced by any model.
Systematic error from overly simplistic assumptions or strong opinion in the model.
e.g. House 🏠 prices 💰 = Rs. 10,000 * Area (sq. ft).
Note: This is over simplified view, because it ignores, amenities, location, age, etc.
Error from sensitivity to small fluctuations 📈 in the data.
e.g. Deep neural 🧠 network trained on a small dataset.
Note: Memorizes everything, including noise.
Say a house 🏠 in XYZ street was sold for very low price 💰.
Reason: Distress selling (outlier), or incorrect entry (noise).
Note: Model will make wrong(lower) price 💰predictions for all houses in XYZ street.
Goal 🎯 is to minimize total error.
Find a sweet-spot balance ⚖️ between Bias and Variance.
A good model ‘generalizes’ well, i.e.,
- Is not too simple or has a strong opinion.
- Does not memorize 🧠 everything in the data, including noise.
- Make model more complex.
- Add more features, add polynomial features.
- Decrease Regularization.
- Train 🏃♂️longer, the model has not yet converged.
- Add more data (most effective).
- Harder to memorize 🧠 1 million examples than 100.
- Use data augmentation, if getting more data is difficult.
- Increase Regularization.
- Early stopping 🛑, prevents memorization 🧠.
- Dropout (DL), randomly kill neurons, prevents co-adaptation.
- Use Ensembles.
- Averaging reduces variance.
Note: Co-adaptation refers to a phenomenon where neurons in a neural network become highly dependent on each other to detect features, rather than learning independently.
End of Section
1.2.1.11 - Regularization
Over-Fitting happens when we have a complex model that creates highly erratic curve to fit every single data point, including the random noise.
- Excellent performance on training 🏃♂️ data, but poor performance on unseen data.
- Penalty for overly complex models, i.e, models with excessively large or numerous parameters.
- Focus on learning general patterns rather than memorizing 🧠 everything, including noise in training 🏃♂️data.
Set of techniques that prevents over-fitting by adding a penalty term to the model’s loss function.
\[ J_{reg}(w) = J(w) + \lambda.\text{Penalty(w)} \]\(\lambda\) = Regularization strength hyper parameter, bias-variance tradeoff knob.
- High \(\lambda\): High ⬆️ penalty, forces weights towards 0, simpler model.
- Low \(\lambda\): Weak ⬇️ penalty, closer to un-regularized model.
- By intentionally simplifying a model (shrinking weights) to reduce its complexity, which prevents it from overfitting.
- Penalty pulls feature weights closer to zero, making the model less faithful representation of the training data’s true complexity.
- L2 Regularization (Ridge Regression)
- L1 Regularization (Lasso Regression)
- Elastic Net Regularization
- Early Stopping
- Dropout (Neural Networks)
- Penalty term: \(\ell_2\) norm - penalizes large weights quadratically.
- Pushes the weights close to 0 (not exactly 0), making models more stable by distributing importance across weights.
- Splits feature importance across co-related features.
- Use case: Best when most features are relevant and co-related.
- Also known as Ridge regression or Tikhonov regularization.
- Penalty term: \(\ell_1\) norm.
- Shrinks some weights exactly to 0, effectively performing feature selection, giving sparse solutions.
- For a group of highly co-related features, arbitrarily selects one feature and shrinks others to 0.
- Use case: Best when using high-dimensional datasets (d»n) where we suspect many features are irrelevant or redundant, or when model interpretability matters.
- Also known as Lasso (Least Absolute Shrinkage and Selection Operator) regression.
- Computational hack: \(\frac{\partial{|w_j|}}{\partial{w_j}} = 0\), since absolute function is not differentiable at 0.
- Penalty term : linear combination of and norm.
- Sparsity(feature-selection) of L1 and stability/grouping effect of L2 regularization.
- Use case: Best when we have high dimensional data with correlated features and we want sparse and stable solution.
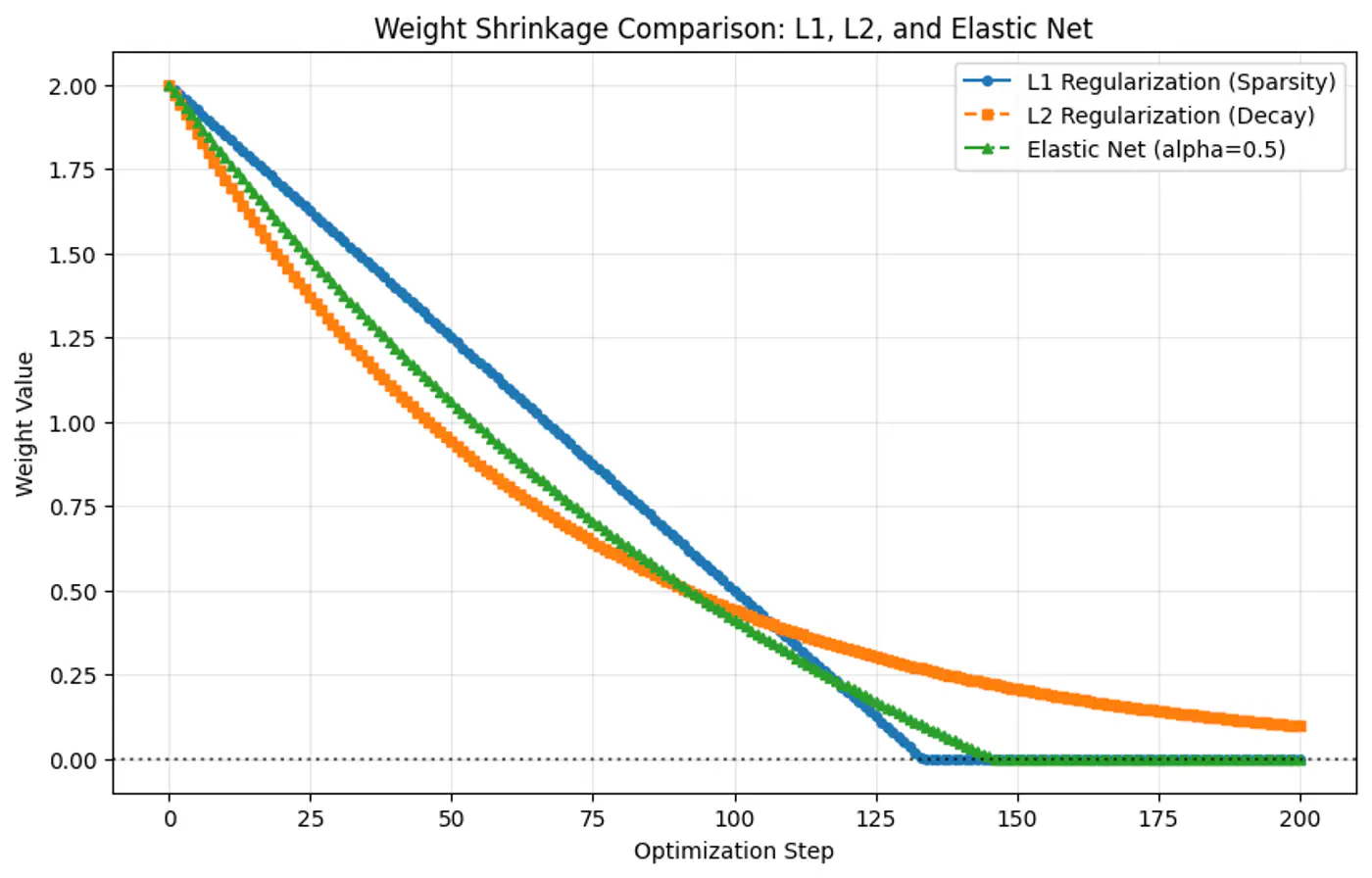
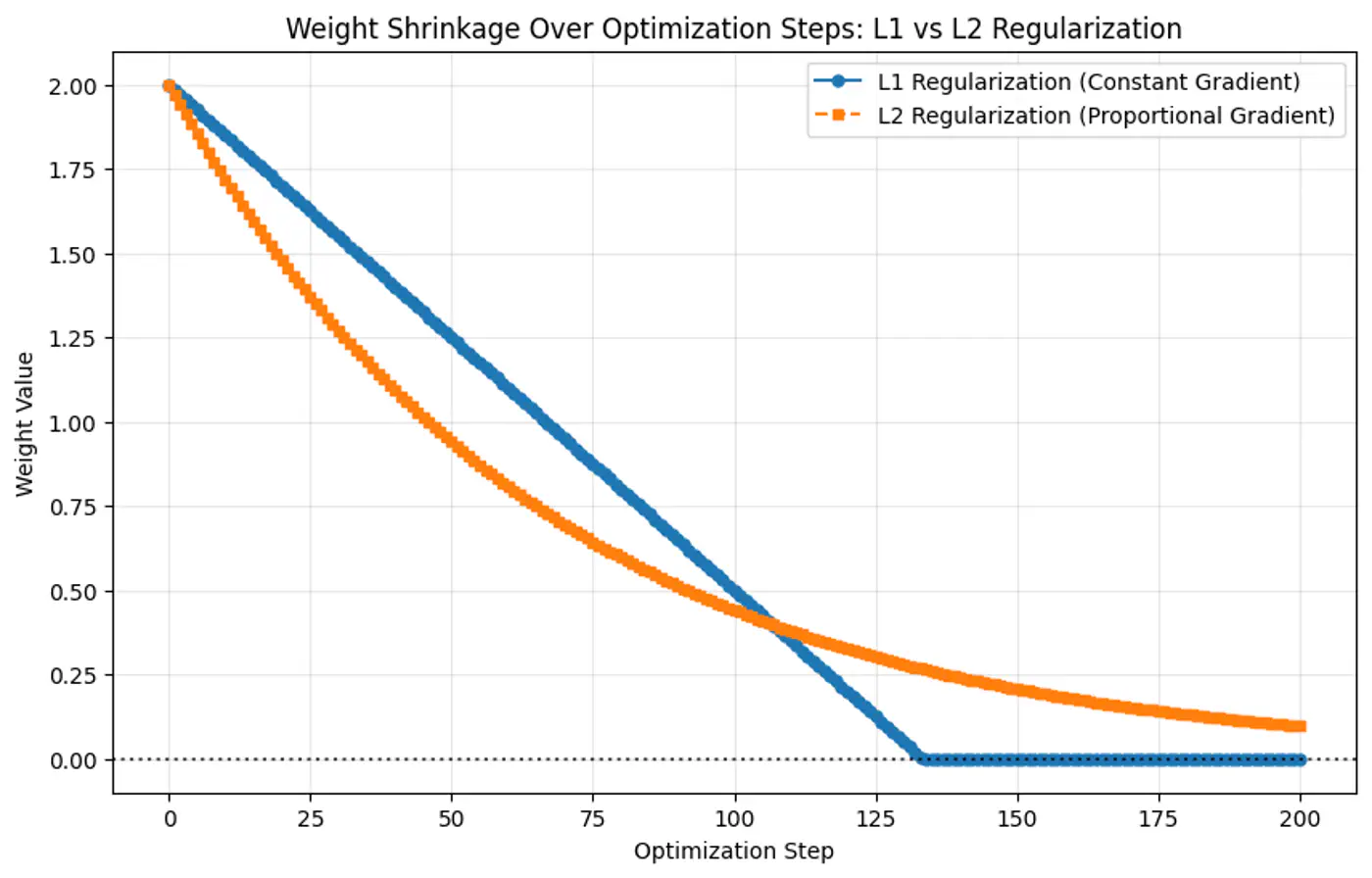
- Because the gradient of L1 penalty (absolute function) is a constant value, i.e, \(\pm 1\), this means a constant reduction in weight at each step, making it gradually reach to 0 in finite steps.
- Whereas, the derivative of L2 penalty is proportional to the weight (\(2w_j\)) and as the weight reaches close to 0, the gradient also becomes very small, this means that the weight will become very close to 0, but not exactly equal to 0.
End of Section
1.2.1.12 - Regression Metrics
MAE = \(\frac{1}{n} \sum_{i=1}^n |y_i - \hat{y_i}|\)
- Treats each error equally.
- Robust to outliers.
- Not differentiable x=0.
- Using gradient descent requires computational hack.
- Easy to interpret, as same units as target variable.
MSE = \(\frac{1}{n} \sum_{i=1}^n (y_i - \hat{y_i})^2\)
- Heavily penalizes large errors.
- Sensitive to outliers.
- Differentiable everywhere.
- Used by gradient descent and most other optimization algorithms.
- Difficult to interpret, as it has squared units.
RMSE = \(\sqrt{\frac{1}{n} \sum_{i=1}^n (y_i - \hat{y_i})^2}\)
- Easy to interpret, as it has same units as target variable.
- Useful when we need outlier-sensitivity of MSE but the interpretability of MAE.
Measures improvement over mean model.
\[ R^2 = 1 - \frac{SS_{res}}{SS_{tot}} = 1 - \frac{\sum_{i=1}^n (y_i - \hat{y_i})^2}{\sum_{i=1}^n (y_i - \bar{y_i})^2} \]Good R^2 value depends upon the use case, e.g. :
- Car 🚗 sale, R =0.8 is good enough.
- Cancer 🧪 prediction R 0.95, as life depends on it.
Range of values:
- Best value = 1
- Baseline value = 0
- Worst value = \(- \infty\)
Note: Example for bad model is all the points lie along x-axis and model predicts y-axis.
Quadratic for small errors; Linear for large errors.
\[ L_{\delta}(y, \hat{y}) = \begin{cases} \frac{1}{2}(y - \hat{y})^2 & \text{for } |y - \hat{y}| \le \delta \\ \\ \delta (|y - \hat{y}| - \frac{1}{2}\delta) & \text{otherwise} \end{cases} \]- Robust to outliers.
- Differentiable at 0; smooth convergence to minima.
- Delta (\(\delta\)) knob(hyper parameter) to control.
- \(\delta\) high: MSE
- \(\delta\) low: MAE
Note: Tune \(\delta\): MAE, for outliers > \(\delta\); MSE, for small errors < \(\delta\).
e.g: = 95th percentile of errors or 1.35\(\sigma\) for standard Gaussian data.
Huber loss (Green) and Squared loss (blue)
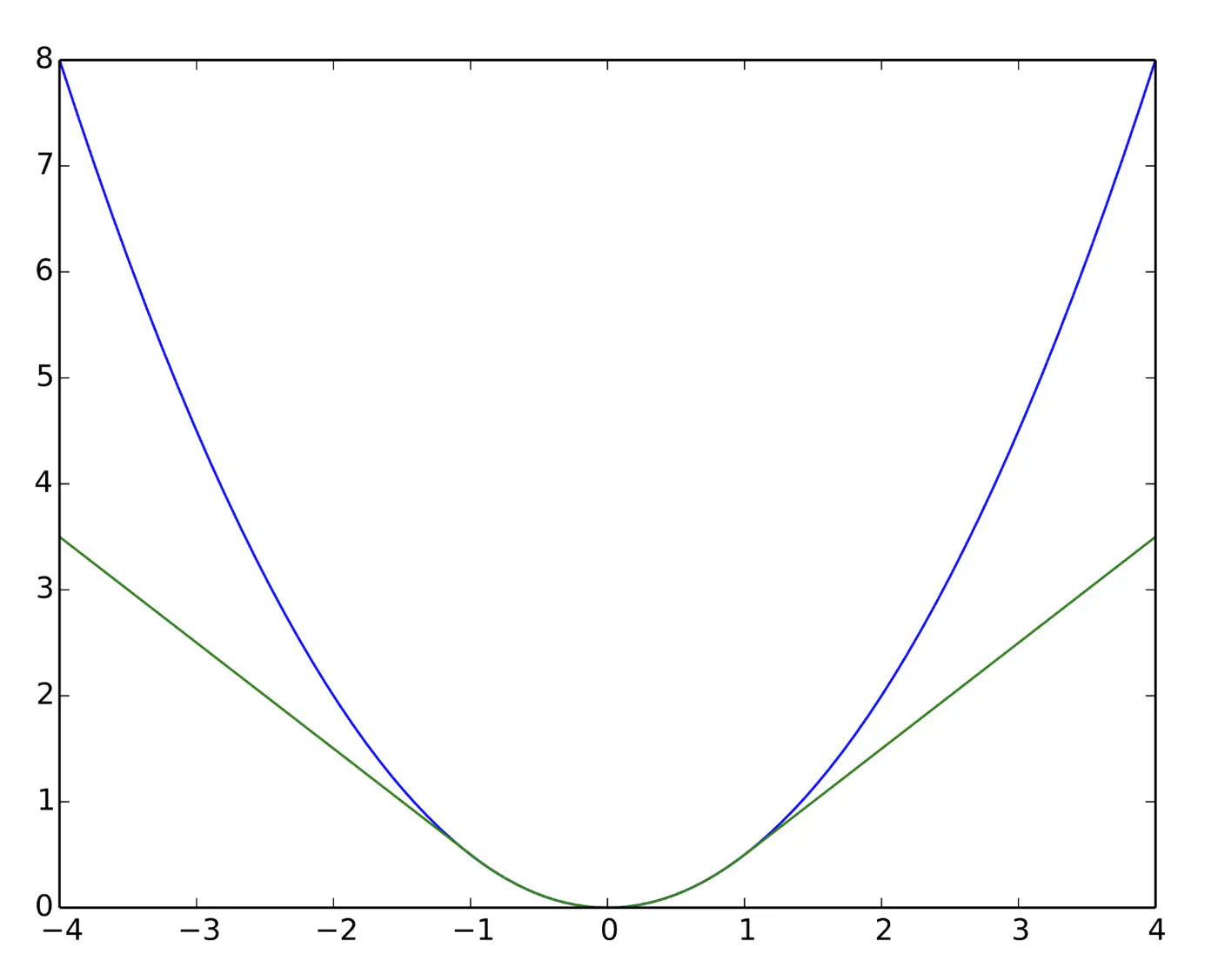
End of Section
1.2.1.13 - Assumptions
Linear Regression works reliably only when certain key 🔑 assumptions about the data are met.
- Linearity
- Independence of Errors (No Auto-Correlation)
- Homoscedasticity (Equal Variance)
- Normality of Errors
- No Multicollinearity
- No Endogeneity (Target not correlated with the error term)
Relationship between features and target 🎯 is linear in parameters.
Note: Polynomial regression is linear regression.
\(y=w_0 +w_1x_1+w_2x_2^2 + w_3x_3^3\)
Residuals (errors) should not have a visible pattern or correlation with one another (most common in time-series ⏰ data).
Risk:
If errors are correlated, standard errors will be underestimated, making variables look ‘statistically significant’ when they are not.
Test:
Durbin–Watson test
Autocorrelation plots (ACF)
Residuals vs time
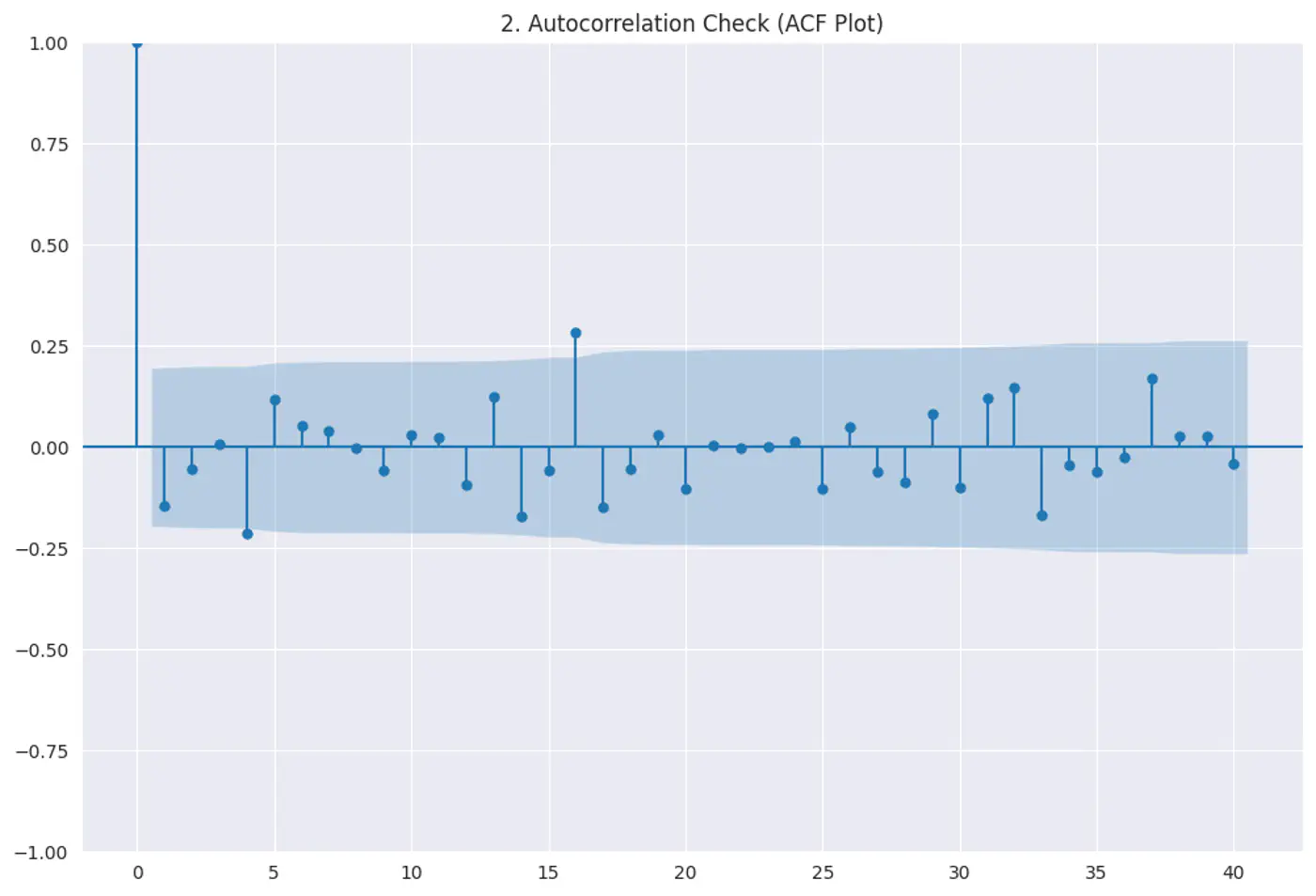
Constant variance of errors; Var(ϵ|X)=σ
Risk:
Standard errors become biased, leading to unreliable hypothesis tests (t-tests, F-tests).
Test:
- Breusch–Pagan test
- White test
Fix:
- Log transform
- Weighted Least Squares(WLS)
Error terms should follow a normal distribution; (Required for small datasets.)
Note: Because of Central Limit Theorem, with a large enough sample size, this becomes less critical for estimation.
Risk: Hypothesis testing (calculating p-values and confidence intervals), we assume the error terms follow a normal distribution.
Test:
Q-Q plot
Shapiro-Wilk Test
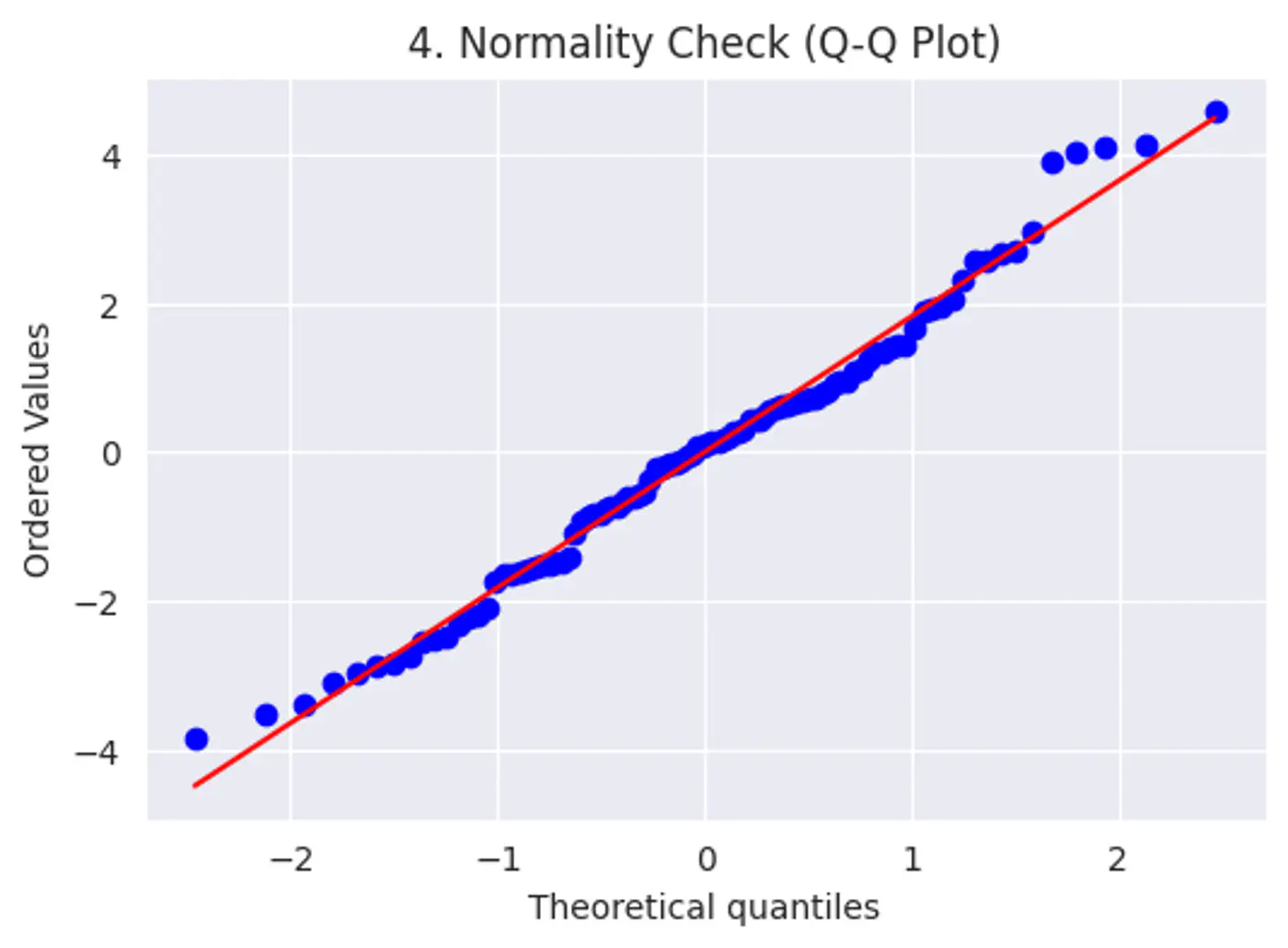
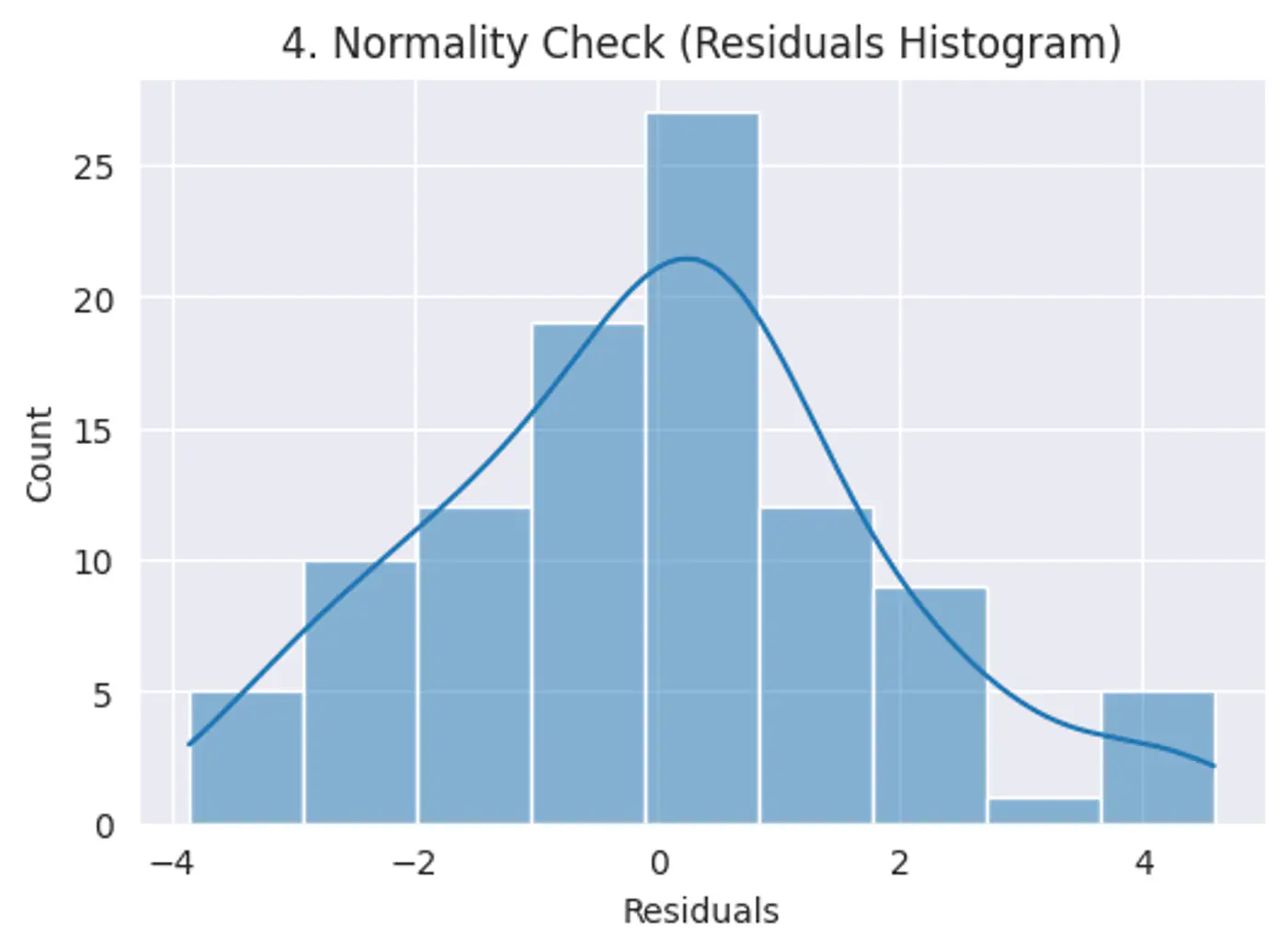
Features should not be highly correlated with each other.
Risk:
- High correlation makes it difficult to determine the unique, individual impact of each feature. This leads to high variance in model parameter estimates, small changes in data cause large swings in parameters.
- Model interpretability issues.
Test:
- Variance Inflation Factor(VIF)VIF > 5 → concern, VIF > 10 → serious issue
Fix:
- PCA
- Remove redundant features
Error term must be uncorrelated with the features; E[ϵ|X] = 0
Risk:
- Parameters will be biased and inconsistent.
Test:
- Hausman Test
- Durbin-Wu-Hausman (DWH) Test
Fix:
- Controlled experiments.
End of Section
1.2.2.1 - Binary Classification
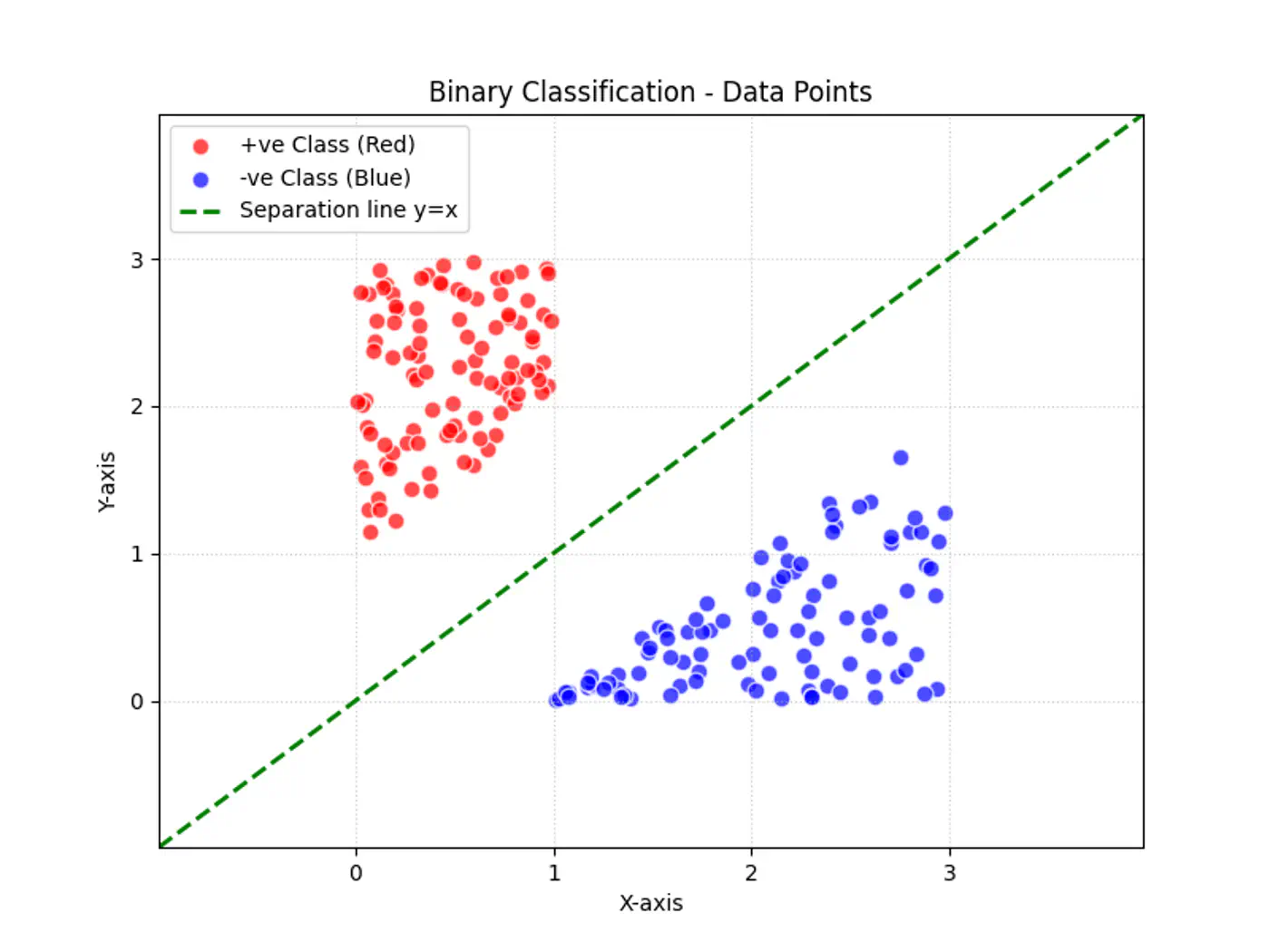
Linear regression tries to find the best fit line, but we want to find the line or decision boundary that clearly separates the two classes.
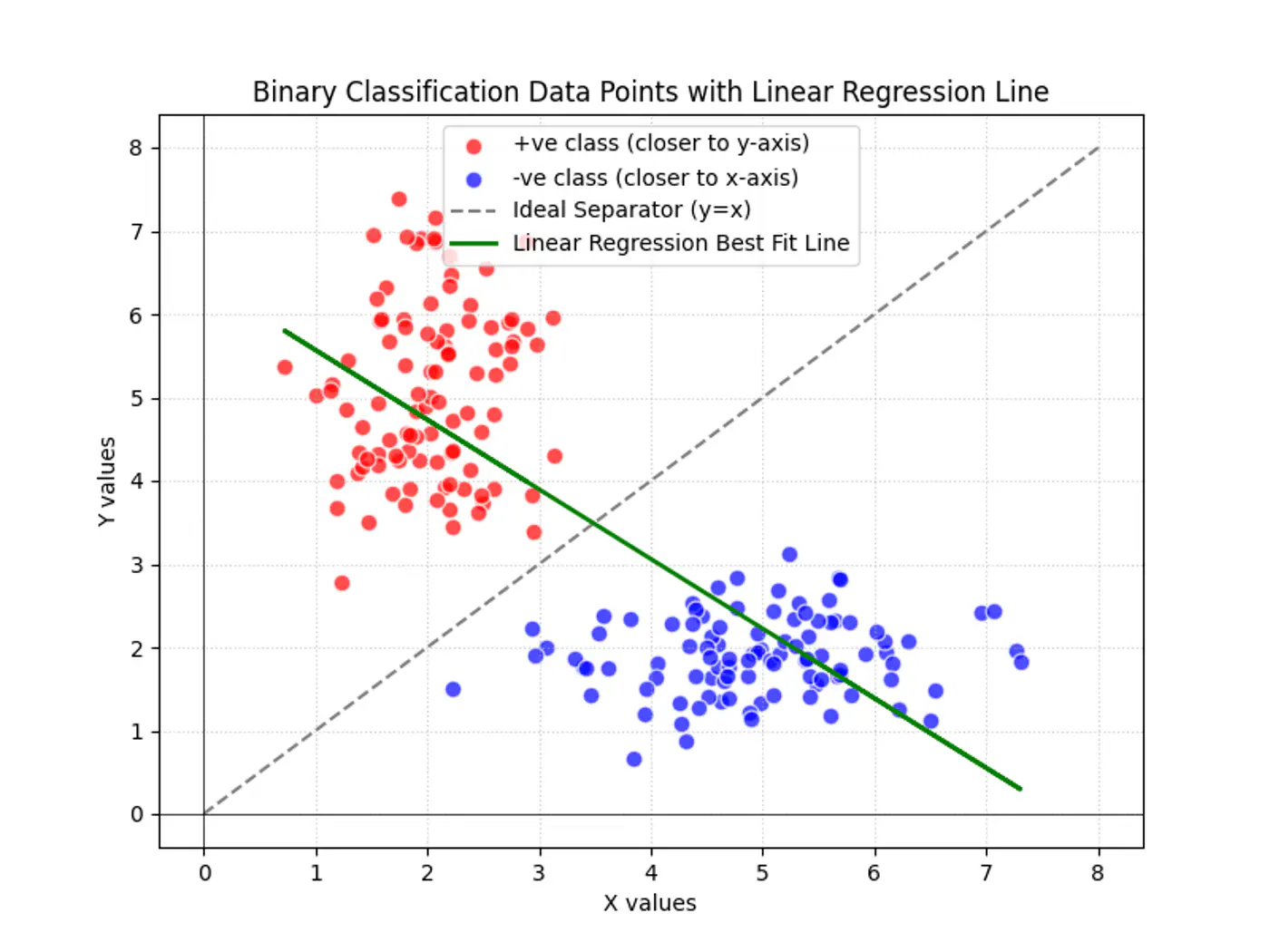
Find the decision boundary, i.e, the equation of the separating hyperplane.
\[z=w^{T}x+w_{0}\]Value of \(z = \mathbf{w^Tx} + w_0\) tells us how far is the point from the decision boundary and on which side.
Note: Weight 🏋️♀️ vector ‘w’ is normal/perpendicular to the hyperplane, pointing towards the positive class (y=1).
- For points exactly on the decision boundary \[z = \mathbf{w^Tx} + w_0 = 0 \]
- Positive (+ve) labeled points \[ z = \mathbf{w^Tx} + w_0 > 0 \]
- Negative (-ve) labeled points \[ z = \mathbf{w^Tx} + w_0 < 0 \]
So we need a link 🔗 to transform the geometric distance to probability.
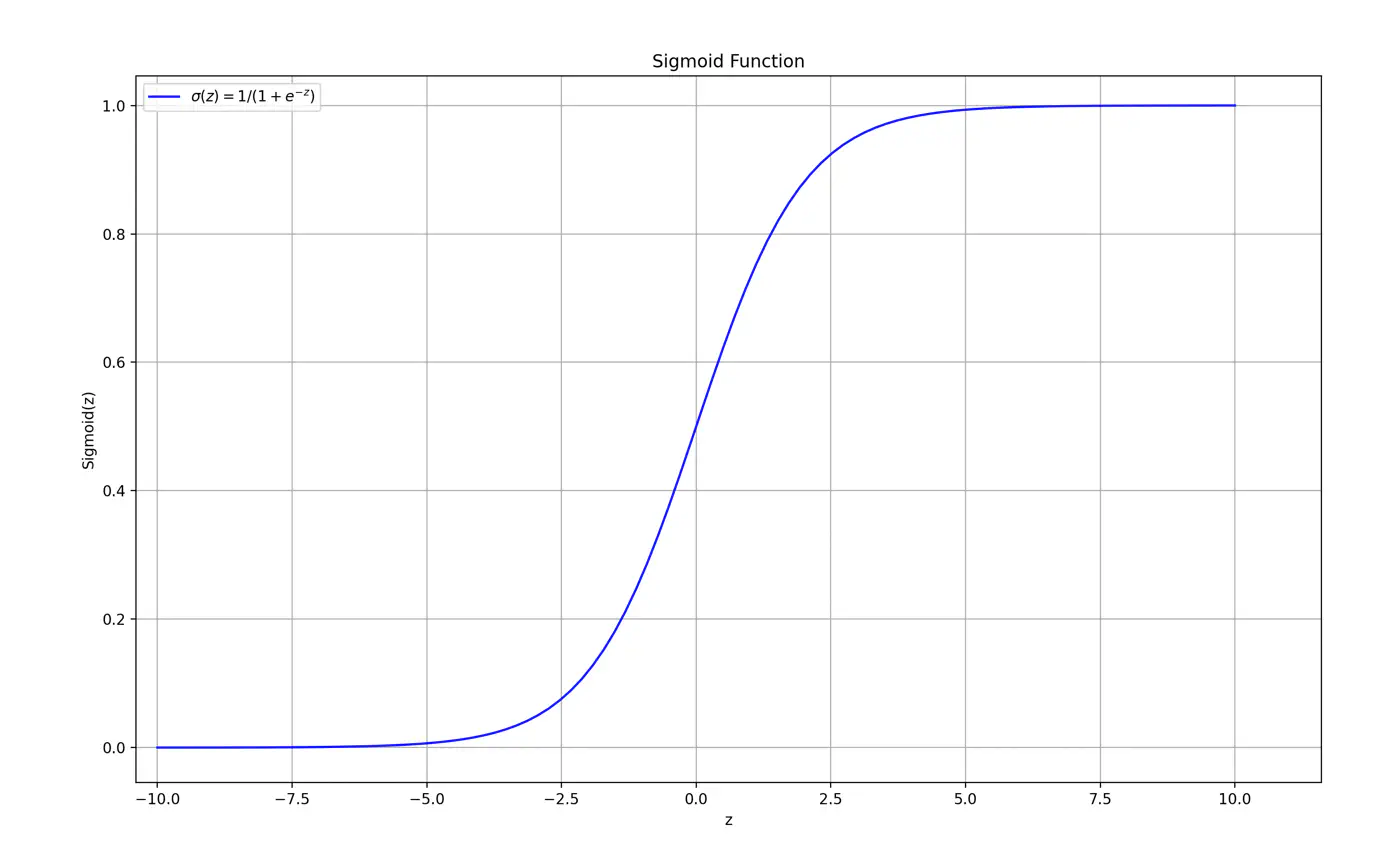
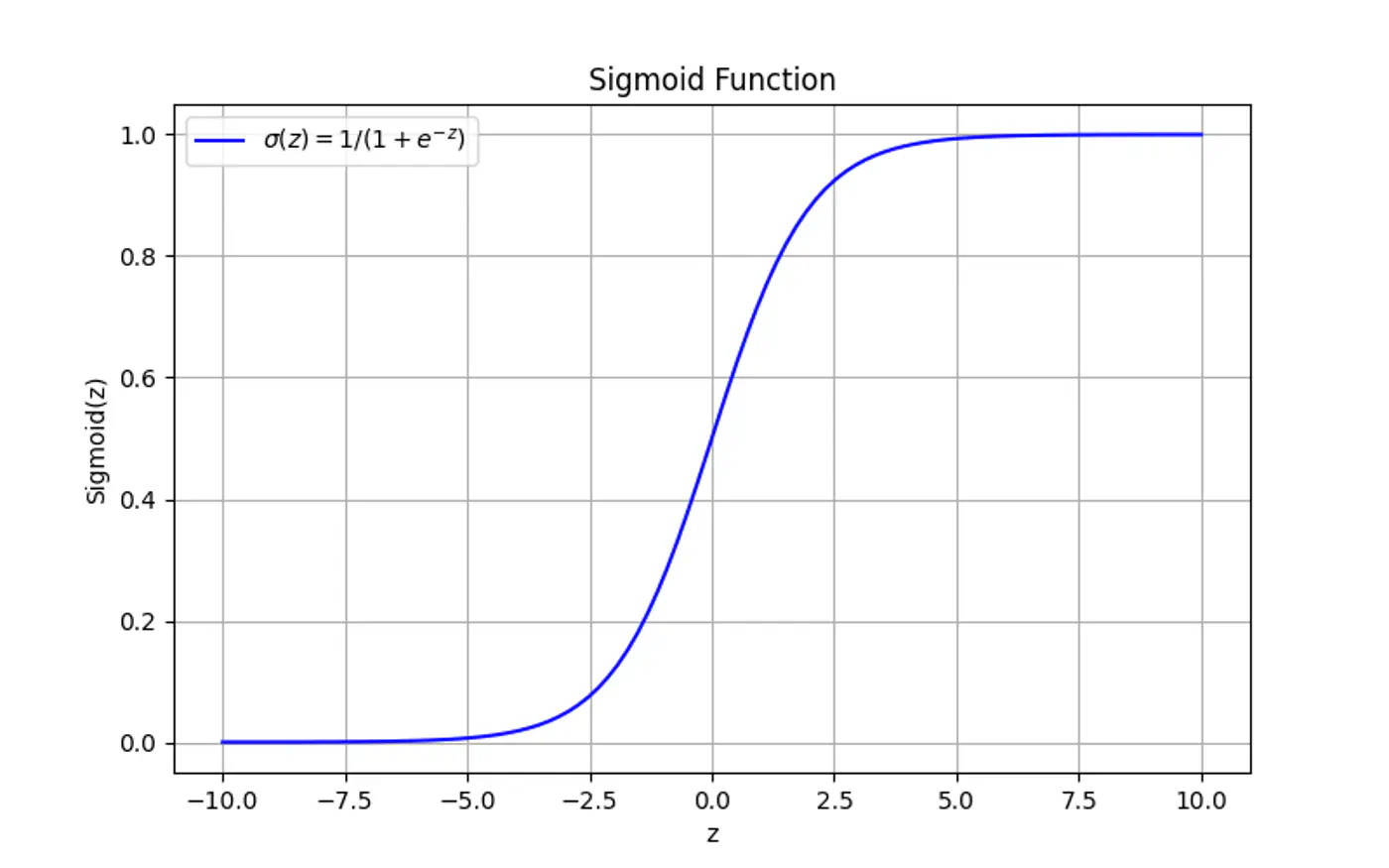
Maps the output of a linear equation to a value between 0 and 1, allowing the result to be interpreted as a probability.
\[\hat{y} = \sigma(z) = \frac{1}{1 + e^{-z}}\]If the distance ‘z’ is large and positive, \(\hat{y} \approx 1\) (High confidence).
If the distance ‘z’ is 0, \(\hat{y} = 0.5\) (Maximum uncertainty).
End of Section
1.2.2.2 - Log Loss
Log Loss = \(\begin{cases} -log(\hat{y_i}) & \text{if } y_i = 1 \\ \\ -log(1-\hat{y_i}) & \text{if } y_i = 0 \end{cases} \)
Combining the above 2 conditions into 1 equation gives:
Log Loss = \(-[y_ilog(\hat{y_i}) + (1-y_i)log(1-\hat{y_i})]\)
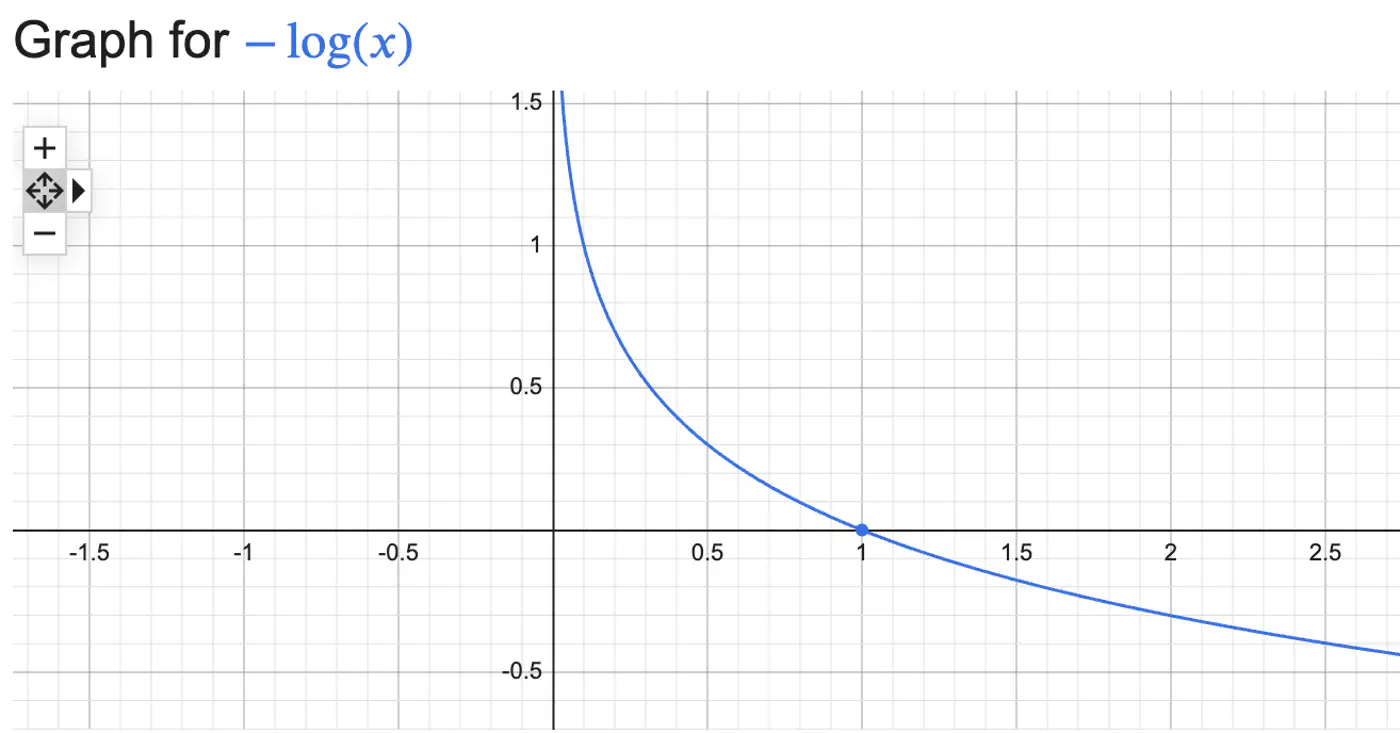
We need to find the weights 🏋️♀️ ‘w’ that minimize the cost 💵 function.
- Weight update: \[w_{new}=w_{old}-η.\frac{∂J(w)}{∂w_{old}}\]
We need to find the gradient of log loss w.r.t weight ‘w’.
Chain Rule:
\[\frac{\partial{J(w)}}{\partial{w}} = \frac{\partial{J(w)}}{\partial{\hat{y}}}.\frac{\partial{\hat{y}}}{\partial{z}}.\frac{\partial{z}}{\partial{w}}\]- Cost Function: \(J(w) = -\frac{1}{n}\sum [y_ilog(\hat{y_i}) + (1-y_i)log(1-\hat{y_i})]\)
- Prediction: \(\hat{y} = \sigma(z) = \frac{1}{1 + e^{-z}}\)
- Distance of Point: \(z = \mathbf{w^Tx} + w_0\)
How loss changes w.r.t prediction ?
\[ \begin{align*} \frac{\partial{J(w)}}{\partial{\hat{y}}} &= - [\frac{y}{\hat{y}} - \frac{1-y}{1-\hat{y}}] \\ &= -[\frac{y- \cancel{y\hat{y}} -\hat{y} + \cancel{y\hat{y}}}{\hat{y}(1-\hat{y})}] \\ \therefore \frac{\partial{J(w)}}{\partial{\hat{y}}} &= \frac{\hat{y} - y}{\hat{y}(1-\hat{y})} \end{align*} \]How prediction changes w.r.t distance ?
\[ \begin{align*} \frac{\partial{\hat{y}}}{\partial{z}} &= \frac{\partial{\sigma(z)}}{\partial{z}} = \sigma'(z) \\ \sigma'(z) &= \sigma(z)(1-\sigma(z)) \\ \therefore \frac{\partial{\hat{y}}}{\partial{z}} &= \hat{y}(1-\hat{y}) \end{align*} \]from equations (1) & (2):
\[ \begin{align*} \because \frac{\partial{\sigma(z)}}{\partial{z}} &= \frac{\partial{\sigma(z)}}{\partial{u}}. \frac{\partial{u}}{\partial{z}} \\ \implies \frac{\partial{\sigma(z)}}{\partial{z}} &= - \frac{1}{(1 + e^{-z})^2}. -e^{-z} = \frac{e^{-z}}{(1 + e^{-z})^2} \\ 1 - \sigma(z) & = 1 - \frac{1}{1 + e^{-z}} = \frac{e^{-z}}{1 + e^{-z}} \\ \frac{\partial{\sigma(z)}}{\partial{z}} &= \frac{1}{1 + e^{-z}}.\frac{e^{-z}}{1 + e^{-z}} \\ \therefore \frac{\partial{\sigma(z)}}{\partial{z}} &= \sigma(z).(1-\sigma(z)) \end{align*} \]How distance changes w.r.t weight 🏋️♀️ ?
\[ \frac{\partial{z}}{\partial{w}} = \mathbf{x} \]\[\because \frac{\partial{(a^T\mathbf{x})}}{\partial{\mathbf{x}}} = a\]Chain Rule:
\[ \begin{align*} \frac{\partial{J(w)}}{\partial{w}} &= \frac{\partial{J(w)}}{\partial{\hat{y}}}.\frac{\partial{\hat{y}}}{\partial{z}}.\frac{\partial{z}}{\partial{w}} \\ &= \frac{\hat{y} - y}{\cancel{\hat{y}(1-\hat{y})}}.\cancel{\hat{y}(1-\hat{y})}.x \\ \therefore \frac{\partial{J(w)}}{\partial{w}} &= (\hat{y} - y).x \end{align*} \]Gradient = Error x Input
- Error = \((\hat{y_i}-y_i)\): how far is prediction from the truth?
- Input = \(x_i\): contribution of specific feature to the error.
Weight update:
\[w_{new} = w_{old} - \eta. \sum_{i=1}^n (\hat{y_i} - y_i).x_i\]Mean Squared Error (MSE) can not be used to quantify error/loss in binary classification because:
- Convexity : MSE combined with Sigmoid is non-convex, so, Gradient Descent can get trapped in local minima.
- Penalty: MSE does not appropriately penalize mis-classifications in binary classification.
- e.g: If actual value is class 1 but the model predicts class 0, then MSE = \((1-0)^2 = 1\), which is very low, whereas los loss = \(-log(0) = \infty\)
End of Section
1.2.2.3 - Regularization
The weights 🏋️♀️ will tend towards infinity, preventing a stable solution.
The model tries to make probabilities exactly 0 or 1, but the sigmoid function never reaches these limits, leading to extreme weights 🏋️♀️ to push probabilities near the extremes.
Distance of Point: \(z = \mathbf{w^Tx} + w_0\)
Prediction: \(\hat{y} = \sigma(z) = \frac{1}{1 + e^{-z}}\)
Log loss: \(-[y_ilog(\hat{y_i}) + (1-y_i)log(1-\hat{y_i})] \)
Model becomes perfectly accurate on training 🏃♂️data but fails to generalize, performing poorly on unseen data.
Adds a penalty term to the loss function, discouraging weights 🏋️♀️ from becoming too large.
End of Section
1.2.2.4 - Log Odds
Odds compare the likelihood of an event happening vs. not happening.
Odds = \(\frac{p}{1-p}\)
- p = probability of success
In logistic regression we assume that Log-Odds (the log of the ratio of positive class to negative class) is a linear function of inputs.
Log-Odds (Logit) = \(log_e \frac{p}{1-p}\)
Log Odds = \(log_e \frac{p}{1-p} = z\)
\[z=w^{T}x+w_{0}\]\[ \begin{align*} &log_{e}(\frac{p}{1-p}) = z \\ &⟹\frac{p}{1-p} = e^{z} \\ &\implies p = e^z - p.e^z \\ &\implies p = \frac{e^z}{1+e^z} \\ &\text { divide numerator and denominator by } e^z \\ &\implies p = \frac{1}{1+e^{-z}} \quad \text { i.e, Sigmoid function} \end{align*} \]- Probability: 0 to 1
- Odds: 0 to + \(\infty\)
- Log Odds: -\(\infty\) to +\(\infty\)
End of Section
1.2.2.5 - Probabilistic Interpretation
We assume that our target variable ‘y’ follows a Bernoulli distribution, i.e, has exactly 2 outputs success/failure.
- P(Y=1|X) = p
- P(Y=0|X) = 1- p
Combining above 2 into 1 equation gives:
- P(Y=y|X) = \(p^y(1-p)^{1-y}\)
‘Find the most plausible explanation for what I see.’
We want to find the weights 🏋️♀️‘w’ that maximize the likelihood of seeing the data.
- Data, D = \(\{ (x_i, y_i)_{i=1}^n , \quad y_i \in \{0,1\} \}\)
We do this by maximizing likelihood function.
Assumption: Training data is I.I.D.
A common simplification is to maximize the log-likelihood function instead, which converts the product into a sum.
Note: Log is a strictly monotonically increasing function.
Maximizing the log-likelihood is same as minimizing the negative of log-likelihood.
\[ \begin{align*} \underset{w}{\mathrm{max}}\ log\mathcal{L}(w) &= \underset{w}{\mathrm{min}} - log\mathcal{L}(w) \\ \underset{w}{\mathrm{min}} - log\mathcal{L}(w) &= - \sum_{i=1}^n [ y_ilog(p_i) + (1-y_i)log(1-p_i)] \\ \underset{w}{\mathrm{min}} - log\mathcal{L}(w) &= \text {Log Loss} \end{align*} \]End of Section
1.2.3 - K Nearest Neighbors
End of Section
1.2.3.1 - KNN Introduction
- Parametric models:
- Rely on assumption that relationships between data points are linear.
- For polynomial regression we need to find the degree of polynomial.
- Training:
- We need to train 🏃♂️the model for prediction.
Simple: Intuitive way to classify data or predict values by finding similar existing data points (neighbors).
Non-Parametric: Makes no assumptions about the underlying data distribution.
No Training Required: KNN is a ‘lazy learner’, it does not require a formal training 🏃♂️ phase.
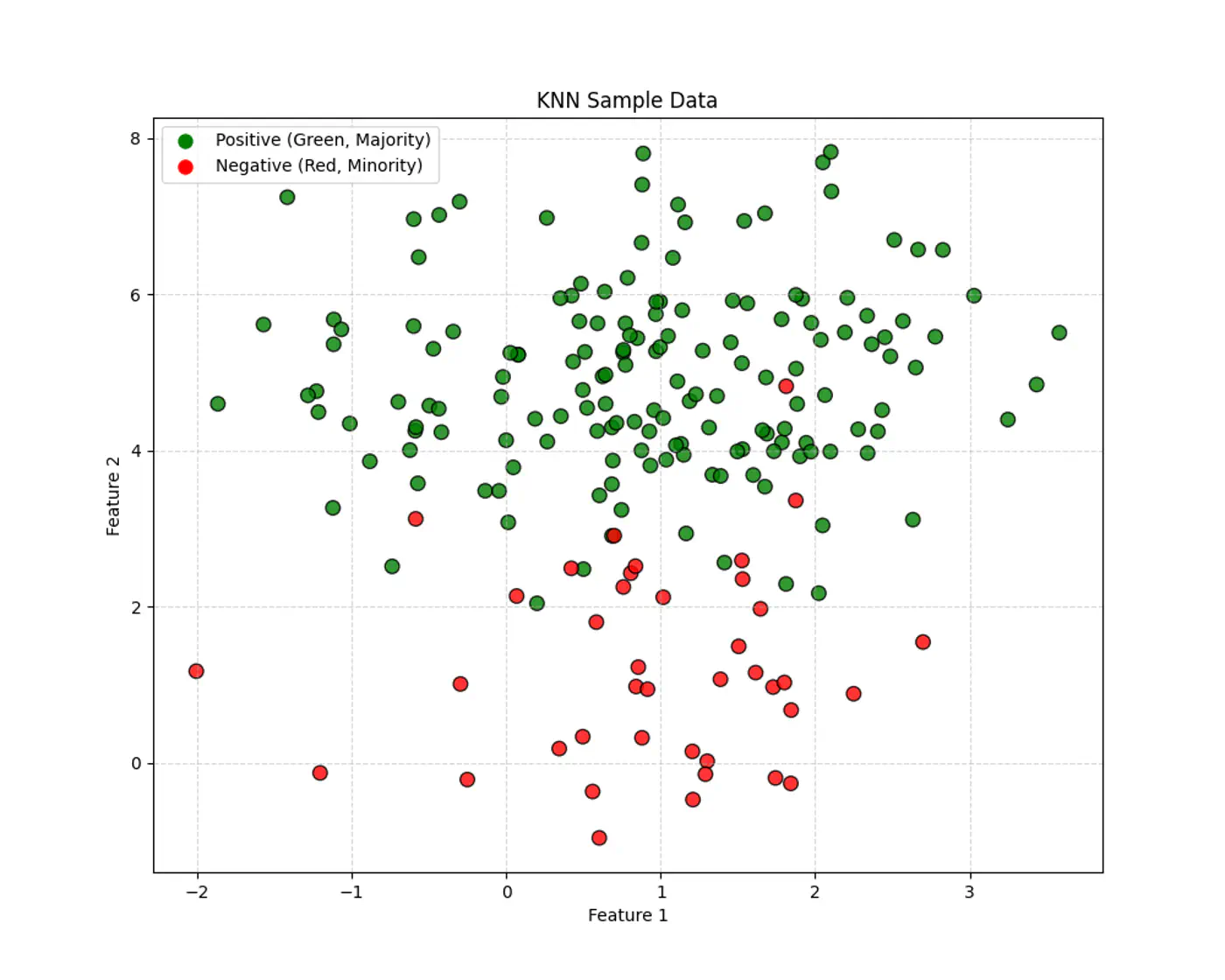

Given a query point \(x_q\) and a dataset, D = {\((x_i,y_i)_{i=1}^n, \quad x_i,y_i \in \mathbb{R}^d\)}, the algorithm finds a set of ‘k’ nearest neighbors \(\mathcal{N}_k(x_q) \subseteq D\).
Inference:
- Choose a value of ‘k’ (hyper-parameter); odd number.
- Calculate distance (Euclidean, Cosine etc.) between and every point in dataset and store in a distance list.
- Sort the distance list in ascending order; choose top ‘k’ data points.
- Make prediction:
- Classification: Take majority vote of ‘k’ nearest neighbors and assign label.
- Regression: Take the mean/median of ‘k’ nearest neighbors.
Note: Store entire dataset.
- Storing Data: Space Complexity: O(nd)
- Inference: Time Complexity ⏰: O(nd + nlogn)
Explanation:
- Distance to all ’n’ points in ‘d’ dimensions: O(nd)
- Sorting all ’n’ data points : O(nlogn)
Note: Brute force 🔨 KNN is unacceptable when ’n’ is very large, say billions.
End of Section
1.2.3.2 - KNN Optimizations
Naive KNN needs some improvements to fix some of its drawbacks.
- Standardization
- Distance-Weighted KNN
- Mahalanobis Distance
⭐️Say one feature is ‘Annual Income’ (0-1M), and another feature is ‘Years of Experience’ (0-40).
👉The Euclidean distance will be almost entirely dominated by income 💵.
💡So, we do standardization of each feature, such that it has a mean, \(\mu\)=0 and variance,\(\sigma\)=1.
\[z=\frac{x-\mu}{\sigma}\]⭐️Vanilla KNN treats the 1st nearest neighbor and the k-th nearest neighbor as equal.
💡A neighbor that is 0.1units away should have more influence than a neighbor that is 10 units away.
👉We assign weight 🏋️♀️ to each neighbor; most common strategy is inverse of squared distance.
\[w_i = \frac{1}{d(x_q, x_i)^2 + \epsilon}\]Improvements:
- Noise/Outlier: Reduces the impact of ‘noise’ or ‘outlier’ (distant neighbors).
- Imbalanced Data: Closer points dominate, mitigating impact of imbalanced data.
- e.g: If you have a query point surrounded by 2 very close ‘Class A’ points and 3 distant ‘Class B’ points, weighted 🏋️♀️ KNN will correctly pick ‘Class A'.
⭐️Euclidean distance makes assumption that all the features are independent and provide unique information.
💡‘Height’ and ‘Weight’ are highly correlated.
👉If we use Euclidean distance, we are effectively ‘double-counting’ the size of the person.
🏇Mahalanobis distance measures distance in terms of standard deviations from the mean, accounting for the covariance between features.
\[d(x, y) = \sqrt{(x - y)^T \Sigma^{-1} (x - y)}\]\(\Sigma\): Covariance matrix of the data
- If \(\Sigma\) is identity matrix, Mahalanobis distance reduces to Euclidean distance.
- If \(\Sigma\) is a diagonal matrix, Mahalanobis distance reduces to Normalized Euclidean distance.
🦀Naive KNN shifts all computation 💻 to inference time ⏰, and it is very slow.
- To find the neighbor for one query, we must touch every single bit of the ‘nxd’ matrix.
- If n=10^9,a single query would take seconds, but we need milliseconds.
- Distance Weighted KNN
- K-D Trees (d<20): Recursively partitions space into axis-aligned hyper-rectangles. O(log N ) search.
- Ball Trees : High dimensional data; Haversine distance for geospatial 🌎 data.
- Approximate Nearest Neighbors (ANN)
- Locality Sensitive Hashing (LSH): Uses ‘bucketizing’ 🗑️ hashes. Points that are close have a high probability of having the same hash.
- Hierarchical Navigable Small World (HNSW); Graph of vectors; Search is a ‘greedy walk’ across levels.
- Product Quantization (Reduce memory 🧠 footprint 👣 of high dimensional vectors)
- ScaNN (Google)
- FAISS (Meta)
- Dimensionality Reduction (Mitigate ‘Curse of Dimensionality’)
- PCA
End of Section
1.2.3.3 - Curse Of Dimensionality
While Euclidean distance(L norm) is the most frequently discussed, ‘Curse of Dimensionality’ impacts all Minkowski norms (\(L_p\))
\[L_p = (\sum |x_i|^p)^{\frac{1}{p}} \]Note: ‘Curse of Dimensionality’ is largely a function of the exponent (p) in the distance calculation.
Coined 🪙 by mathematician John Bellman in the 1960s while studying dynamic programming.
High dimensional data created following challenges:
- Distance Concentration
- Data Sparsity
- Exponential Sample Requirement
💡Consider a hypercube in d-dimensions of side length = 1; Volume = \(1^d\) = 1
🧊 A smaller inner cube with side length = 1 - \(\epsilon\) ; Volume = \((1 -\epsilon)^d\)
🧐 This implies that almost all the volume of the high-dimensional cube lies near the ‘crust’.
👉e.g: if \(\epsilon\)= 0.01, d = 500; Volume of inner cube = \((1 -0.01)^{500}\) = \(0.99^{500}\) = 0.006 = 0.6%
🤔Consequently, all points become nearly equidistant, and the concept of ‘nearest’ or ‘neighborhood’ loses
its meaning.
⭐️The volume of the feature space increases exponentially with each added dimension.
👉To maintain the same data density found in a 1D space with 10 points, we would need \(10^{10}\)(10 billion) points in 10D space.
💡Because real-world datasets are rarely this large, the data becomes “sparse,” making it difficult to find truly similar neighbors.
⭐️To maintain a reliable result, the amount of training data needed must grow exponentially with the number of dimensions.
👉Without this growth, the model is highly prone to overfitting, where it learns from noise in the ‘sparse’ data rather than actual underlying patterns.
Note: For modern embeddings (often 768 or 1536 dimensions), it is mathematically impossible to collect enough data to ‘fill’ the space.
- Cosine Similarity
- Normalization
Cosine similarity measures the cosine of the angle between 2 vectors.
\[\text{cos}(\theta) = \frac{A \cdot B}{\|A\|\|B\|} = \frac{\sum_{i=1}^{D} A_i B_i}{\sqrt{\sum_{i=1}^{D} A_i^2} \sqrt{\sum_{i=1}^{D} B_i^2}}\]Note: Cosine similarity mitigates the ‘curse of dimensionality" problem.
⭐️Normalize the vector, i.e, make its length =1, a unit vector.
💡By normalizing, we project all points onto the surface of a unit hypersphere.
- We are no longer searching in the ‘empty’ high-dimensional volume of a hypercube.
- Now, we are searching on a constrained manifold (the shell).
Note: By normalizing, we move the data from the volume of the D-dimensional space onto the surface of a (D-1)-dimensional hypersphere.
Euclidean Distance Squared of Normalized Vectors:
\[ \begin{align*} \|A - B\|^2 &= (A - B) \cdot (A - B) \\ &= \|A\|^2 + \|B\|^2 - 2(A \cdot B)\\ \because \|A\| &= \|B\| = 1 \\ \|A - B\|^2 &= 1 + 1 - 2\cos(\theta) \\ \therefore \|A - B\|^2 &= 2(1 - \cos(\theta))\\ \end{align*} \]Note: This formula proves that maximizing ‘Cosine similarity’ is identical to minimizing ‘Euclidean distance’ on the hypersphere.
End of Section
1.2.3.4 - Bias Variance Tradeoff
Let’s use this dataset to understand the impact of number of neighbours ‘k’.
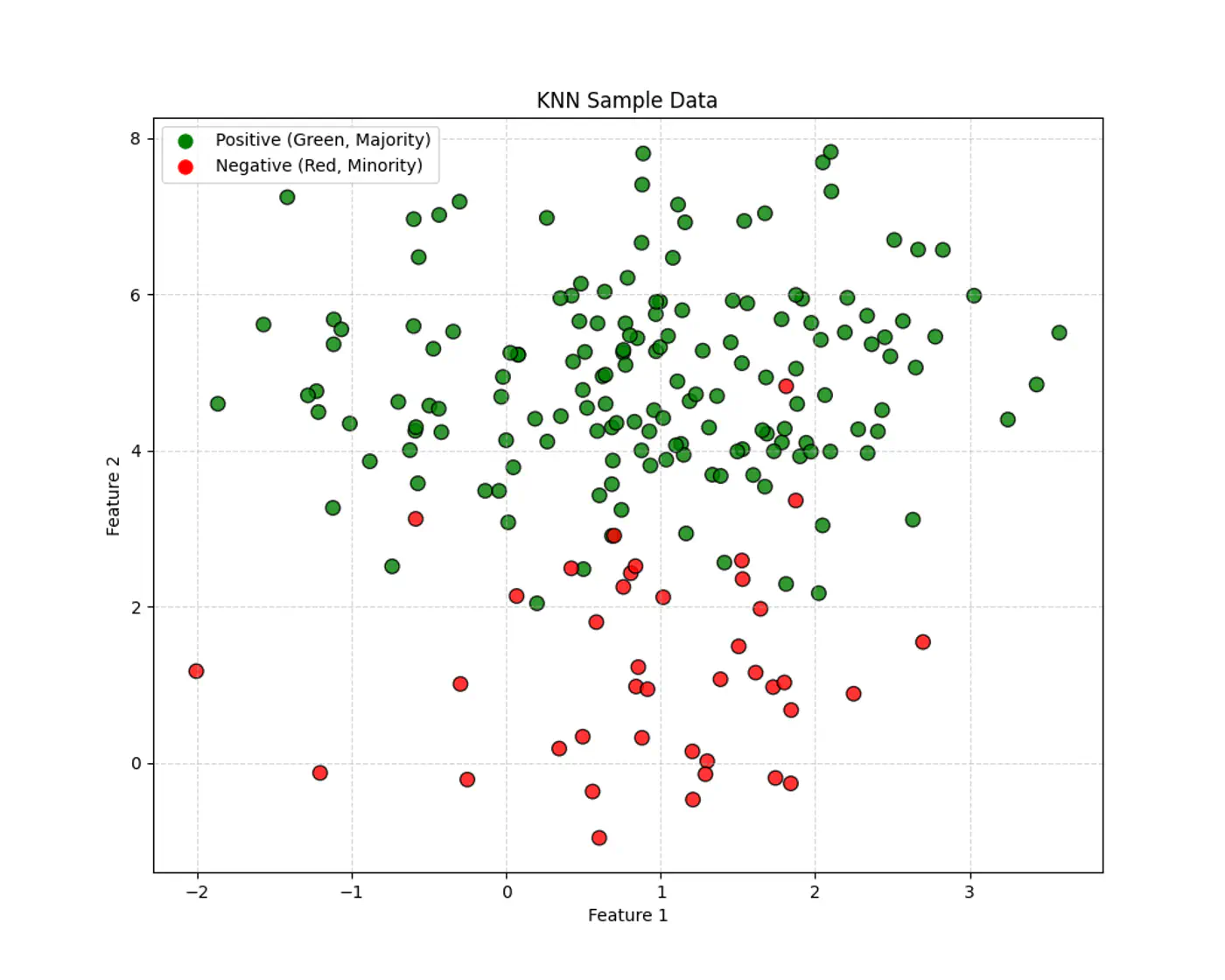
👉If ‘k’ is very large, say, k=n,
- model simply predicts the majority class of the entire dataset for every query point , i.e, under-fitting.
👉If ‘k’ is very small, say, k=1,
- model is highly sensitive to noise or outliers, as it looks at only 1 nearest neighbor, i.e, over-fitting.
Let’s plot Error vs ‘K’ neighbors:
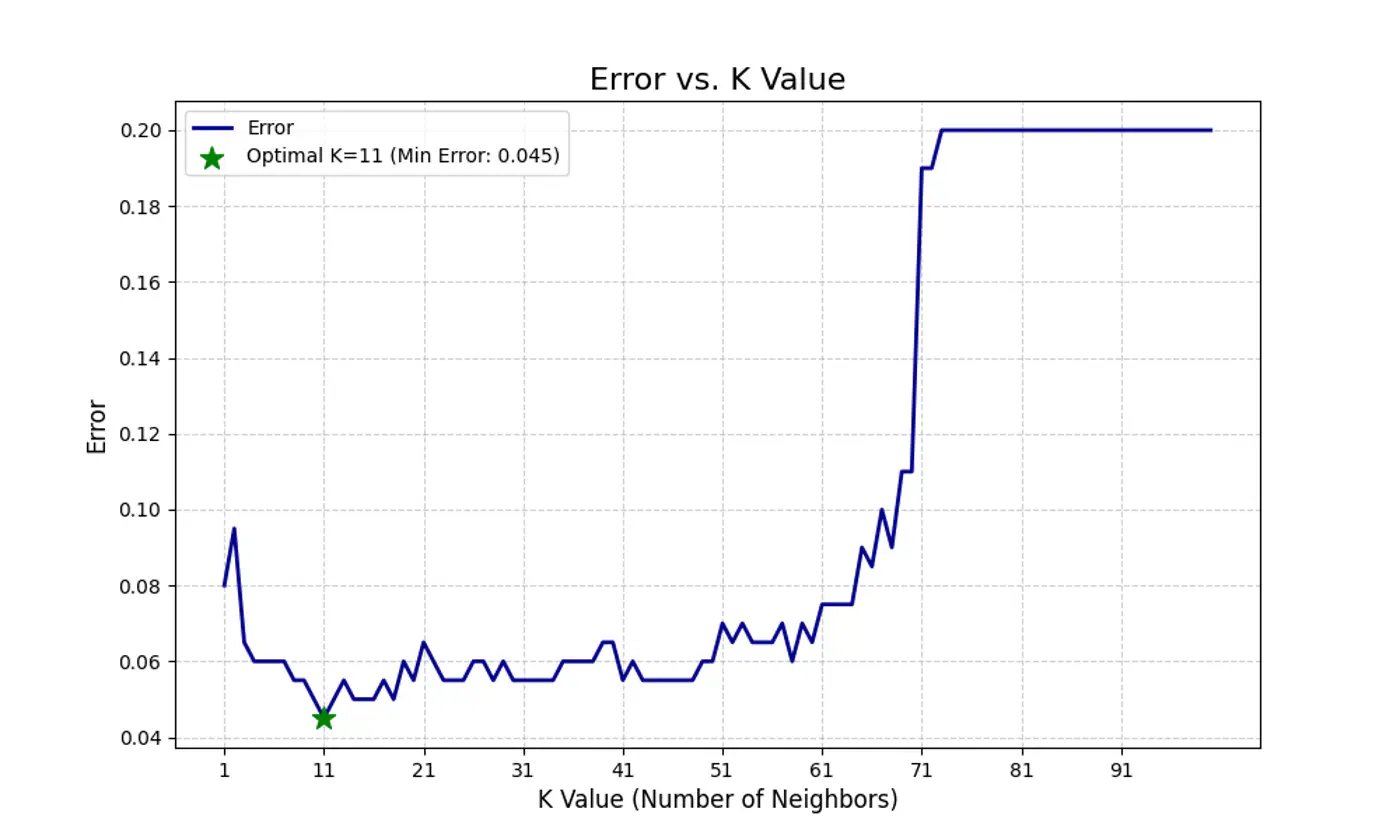
Figure 1: k=1, Over-fitting
Figure 2: k=n, Under-fitting
Figure 3: k=11, Lowest Error (Optimum)
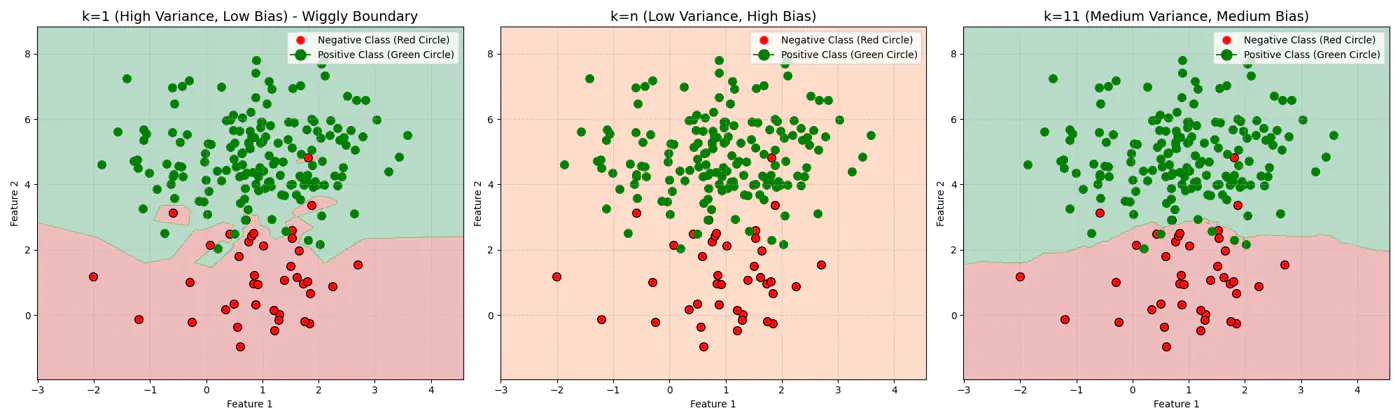
End of Section
1.2.4.1 - Decision Trees Introduction
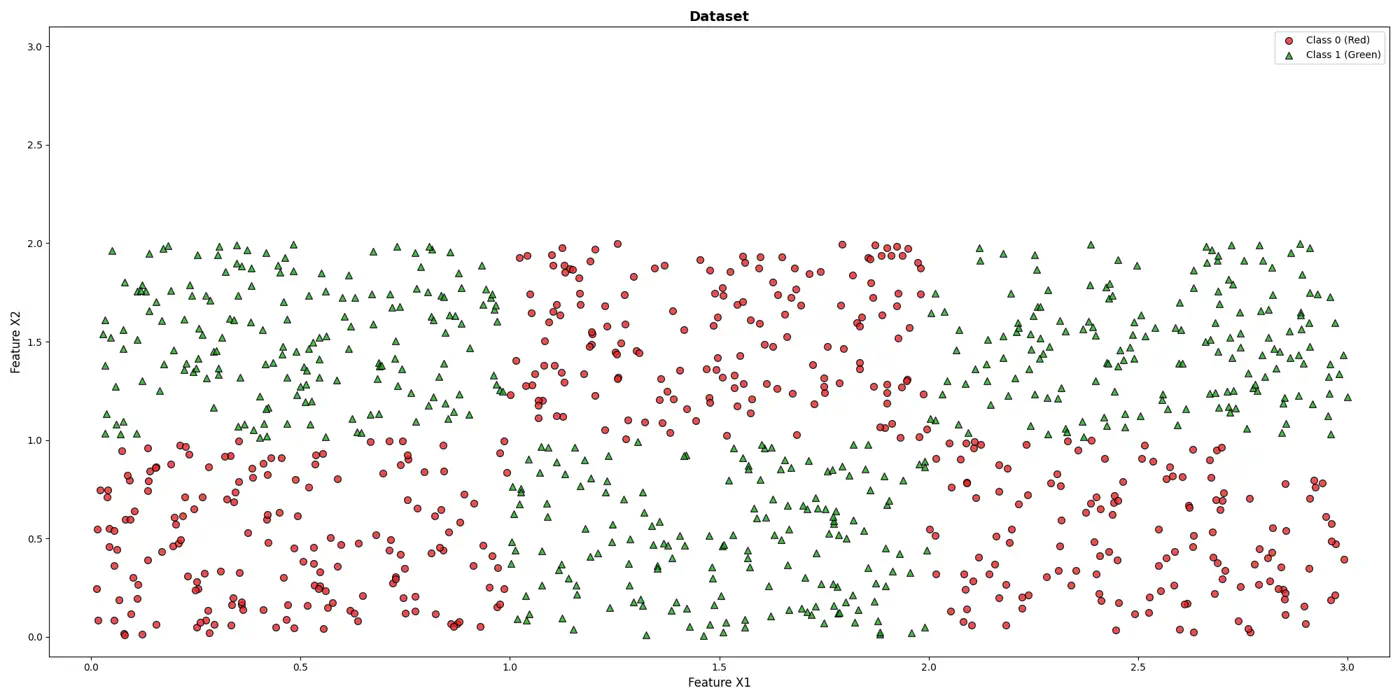
💡It can be written as nested 🕸️ if else statements.
e.g: To classify the left bottom corner red points we can write:
👉if (FeatureX1 <1 & FeatureX2 <1)
⭐️Extending the logic for all, we have an if else ladder like below:
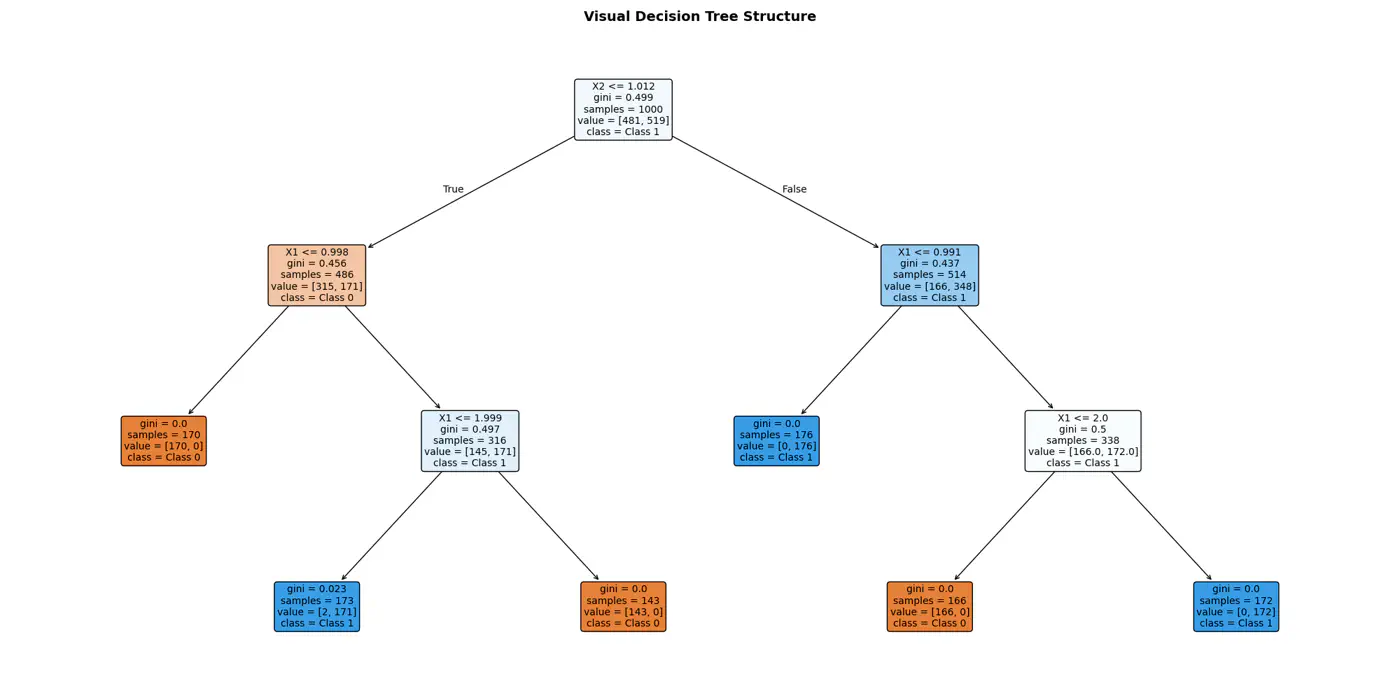
👉Final decision boundaries will be something like below:
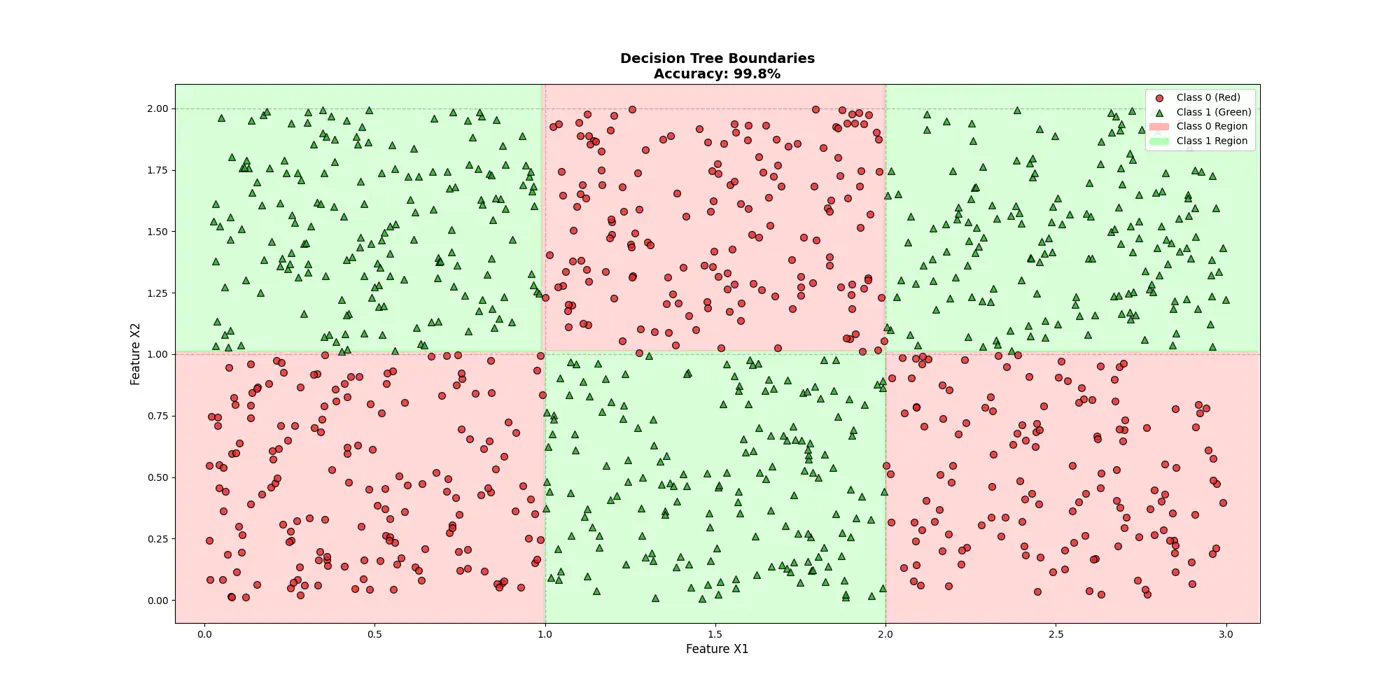
- Non-parametric model.
- Recursively partitions the feature space.
- Top-down, greedy approach to iteratively select feature splits.
- Maximize purity of a node, based on metrics, such as, Information Gain 💵 or Gini 🧞♂️Impurity.
Note: We can extract the if/else logic of the decision tree and write in C++/Java for better performance.
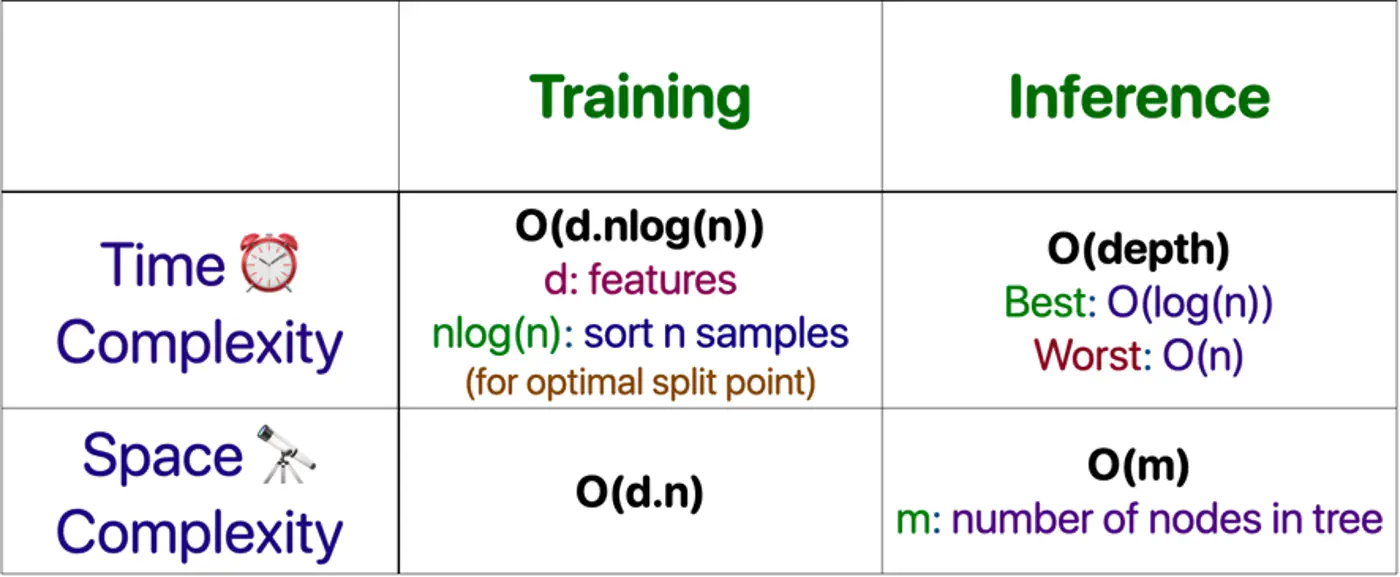
⭐️Building an optimal decision tree 🌲 is a NP-Hard problem.
👉(Time Complexity: Exponential; combinatorial search space)
- Pros
- No standardization of data needed.
- Highly interpretable.
- Good runtime performance.
- Works for both classification & regression.
- Cons
- Number of dimensions should not be too large. (Curse of dimensionality)
- Overfitting.
- As base learners in ensembles, such as, bagging(RF), boosting(GBDT), stacking, cascading, etc.
- As a baseline, interpretable, model or for quick feature selection.
- Runtime performance is important.
End of Section
1.2.4.2 - Purity Metrics
Decision trees recursively partition the data based on feature values.
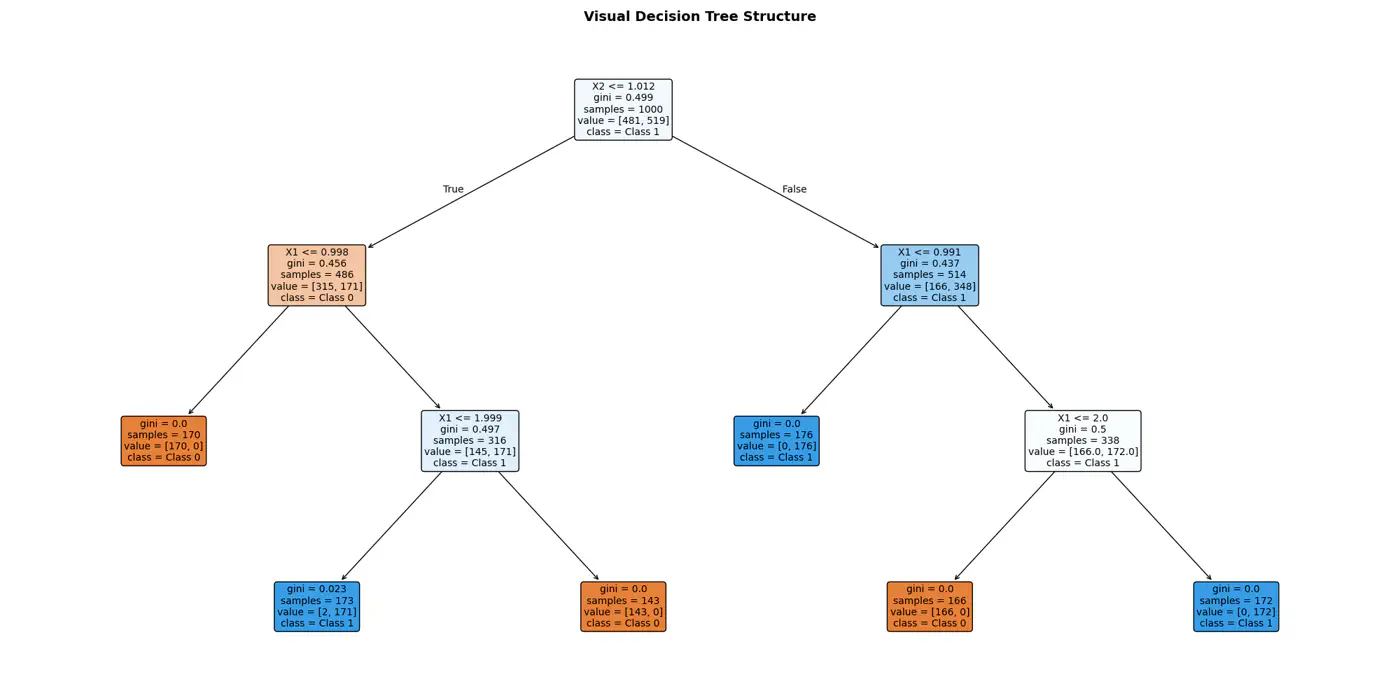
Pure Leaf 🍃 Node: Terminal node where every single data point belongs to the same class.
💡Zero Uncertainty.
The goal of a decision tree algorithm is to find the split that maximizes information gain, meaning it removes the most uncertainty from the data.
So, what is information gain ?
How do we reduce uncertainty ?
Let’s understand few terms first, before we understand information gain.
Measure ⏱ of uncertainty, randomness, or impurity in a data.
\[H(S)=-\sum _{i=1}^{n}p_{i}\log(p_{i})\]Binary Entropy:
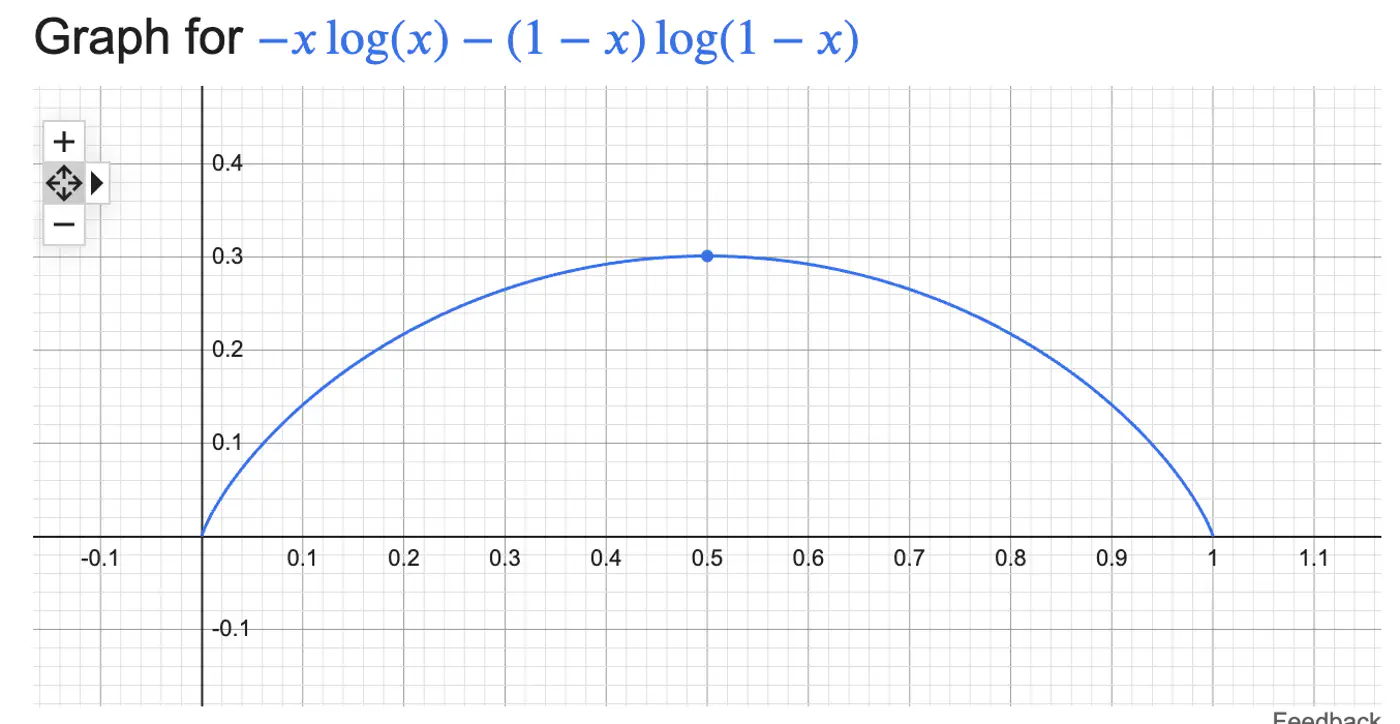
💡Entropy can also be viewed as the ‘average surprise'.
A highly certain event provides little information when it occurs (low surprise).
An unlikely event provides a lot of information (high surprise).

⭐️ Measures the reduction in entropy (uncertainty) achieved by splitting a dataset based on a specific attribute.
\[IG=Entropy(Parent)-\left[\frac{N_{left}}{N_{parent}}Entropy(Child_{left})+\frac{N_{right}}{N_{parent}}Entropy(Child_{right})\right] \]Note: The goal of a decision tree algorithm is to find the split that maximizes information gain, meaning it removes the most uncertainty from the data.
⭐️ Measures the probability of an element being incorrectly classified if it were randomly labeled according to the distribution of labels in a node.
\[Gini(S)=1-\sum_{i=1}^{n}(p_{i})^{2}\]- Range: 0 (Pure) - 0.5 (Maximum impurity)
Note: Gini is used in libraries like Scikit-Learn (as the default), because it avoids the computationally expensive 💰 log function.
Gini Impurity is a first-order approximation of Entropy.
For most of the real-world cases, choosing one over the other results in the exact same tree structure or negligible differences in accuracy.
When we plot the two functions, they follow nearly identical shapes.
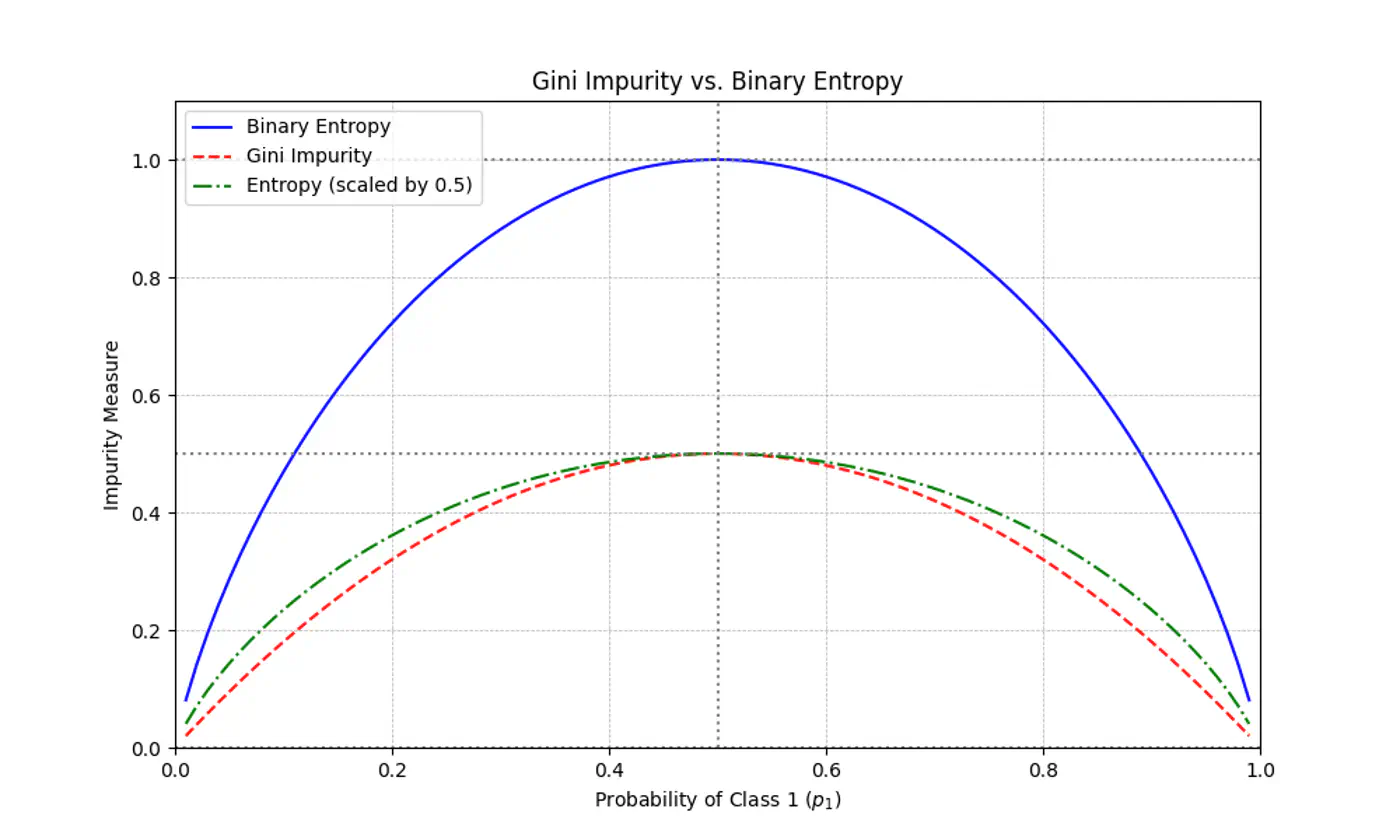
End of Section
1.2.4.3 - Decision Trees For Regression
Decision Trees can also be used for Regression tasks but using a different metrics.
⭐️Metric:
- Mean Squared Error (MSE)
- Mean Absolute Error (MAE)
👉Say we have a following dataset, that we need to fit using decision trees:
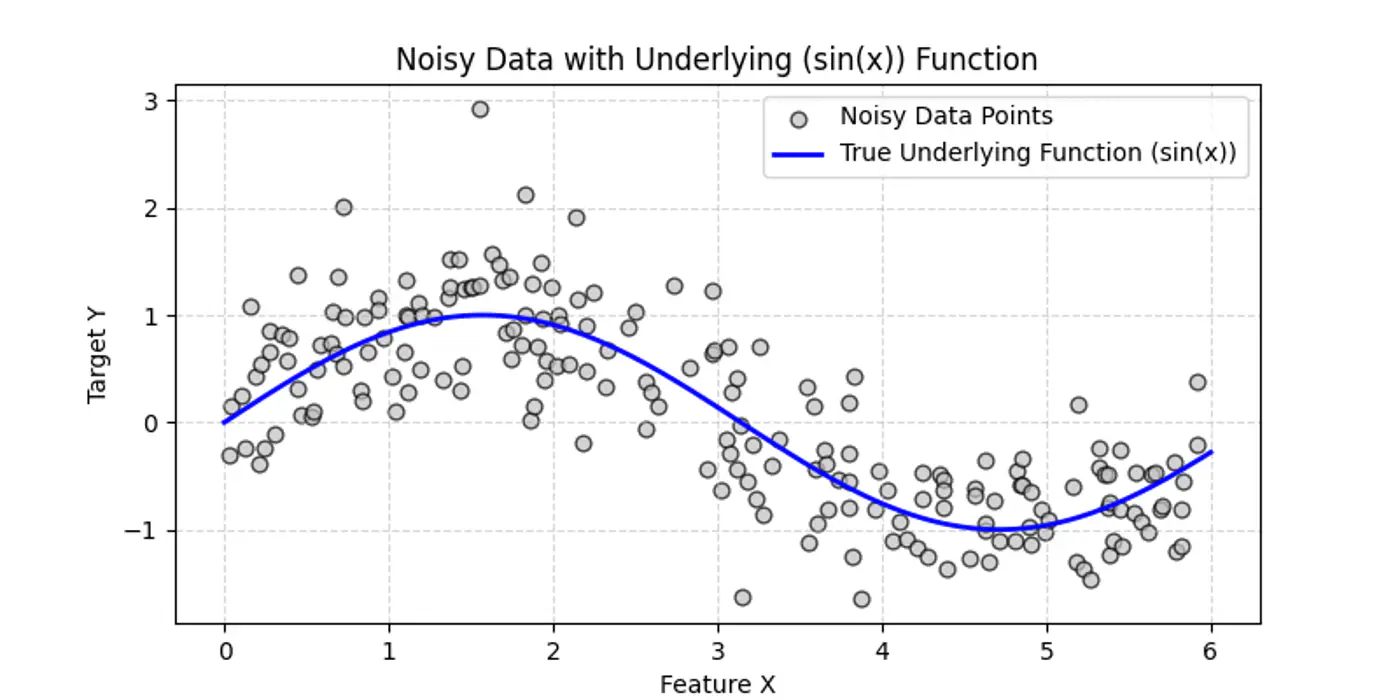
👉Decision trees try to find the decision splits, building step functions that approximate the actual curve, as shown below:
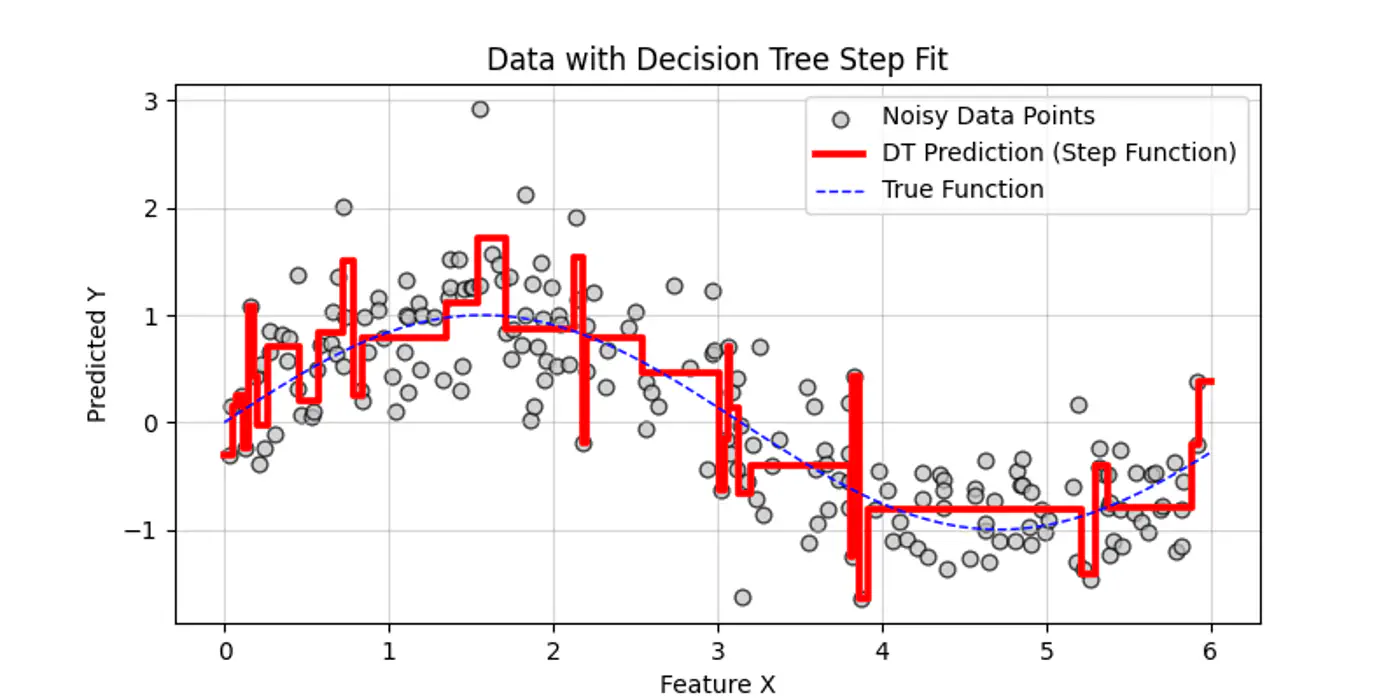
👉Internally the decision tree (if else ladder) looks like below:
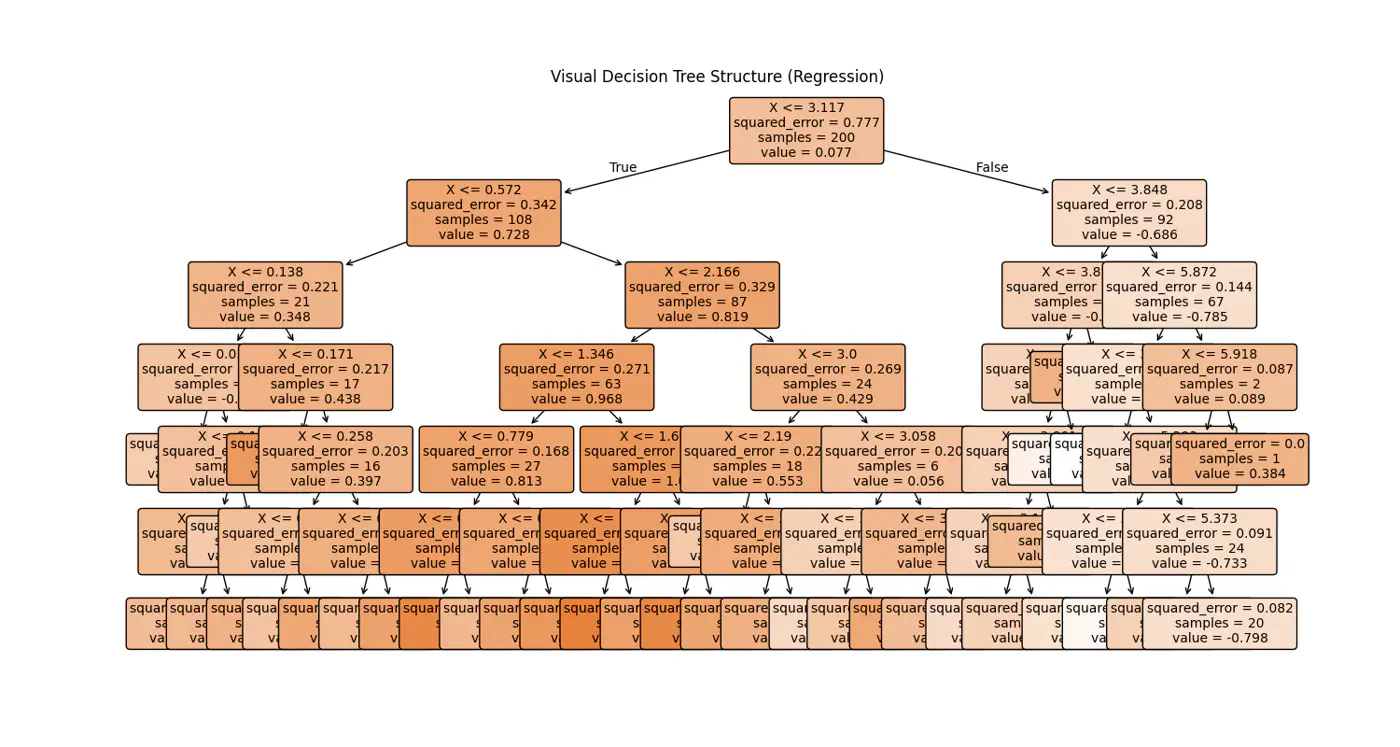
Decision trees cannot predict values outside the range of the training data, i.e, extrapolation.
Let’s understand the interpolation and extrapolation cases one by one.
⭐️Predicting values within the range of features and targets observed during training 🏃♂️.
- Trees capture discontinuities perfectly, because they are piece-wise constant.
- They do not try to force a smooth line where a ‘jump’ exists in reality.
e.g: Predicting a house 🏡 price 💰 for a 3-BHK home when you have seen 2-BHK and 4-BHK homes in that same neighborhood.
⭐️Predicting values outside the range of training 🏃♂️data.
Problem:
Because a tree outputs the mean of training 🏃♂️ samples in a leaf, it cannot predict a value higher than the
highest ‘y’ it saw during training 🏃♂️.
- Flat-Line: Once a feature ‘X’ goes beyond the training boundaries, the tree falls into the same ‘last’ leaf forever.
e.g: Predicting the price 💰 of a house 🏡 in 2026 based on data from 2010 to 2025.
End of Section
1.2.4.4 - Regularization
- Pre-Pruning ✂️ &
- Post-Pruning ✂️
⭐️ ‘Early stopping’ heuristics (hyper-parameters).
- max_depth: Limits how many levels of ‘if else’ the tree can have; most common.
- min_samples_split: A node will only split, if it has at least ‘N’ samples; smooths the model (especially in regression), by ensuring predictions are based on an average of multiple points.
- max_leaf_nodes: Limiting the number of leaves reduces the overall complexity of the tree, making it simpler and less likely to memorize the training data’s noise.
- min_impurity_decrease: A split is only made if it reduces the impurity (Gini/MSE) by at least a certain threshold.
Below is an example for one of the hyper-parameter’s max_depth tuning.
As we can see below the cross-validation error decreases till depth=6 and after that reduction in error is not so significant.
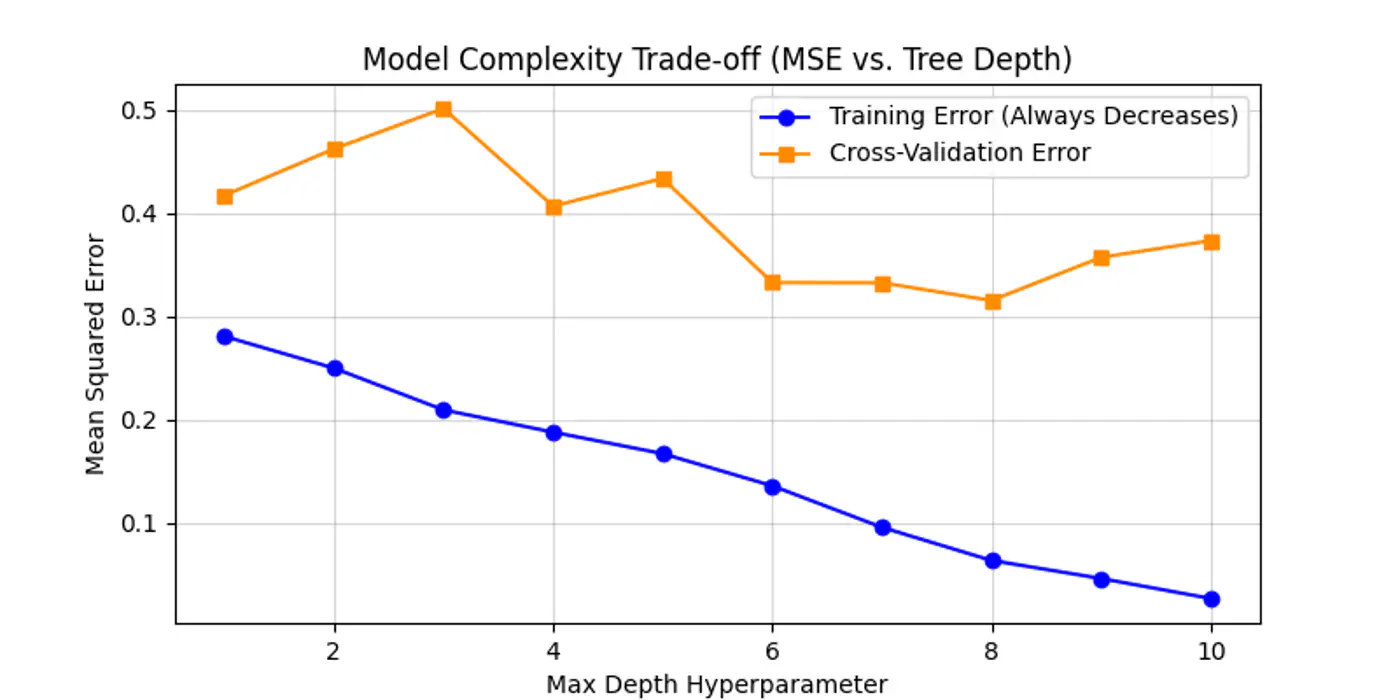
Let the tree🌲 grow to its full depth (overfit) and then ‘collapse’ nodes that provide little predictive value.
Most common algorithm:
- Minimal Cost Complexity Pruning
💡Define a cost-complexity 💰 measure that penalizes the tree 🌲 for having too many leaves 🍃.
\[R_\alpha(T) = R(T) + \alpha |T|\]- R(T): total misclassification rate (or MSE) of the tree
- |T|: number of terminal nodes (leaves)
- \(\alpha\): complexity parameter (the ‘tax’ 💰 on complexity)
Logic:
- If \(\alpha\)=0, the tree is the original overfit tree.
- As \(\alpha\) increases 📈, the penalty for having many leaves grows 📈.
- To minimize the total cost 💰, the model is forced to prune branches that do not significantly reduce R(T).
- Use cross-validation to find the ‘sweet spot’ \(\alpha\) that minimizes validation error.
End of Section
1.2.4.5 - Bagging
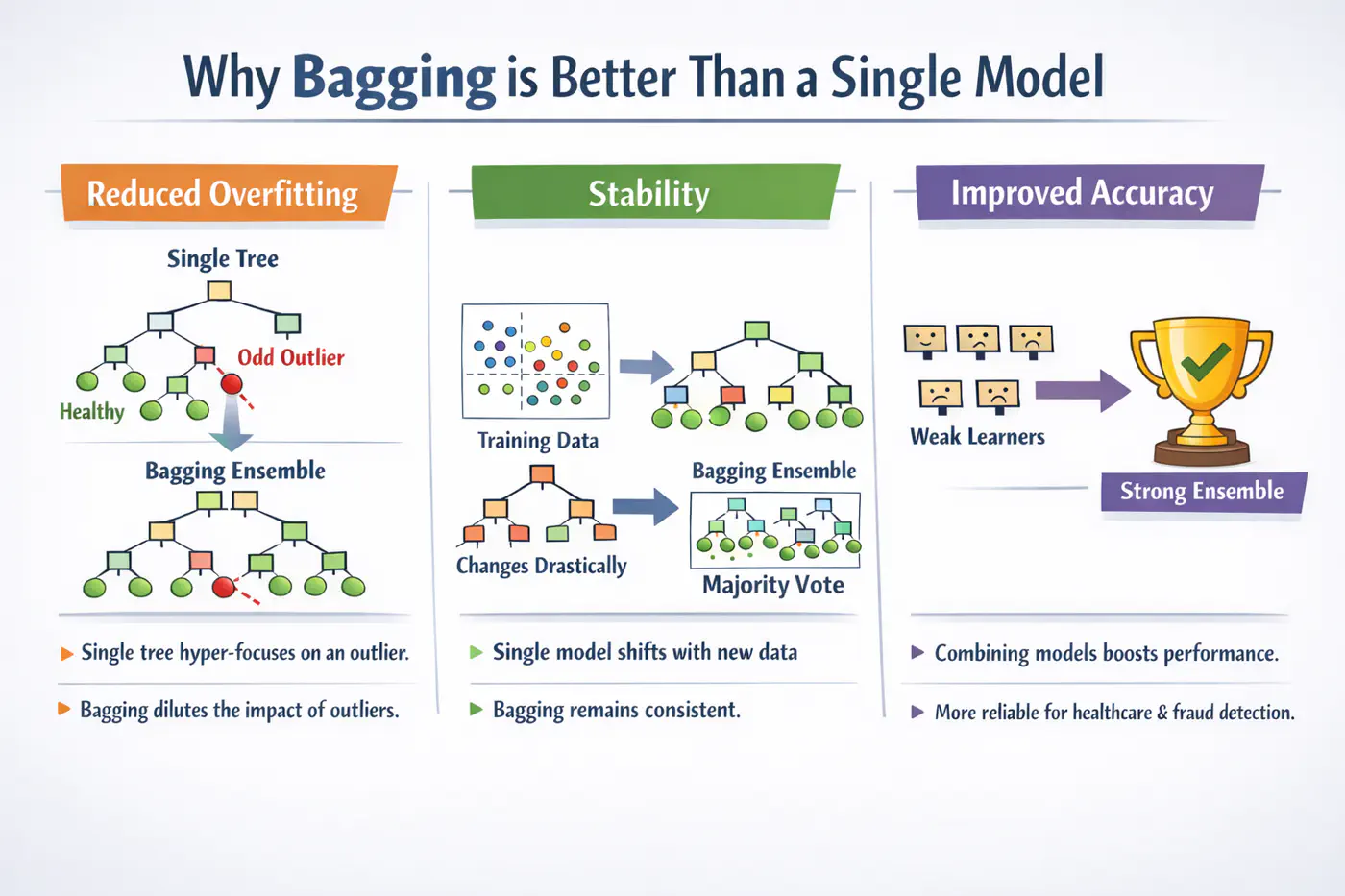
A single decision tree is highly sensitive to the specific training dataset.
Small changes, such as, a few different rows or the presence of an outlier, can lead to a completely different tree structure.Unpruned decision trees often grow until they perfectly classify the training set, essentially ‘memorizing’ noise and outliers, i.e, high variance, rather than finding general patterns.
Bagging = ‘Bootstrapped Aggregation’
Bagging 🎒is a parallel ensemble technique that reduces variance (without significantly increasing the bias) by training multiple versions of the same model on different random subsets of data and then combining their results.
Note: Bagging uses deep trees (overfit) and combines them to reduce variance.
Bootstrapping = ‘Without external help’
Given a training 🏃♂️set D of size ’n’, we create B new training sets D by sampling ’n’ observations from D ‘with replacement'.
💡Since, we are sampling ‘with replacement’, so, some data points may be picked multiple times, while others may not be picked at all.
- The probability that a specific observation is not selected in a bootstrap sample of size ’n’ is: \[\lim_{n \to \infty} \left(1 - \frac{1}{n}\right)^n = \frac{1}{e} \approx 0.368\]
🧐This means each tree is trained on roughly 63.2% of the unique data, while the remaining 36.8% (the Out-of-Bag or OOB set) can be used for cross validation.
⭐️Say we train ‘B’ models (base-learners), each with variance \(\sigma^2\) .
👉Average variance of ‘B’ models (trees) if all are independent:
\[Var(X)=\frac{\sigma^{2}}{B}\]👉Since, bootstrap samples are derived from the same dataset, the trees are correlated with some correlation coefficient ‘\(\rho\)'.
So, the true variance of bagged ensemble is:
\[Var(f_{bag}) = \rho \sigma^2 + \frac{1-\rho}{B} \sigma^2\]- \(\rho\)= 0; independent models, most reduction in variance.
- \(\rho\)= 1; fully correlated models, no improvement in variance.
- 0<\(\rho\)<1; As correlation decreases, variance reduces .
End of Section
1.2.4.6 - Random Forest
💡If one feature is extremely predictive (e.g., ‘Area’ for house prices), almost every bootstrap tree will split on that feature at the root.
👉This makes the trees(models) very similar, leading to a high correlation ‘\(\rho\)’.
\[Var(f_{bagging})=ρ\sigma^{2}+\frac{1-ρ}{B}\sigma^{2}\]💡Choose a random subset of ‘m’ features from the total ‘d’ features, reducing the correlation ‘\(\rho\)’ between trees.
👉By forcing trees to split on ‘sub-optimal’ features, we intentionally increase the variance of individual trees; also the bias is slightly increased (simpler trees).
Standard heuristics:
- Classification: \(m = \sqrt{d}\)
- Regression: \(m = \frac{d}{3}\)
💡Because ‘\(\rho\)’ is the dominant factor in the variance of the ensemble when B is large, the overall ensemble variance Var(\(f_{rf}\)) drops significantly lower than standard Bagging.
\[Var(f_{rf})=ρ\sigma^{2}+\frac{1-ρ}{B}\sigma^{2}\]💡A Random Forest will never overfit by adding more trees (B).
It only converges to the limit: ‘\(\rho\sigma^2\)’.
Overfitting is controlled by:
- depth of the individual trees.
- size of the feature subset ‘m'.
- High Dimensionality: 100s or 1000s of features; RF’s feature sampling prevents a few features from masking others.
- Tabular Data (with Complex Interactions): Captures non-linear relationships without needing manual feature engineering.
- Noisy Datasets: The averaging process makes RF robust to outliers (especially if using min_samples_leaf).
- Automatic Validation: Need a quick estimate of generalization error without doing 10-fold CV (via OOB Error).
End of Section
1.2.4.7 - Extra Trees
💡In a standard Decision Tree or Random Forest, the algorithm searches for the optimal split point (the threshold ’s’) that maximizes Information Gain or minimizes MSE.
👉This search is:
- computationally expensive (sort + mid-point) and
- tends to follow the noise in the training 🏃♂️data.
Adding randomness (right kind) in ensemble averaging reduces correlation/variance.
\[Var(f_{bag})=ρ\sigma^{2}+\frac{1-ρ}{B}\sigma^{2}\]- Random Thresholds: Instead of searching for the best split point (computationally expensive 💰) for a feature, it picks a threshold at random from a uniform distribution between the feature’s local minimum and maximum.
- Entire Dataset: Uses entire training dataset (default) for every tree; no bootstrapping.
- Random Feature Subsets: Random subset of m<n features is used in each decision tree.
Picking thresholds randomly has two effects:
- Structural correlation between trees becomes extremely low.
- Individual trees are ‘weaker’ and have higher bias than a standard optimized tree.
👉The massive drop in ‘\(\rho\)’ often outweighs the slight increase in bias, leading to an overall ensemble that is smoother and more robust to noise than a standard Random Forest.
Note: Extra Trees are almost always grown to full depth, as they may need extra splits to find the same decision boundary.
- Performance: Significantly faster to train, as it does not sort data to find optimal split.
Note: If we are working with billions of rows or thousands of features, ET can be 3x to 5x faster than a Random Forest(RF). - Robustness to Noise: By picking thresholds randomly, tends to ‘handle’ the noise more effectively than RF.
- Feature Importance: Because ET is so randomized, it often provides more ‘stable’ feature importance scores.
Note: It is less likely to favor a high-cardinality feature (e.g. zip-code) just because it has more potential split points.
End of Section
1.2.4.8 - Boosting
⭐️In Bagging 🎒we trained multiple strong(over-fit, high variance) models (in parallel) and then averaged them out to reduce variance.
💡Similarly, we can train many weak(under-fit, high bias) models sequentially, such that, each new model corrects the errors of the previous ones to reduce bias.
⚔️ An ensemble learning approach where multiple ‘weak learners’ (typically simple models like shallow decision trees 🌲 or ‘stumps’) are sequentially combined to create a single strong predictive model.
⭐️The core principle is that each subsequent model focuses 🎧 on correcting the errors made by its predecessors.
- AdaBoost(Adaptive Boosting)
- Gradient Boosting Machine (GBM)
- XGBoost
- LightGBM (Microsoft)
- CatBoost (Yandex)
End of Section
1.2.4.9 - AdaBoost
💡Works by increasing 📈 the weight 🏋️♀️ of misclassified data points after each iteration, forcing the next weak learner to ‘pay more attention’🚨 to the difficult cases.
⭐️ Commonly used for classification.
👉Weak learners are typically ‘Decision Stumps’, i.e, decision trees🌲with a depth of only one (1 split, 2 leaves 🍃).
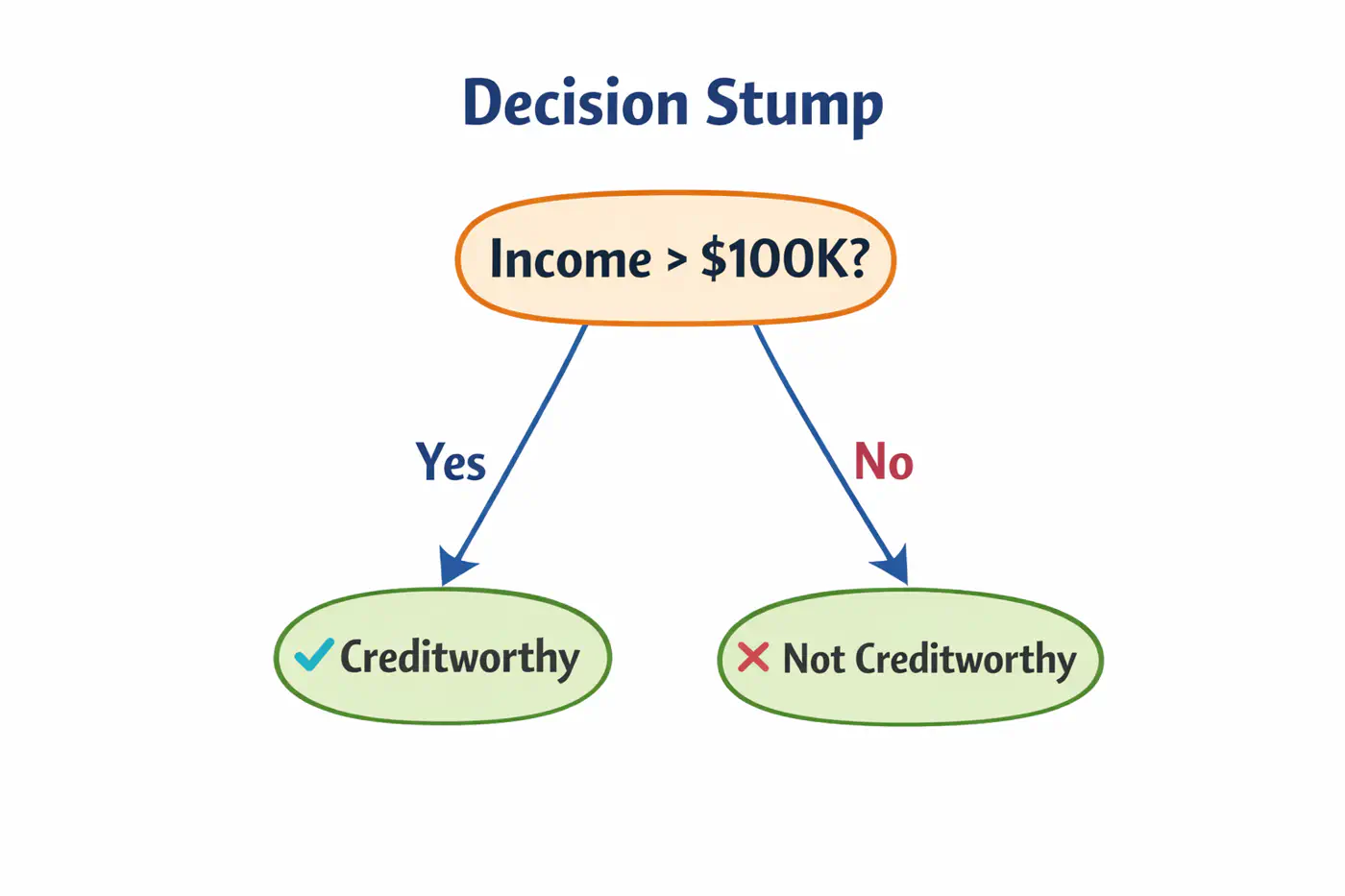
- Assign an equal weight 🏋️♀️to every data point; \(w_i = 1/n\), where ’n’=number of samples.
- Build a decision stump that minimizes the weighted classification error.
- Calculate total error; \(E_m = \Sigma w_i\).
- Determine ‘amount of say’, i.e, the weight 🏋️♀️ of each stump in final decision.
\[\alpha_m = \frac{1}{2}ln\left( \frac{1-E_m}{E_m} \right)\]
- Low error results in a high positive \(\alpha\) (high influence).
- 50% error (random guessing) results in an \(\alpha = 0\) (no influence).
- Update sample weights 🏋️♀️.
- Misclassified samples: Weight 🏋️♀️ increases by \(e^{\alpha_m}\).
- Correctly classified samples: Weight 🏋️♀️ decreases by \(e^{-\alpha_m}\).
- Normalization: All new weights 🏋️♀️ are divided by their total sum so they add up back to 1.
- Iterate for a specified number of estimators (n_estimators).
👉 To classify a new data point, every stump makes a prediction (+1 or -1).
These are multiplied by their respective ‘amount of say’ \(\alpha_m\) and summed.
\[H(x)=sign\sum_{m=1}^{M}\alpha_{m}⋅h_{m}(x)\]👉 If the total weighted 🏋️♀️ sum is positive, the final class is +1; otherwise -1.
Note: Sensitive to outliers; Because AdaBoost aggressively increases weights 🏋️♀️ on misclassified points, it may ‘over-focus’ on noisy outliers, hurting performance.
End of Section
1.2.4.10 - Gradient Boosting Machine
GBM treats the final model \(F_m(x)\) as weighted 🏋️♀️ sum of ‘m’ weak learners:
\[ F_{M}(x)=\underbrace{F_{0}(x)}_{\text{Initial\ Guess}}+\nu \sum _{m=1}^{M}\underbrace{\left(\sum _{j=1}^{J_{m}}\gamma _{jm}\mathbb{I}(x\in R_{jm})\right)}_{\text{Weak\ Learner\ }h_{m}(x)}\]- \(F_0(x)\): The initial base model (usually a constant).
- M: The total number of boosting iterations (number of trees).
- \(\gamma_{jm}\)(Leaf Weight): The optimized value for leaf in tree .
- \(\nu\)(Nu): The Learning Rate or Shrinkage; prevent overfitting.
- \(\mathbb{I}(x\in R_{jm})\): ‘Indicator Function’; It is 1 if data point falls into leaf of the tree, and 0 otherwise.
- \(R_{jm}\)(Regions): Region of \(j_{th}\) leaf in \(m_{th}\)tree.
📍In Gradient Descent, we update parameters ‘\(\Theta\)';
📍In GBM, we update the predictions F(x) themselves.
🦕We move the predictions in the direction of the negative gradient of the loss function L(y, F(x)).
🎯We want to minimize loss:
\[\mathcal{L}(F) = \sum_{i=1}^n L(y_i, F(x_i))\]✅ In parameter optimization we update weights 🏋️♀️:
\[w_{t+1} = w_t - \eta \cdot \nabla_{w}\mathcal{L}(w_t)\]✅ In gradient boosting, we update the prediction function:
\[F_m(x) = F_{m-1}(x) -\eta \cdot \nabla_F \mathcal{L}(F_{m-1}(x))\]➡️ The gradient is calculated w.r.t. predictions, not weights.
In GBM we can use any loss function as long as it is differentiable, such as, MSE, log loss, etc.
Loss(MSE) = \((y_i - F_m(x_i))^2\)
\[\frac{\partial L}{F_m(x_i)} = -2 (y-F_m(x_i))\]\[\implies \frac{\partial L}{F_m(x_i)} \propto - (y-F_m(x_i))\]👉Pseudo Residual (Error) = - Gradient
💡To minimize loss, take derivative of loss function w.r.t ‘\(\gamma\)’ and equate to 0:
\[F_0(x) = \arg\min_{\gamma} \sum_{i=1}^n L(y_i, \gamma)\]MSE Loss = \(\mathcal{L}(y_i, \gamma) = \sum_{i=1}^n(y_i -\gamma)^2\)
\[ \begin{aligned} &\frac{\partial \mathcal{L}(y_i, \gamma)}{\partial \gamma} = -2 \cdot \sum_{i=1}^n(y_i -\gamma) = 0 \\ &\implies \sum_{i=1}^n (y_i -\gamma) = 0 \\ &\implies \sum_{i=1}^n y_i = n.\gamma \\ &\therefore \gamma = \frac{1}{n} \sum_{i=1}^n y_i \end{aligned} \]💡To minimize cost, take derivative of cost function w.r.t ‘\(\gamma\)’ and equate to 0:
Cost Function = \(J(\gamma )\)
\[J(\gamma )=\sum _{x_{i}\in R_{jm}}\frac{1}{2}(y_{i}-(F_{m-1}(x_{i})+\gamma ))^{2}\]We know that:
\[ r_{i}=y_{i}-F_{m-1}(x_{i})\]\[\implies J(\gamma )=\sum _{x_{i}\in R_{jm}}\frac{1}{2}(r_{i}-\gamma )^{2}\]\[\frac{d}{d\gamma }\sum _{x_{i}\in R_{jm}}\frac{1}{2}(r_{i}-\gamma )^{2}=\sum _{x_{i}\in R_{jm}}-(r_{i}-\gamma )=0\]\[\implies \sum _{x_{i}\in R_{jm}}\gamma -\sum _{x_{i}\in R_{jm}}r_{i}=0\]👉Since, \(\gamma\) is constant for all \(n_j\) samples in the leaf, \(\sum _{x_{i}\in R_{jm}}\gamma =n_{j}\gamma \)
\[n_{j}\gamma =\sum _{x_{i}\in R_{jm}}r_{i}\]\[\implies \gamma =\frac{\sum _{x_{i}\in R_{jm}}r_{i}}{n_{j}}\]Therefore, \(\gamma\) = average of all residuals in the leaf.
Note: \(R_{jm}\)(Regions): Region of \(j_{th}\) leaf in \(m_{th}\)tree.
End of Section
1.2.4.11 - GBDT Algorithm
Gradient Boosted Decision Tree (GBDT) is a decision tree based implementation of Gradient Boosting Machine (GBM).
GBM treats the final model \(F_m(x)\) as weighted 🏋️♀️ sum of ‘m’ weak learners (decision trees):
\[ F_{M}(x)=\underbrace{F_{0}(x)}_{\text{Initial\ Guess}}+\nu \sum _{m=1}^{M}\underbrace{\left(\sum _{j=1}^{J_{m}}\gamma _{jm}\mathbb{I}(x\in R_{jm})\right)}_{\text{Decision\ Tree\ }h_{m}(x)}\]- \(F_0(x)\): The initial base model (usually a constant).
- M: The total number of boosting iterations (number of trees).
- \(\gamma_{jm}\)(Leaf Weight): The optimized value for leaf in tree .
- \(\nu\)(Nu): The Learning Rate or Shrinkage; prevent overfitting.
- \(\mathbb{I}(x\in R_{jm})\): ‘Indicator Function’; It is 1 if data point falls into leaf of the tree, and 0 otherwise.
- \(R_{jm}\)(Regions): Region of \(j_{th}\) leaf in \(m_{th}\)tree.
- Step 1: Initialization.
- Step 2: Iterative loop 🔁 : for i=1 to m.
- 2.1: Calculate pseudo residuals ‘\(r_{im}\)'.
- 2.2: Fit a regression tree 🌲‘\(h_m(x)\)'.
- 2.3:Compute leaf 🍃weights 🏋️♀️ ‘\(\gamma_{jm}\)'.
- 2.4:Update the model.
Start with a function that minimizes our loss function;
for MSE its mean.
MSE Loss = \(\mathcal{L}(y_i, \gamma) = \sum_{i=1}^n(y_i -\gamma)^2\)
Find the ‘gradient’ of error;
Tells us the direction and magnitude needed to reduce the loss.
Train a tree to predict the residuals ‘\(h_m(x)\)';
- Tree 🌲 partitions the data into leaves 🍃 (\(R_{jm}\)regions )
Within each leaf 🍃, we calculate the optimal value ‘\(\gamma_{jm}\)’ that minimizes the loss for the samples in that leaf 🍃.
\[\gamma_{jm} = \arg\min_{\gamma} \sum_{x_i \in R_{jm}} L(y_i, F_{m-1}(x_i) + \gamma)\]➡️ The optimal leaf 🍃value is the ‘Mean’(MSE) of the residuals; \(\gamma = \frac{\sum r_i}{n_j}\)
Add the new ‘correction’ to the existing model, scaled by the learning rate.
\[F_{m}(x)=F_{m-1}(x)+\nu \cdot \underbrace{\sum _{j=1}^{J_{m}}\gamma _{jm}\mathbb{I}(x\in R_{jm})}_{h_{m}(x)}\]- \(\mathbb{I}(x\in R_{jm})\): ‘Indicator Function’; It is 1 if data point falls into leaf of the tree, and 0 otherwise.
End of Section
1.2.4.12 - GBDT Example
Gradient Boosted Decision Tree (GBDT) is a decision tree based implementation of Gradient Boosting Machine (GBM).
GBM treats the final model \(F_m(x)\) as weighted 🏋️♀️ sum of ‘m’ weak learners (decision trees):
\[ F_{M}(x)=\underbrace{F_{0}(x)}_{\text{Initial\ Guess}}+\nu \sum _{m=1}^{M}\underbrace{\left(\sum _{j=1}^{J_{m}}\gamma _{jm}\mathbb{I}(x\in R_{jm})\right)}_{\text{Decision\ Tree\ }h_{m}(x)}\]- \(F_0(x)\): The initial base model (usually a constant).
- M: The total number of boosting iterations (number of trees).
- \(\gamma_{jm}\)(Leaf Weight): The optimized value for leaf in tree .
- \(\nu\)(Nu): The Learning Rate or Shrinkage; prevent overfitting.
- \(\mathbb{I}(x\in R_{jm})\): ‘Indicator Function’; It is 1 if data point falls into leaf of the tree, and 0 otherwise.
- \(R_{jm}\)(Regions): Region of \(j_{th}\) leaf in \(m_{th}\)tree.
- Step 1: Initialization.
- Step 2: Iterative loop 🔁 : for i=1 to m.
- 2.1: Calculate pseudo residuals ‘\(r_{im}\)'.
- 2.2: Fit a regression tree 🌲‘\(h_m(x)\)'.
- 2.3:Compute leaf 🍃weights 🏋️♀️ ‘\(\gamma_{jm}\)'.
- 2.4:Update the model.
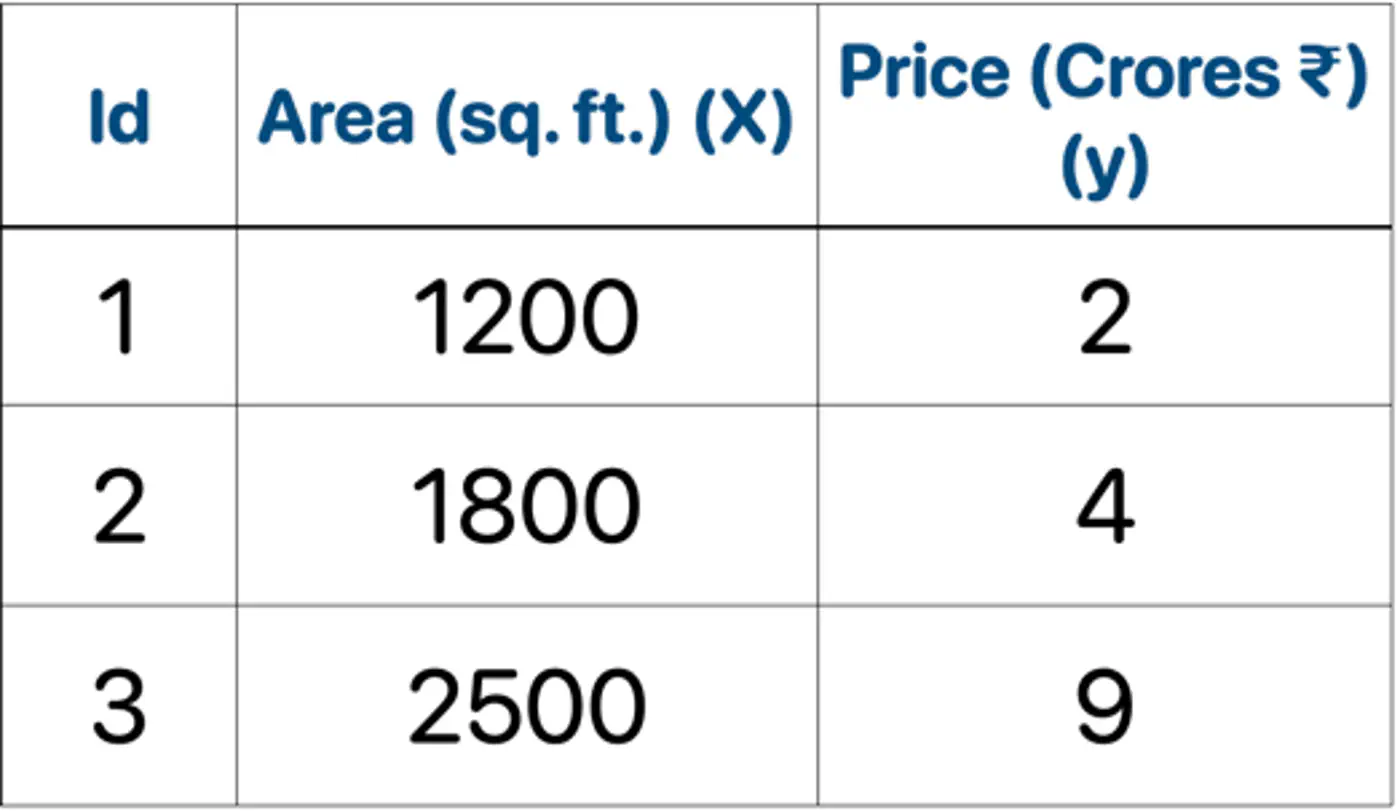
👉Loss = MSE, Learning rate (\(\nu\)) = 0.5
- Initialization : \(F_0(x) = mean(2,4,9) = 5.0\)
- Iteration 1(m=1):
2.1: Calculate residuals ‘\(r_{i1}\)'
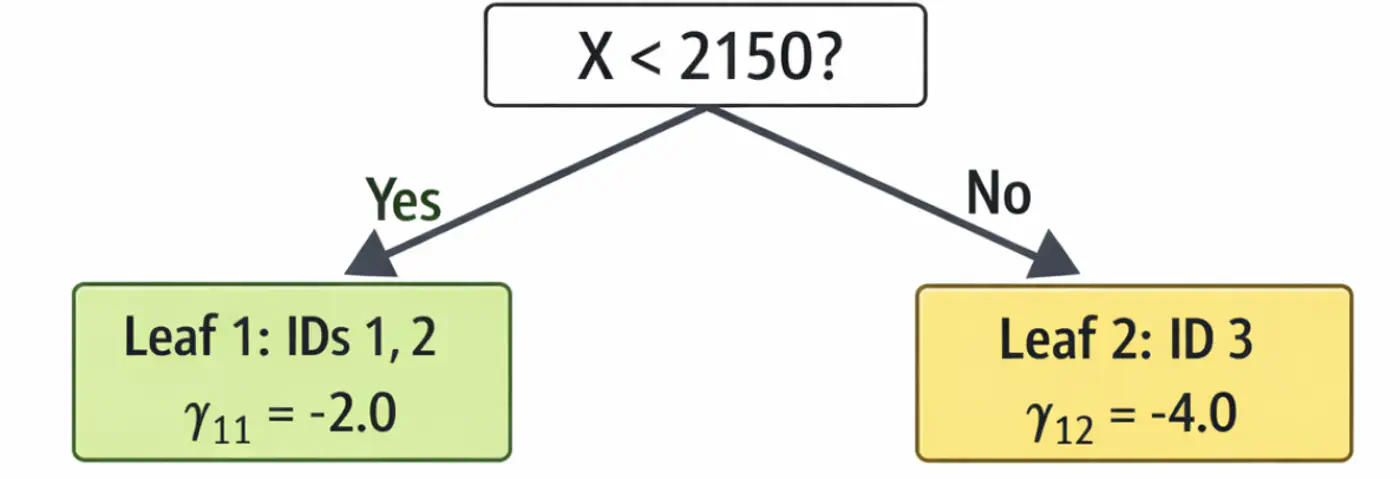
\[\begin{aligned} r_{11} &= 2-5 = -3.0 \\ r_{21} &= 4-5 = -1.0 \\ r_{31} &= 9-5 = 4.0 \\ \end{aligned} \]2.2: Fit tree(\(h_1\)); Split at X<2150 (midpoint of 1800 and 2500)
2.3: Compute leaf weights \(\gamma_{j1}\)
- Y-> Leaf 1: Ids 1, 2 ( \(\gamma_{11}\)= -2.0)
- N-> Leaf 2: Id 3 ( \(\gamma_{21}\)= 4.0)
2.4: Update predictions (\(F_1 = F_0 + 0.5 \cdot \gamma\))
\[ \begin{aligned} F_1(x_1) &= 5.0 + 0.5(-2.0) = \mathbf{4.0}\ \\F_1(x_2) &= 5.0 + 0.5(-2.0) = \mathbf{4.0}\ \\F_1(x_3) &= 5.0 + 0.5(4.0) = \mathbf{7.0}\ \\ \end{aligned} \]Tree 1:
- Iteration 2(m=2):
2.1: Calculate residuals ‘\(r_{i2}\)'
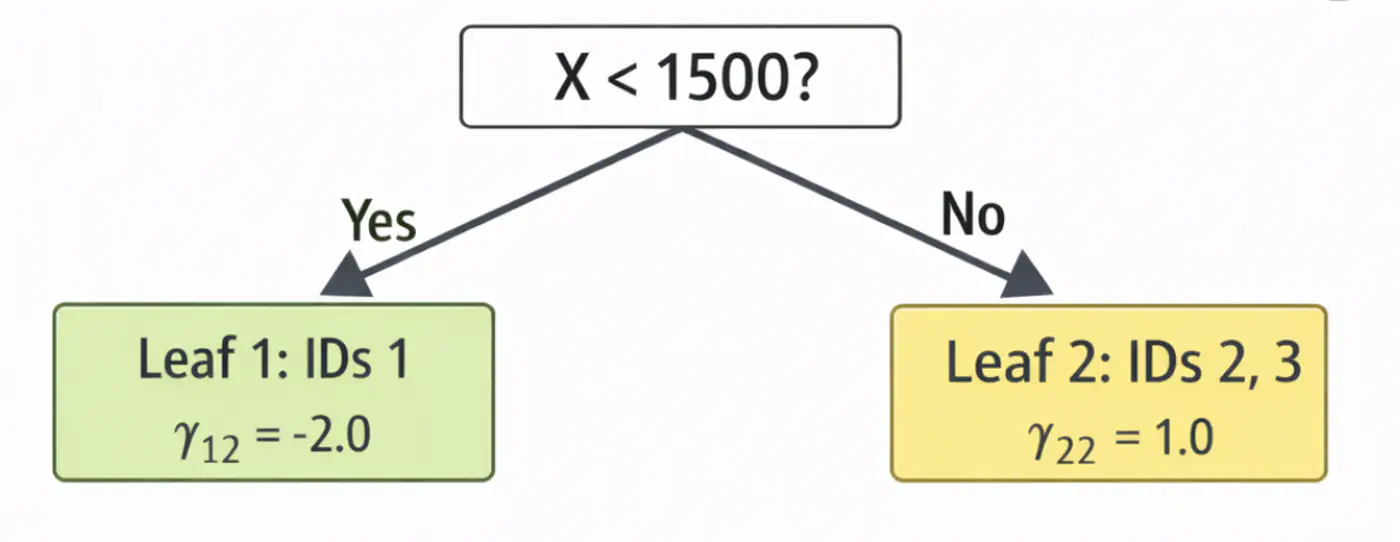
\[ \begin{aligned} r_{12} &= 2-4.0 = -2.0 \\ r_{22} &= 4-4.0 = 0.0 \\ r_{32} &= 9-7.0 = 2.0 \\ \end{aligned} \]2.2: Fit tree(\(h_2\)); Split at X<1500 (midpoint of 1200 and 1800)
2.3: Compute leaf weights \(\gamma_{j2}\)
- Y-> Leaf 1: Ids 1 ( \(\gamma_{12}\)= -2.0)
- N-> Leaf 2: Id 2, 3 ( \(\gamma_{22}\)= 1.0)
2.4: Update predictions (\(F_1 = F_0 + 0.5 \cdot \gamma\))
\[ \begin{aligned} F_2(x_1) &= 4.0 + 0.5(-2.0) = \mathbf{3.0} \\F_2(x_2) &= 4.0 + 0.5(1.0) = \mathbf{4.5} \\ F_2(x_3) &= 7.0 + 0.5(1.0) = \mathbf{7.5}\ \\ \end{aligned} \]Tree 2:
Note: We can keep adding more trees with every iteration;
ideally, learning rate \(\nu\) is small, say 0.1, so that we do not overshoot and converge slowly.
Let’s predict the price of a house with area = 2000 sq. ft.
- \(F_{0}=5.0\)
- Pass though tree 1 (\(h_1\)): is 2000 < 2150 ? Yes, \(\gamma_{11}\)= -2.0
- Contribution (\(h_1\)) = 0.5 x (-2.0) = -1.0
- Pass though tree 2 (\(h_2\)): is 2000 < 1500 ? No, \(\gamma_{22}\) = 1.0
- Contribution(\(h_2\)) = 0.5 x (1.0) = 0.5
- Final prediction = 5.0 - 1.0 + 0.5 = 4.5
Therefore, the price of a house with area = 2000 sq. ft is Rs 4.5 crores, which is very close.
In just 2 iterations, although with higher learning rate (\(\nu=0.5\)), we were able to get a fairly good estimate.
End of Section
1.2.4.13 - Advanced GBDT Algorithms
🔵 LightGBM (Light Gradient Boosting Machine)
⚫️ CatBoost (Categorical Boosting)
⭐️An optimized and highly efficient implementation of gradient boosting.
👉 Widely used in competitive data science (like Kaggle) due to its speed and performance.
Note: Research project developed by Tianqi Chen during his doctoral studies at the University of Washington.
⭐️Developed by Microsoft, this framework is designed for high speed and efficiency with large datasets.
👉It grows trees leaf-wise rather than level-wise and uses Gradient-based One-Side Sampling (GOSS) to speed 🐇 up the finding of optimal split points.
End of Section
1.2.4.14 - XgBoost
⭐️An optimized and highly efficient implementation of gradient boosting.
👉 Widely used in competitive data science (like Kaggle) due to its speed and performance.
Note: Research project developed by Tianqi Chen during his doctoral studies at the University of Washington.
🔵 Regularization
🔵 Sparsity-Aware Split Finding
⭐️Uses second derivative (Hessian), i.e, curvature, in addition to first derivative (gradient) to optimize the objective function more quickly and accurately than GBDT.
Let’s understand this with the problem to minimize \(f(x) = x^4\), using:
Gradient descent (uses only 1st order derivative, \(f'(x) = 4x^3\))
Newton’s method (uses both 1st and 2nd order derivatives \(f''(x) = 12x^2\))
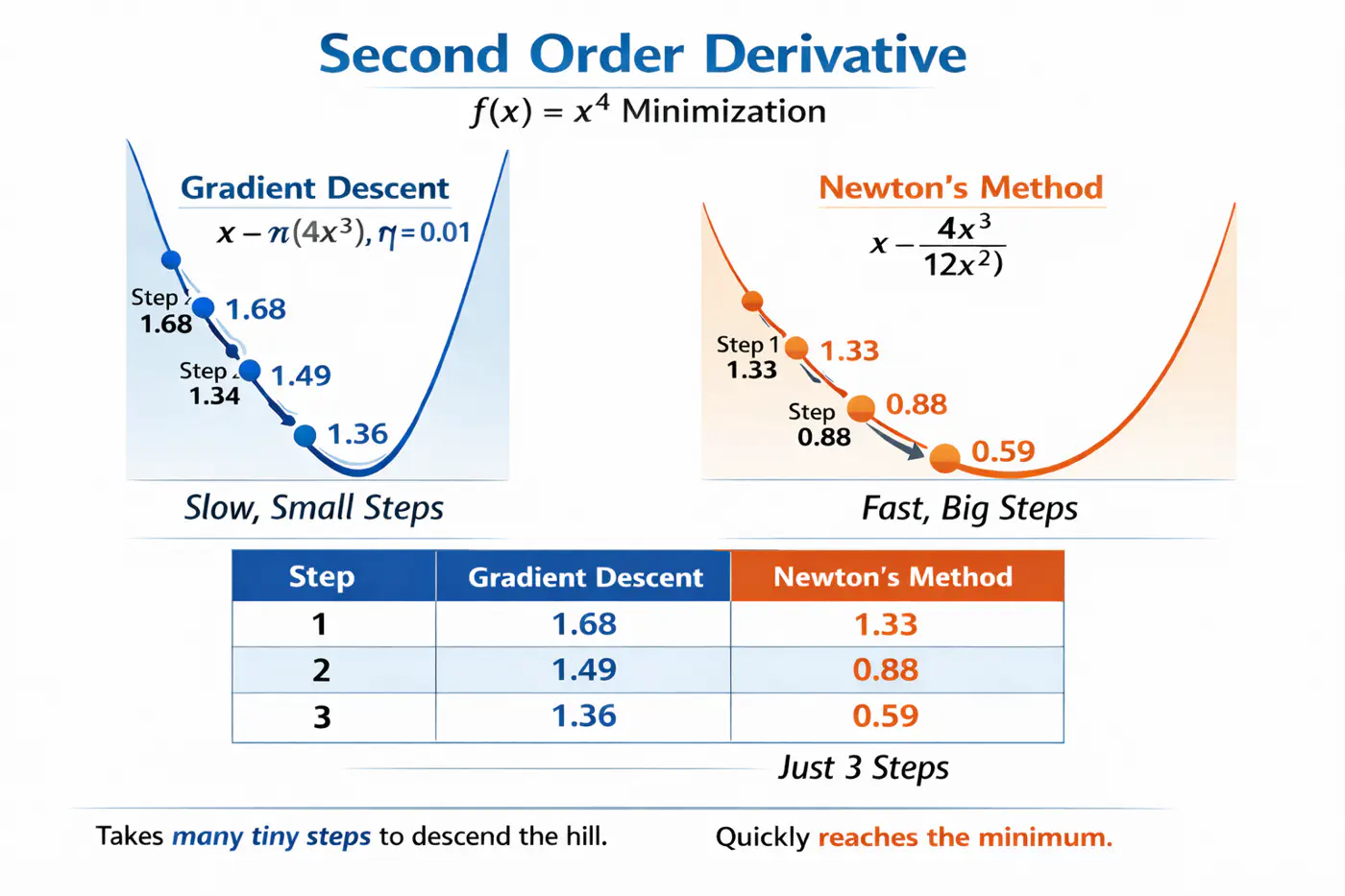
- Adds explicit regularization terms (L1/L2 on leaf weights and tree complexity) into the boosting objective, helping reduce over-fitting. \[ \text{Objective} = \underbrace{\sum_{i=1}^{n} L(y_i, \hat{y}_i)}_{\text{Training Loss}} + \underbrace{\gamma T + \frac{1}{2}\lambda \sum_{j=1}^{T} w_j^2 + \alpha \sum_{j=1}^{T} |w_j|}_{\text{Regularization (The Tax)}} \]
💡Real-world data often contains many missing values or zero-entries (sparse data).
👉 XGBoost introduces a ‘default direction’ for each node.
➡️During training, it learns the best direction (left or right) for missing values to go, making it significantly faster and more robust when dealing with sparse or missing data.
End of Section
1.2.4.15 - LightGBM
⭐️Developed by Microsoft, this framework is designed for high speed and efficiency with large datasets.
👉It grows trees leaf-wise rather than level-wise and uses Gradient-based One-Side Sampling (GOSS) to speed 🐇 up the finding of optimal split points.
🔵 Exclusive Feature Bundling (EFB)
🔵 Leaf-wise Tree Growth Strategy
- ❌ Traditional GBDT uses all data instances for gradient calculation, which is inefficient.
- ✅ GOSS focuses 🔬on instances with larger gradients (those that are less well-learned or have higher error).
- 🐛 Keeps all instances with large gradients but randomly samples from those with small gradients.
- 🦩This way, it prioritizes the most informative examples for training, significantly reducing the data used and speeding up 🐇 the process while maintaining accuracy.
- 🦀 High-dimensional data often contains many sparse, mutually exclusive features (features that never take a non-zero value simultaneously, such as, One Hot Encoding (OHE)).
- 💡 EFB bundles the non-overlapping features into a single, dense feature, reducing the number of features, without losing much information, saving computation.
- ❌ Traditional gradient boosting machines (like XGBoost), the trees are built level-wise (BFS-like), meaning all nodes at the current level are split before moving to the next level.
- ✅ LightGBM maintains a set of all potential leaves that can be split at any given time and selects the leaf (for splitting) that provides the maximum gain across the entire tree, regardless of its depth.
Note: Need mechanisms to avoid over-fitting.
End of Section
1.2.4.16 - CatBoost
🔵 Symmetric(Oblivious) Trees
🔵 Handling Missing Values
- ❌ Standard target encoding can lead to target leakage, where the model uses information from the target variable during training that would not be available during inference.
👉(model ‘cheats’ by using a row’s own label to predict itself). - ✅ CatBoost calculates the target statistics (average target value) for each category based only on the history of previous training examples in a random permutation of the data.
🦋 Uses symmetric decision trees by default.
👉 In symmetric trees, the same split condition is applied at each level across the entire tree structure.🦘Does not walk down the tree using ‘if-else’ logic, instead it evaluates decision conditions to create a binary index (e.g 101) and jumps directly to that leaf 🍃 in memory 🧠.
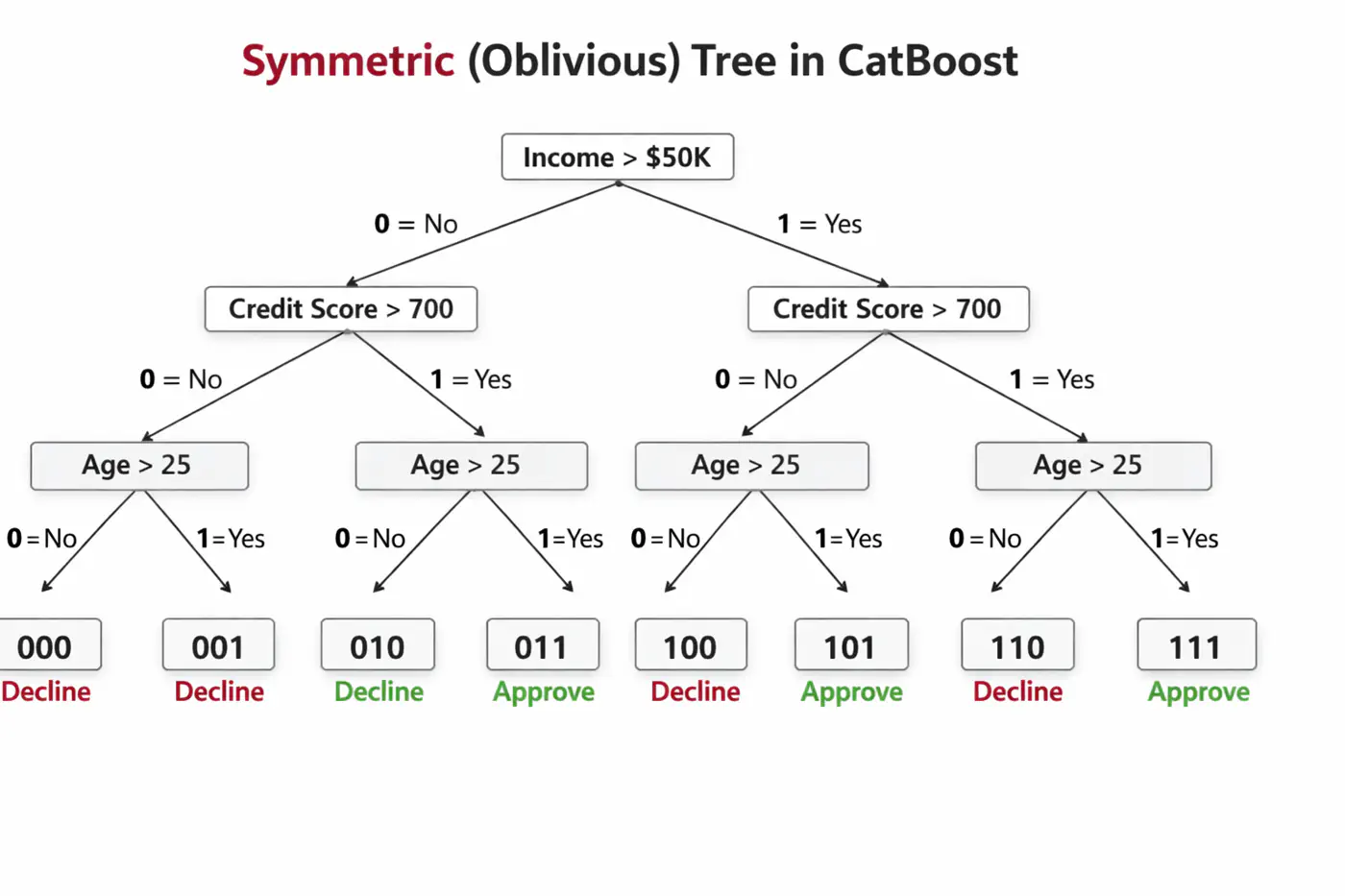
⚙️ CatBoost offers built-in, intelligent handling of missing values and sparse features, which often require manual preprocessing in other GBDT libraries.
💡Treats ‘NaN’ as a distinct category, reducing the need for imputation.
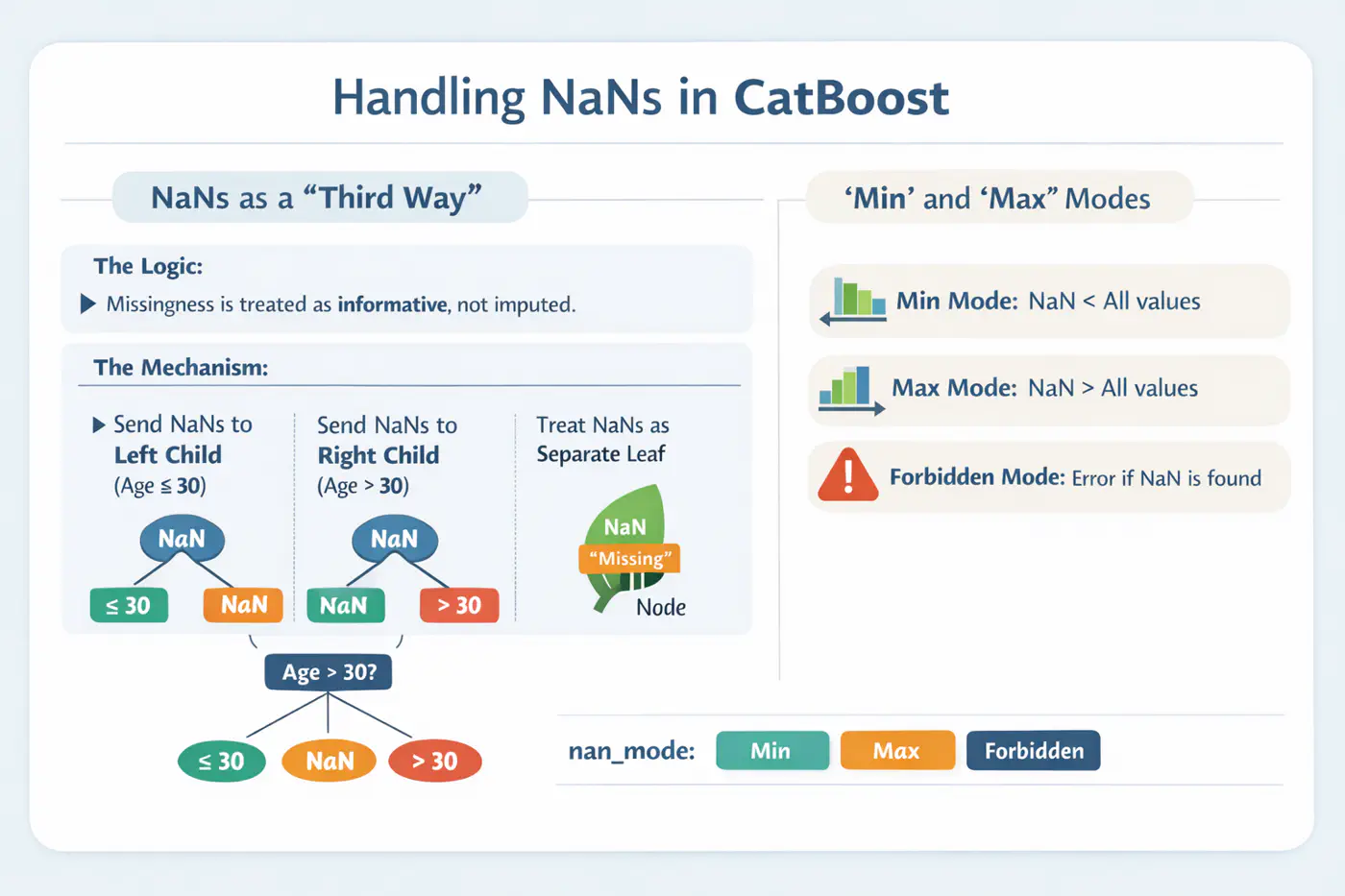
End of Section
1.2.5 - Support Vector Machine
End of Section
1.2.5.1 - SVM Intro
⭐️We have two classes of points (e.g. Cats 😸vs. Dogs 🐶) that can be separated by a straight line.
👉 Many such lines exist !
💡SVM asks: “Which line is the safest?”
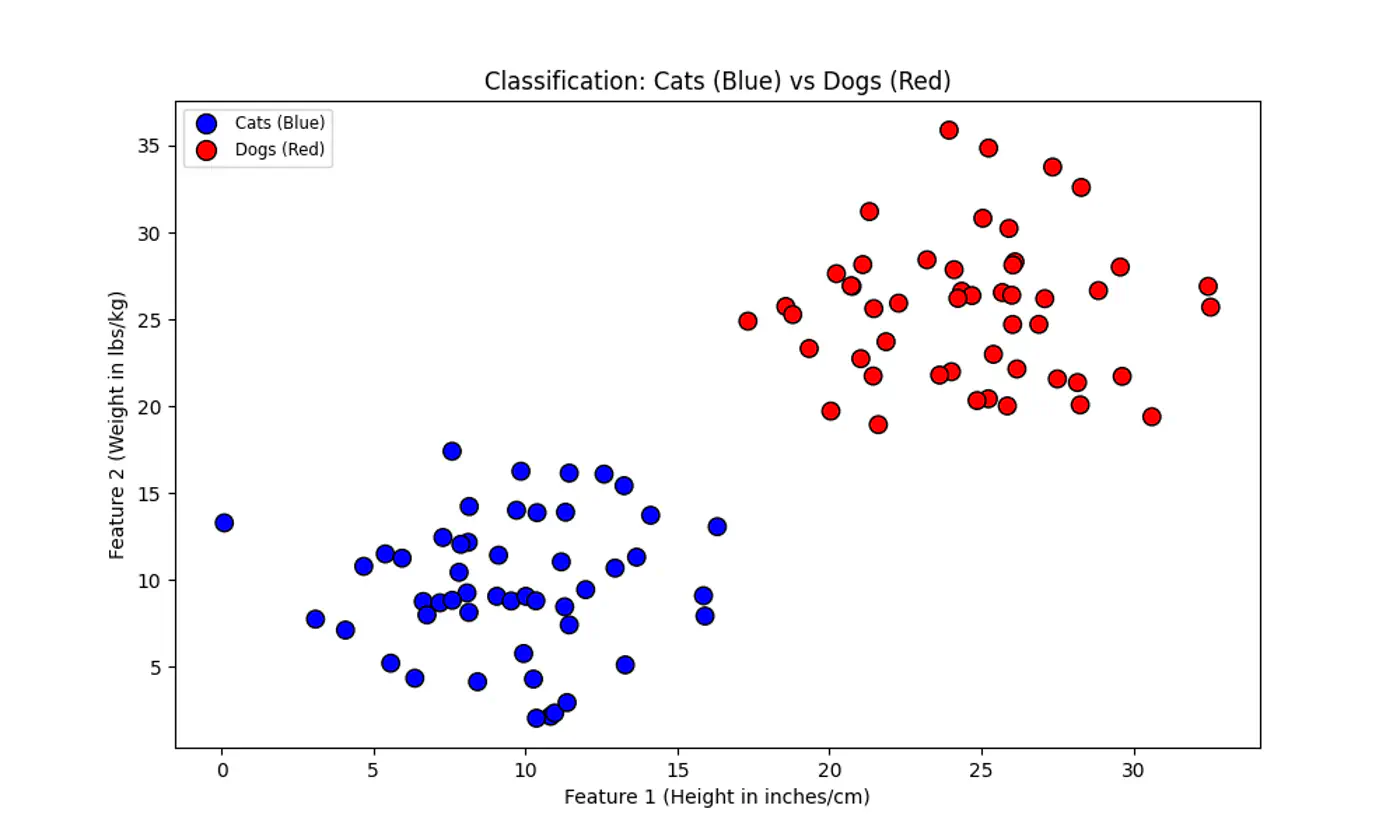
💡Think of the decision boundary as the center-line of a highway 🛣️.
SVM tries to make this highway 🛣️ as wide as possible without hitting any ‘buildings’ 🏡 (data points) on either side.
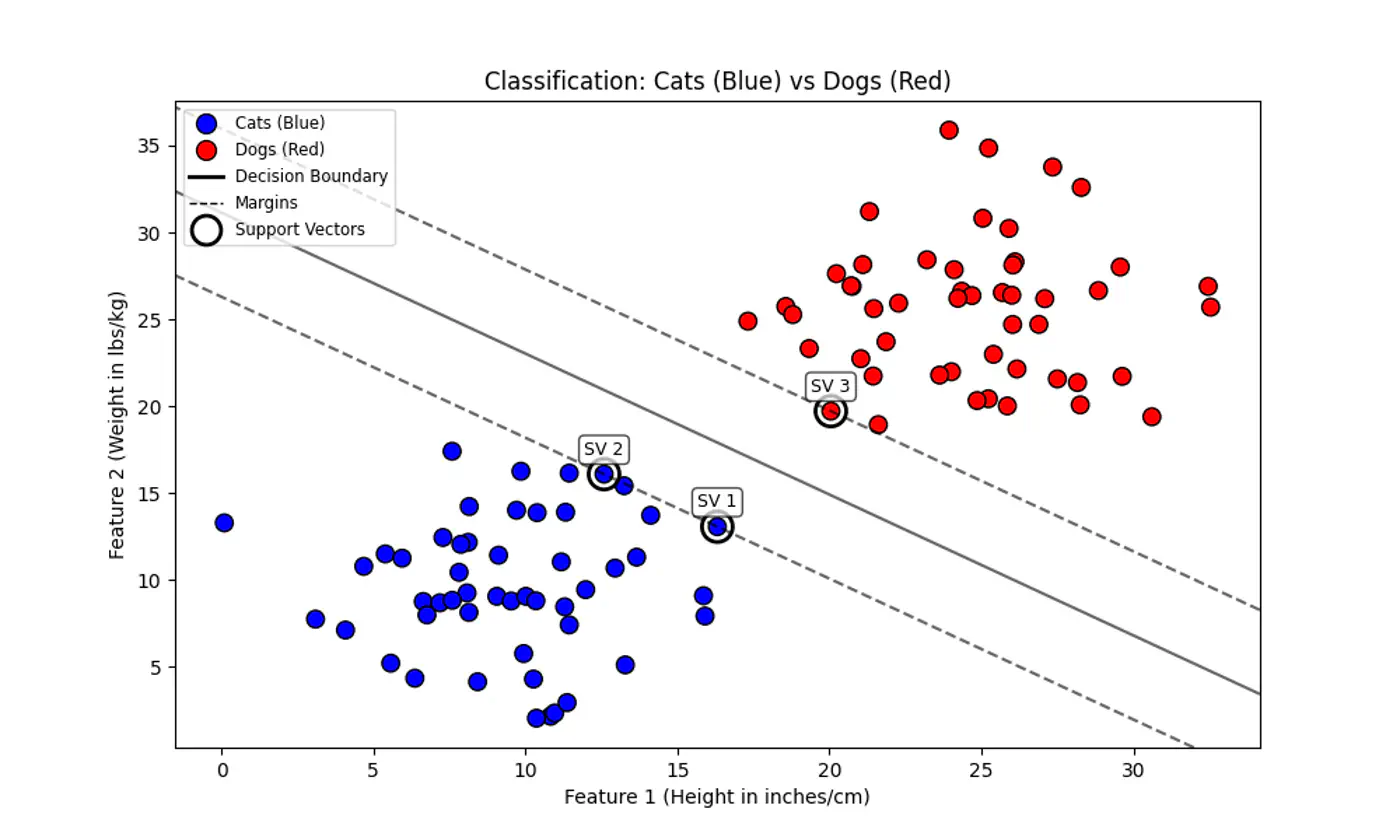
End of Section
1.2.5.2 - Hard Margin SVM
Data is perfectly linearly separable, i.e, there must exist a hyperplane that can perfectly separate the data into two distinct classes without any misclassification.
No noise or outliers that fall within the margin or on the wrong side of the decision boundary. Note: Even a single outlier can prevent the algorithm from finding a valid solution or drastically affect the boundary’s position, leading to poor generalization.
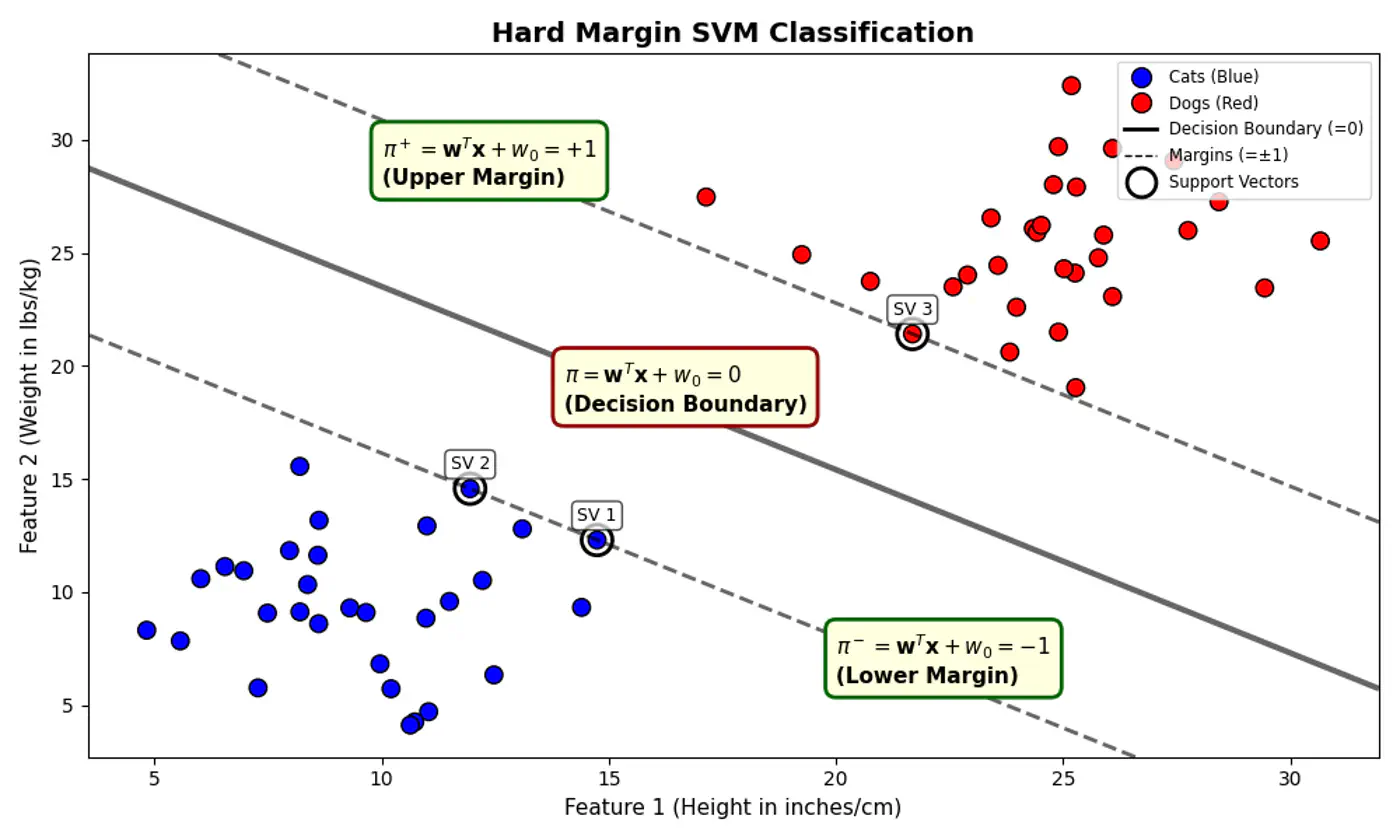
- 🐎 Distance(signed) of a hyperplane from origin = \(\frac{-w_0}{\|w\|}\)
- 🦣 Margin width = distance(\(\pi^+, \pi^-\))
- = \(\frac{1-w_0 - (-1 -w_0)}{\|w\|}\) = \(\frac{1-\cancel{w_0} + 1 + \cancel{w_0})}{\|w\|}\)
- distance(\(\pi^+, \pi^-\)) = \(\frac{2}{\|w\|}\)
Figure: Distance of Hyperplane from Origin
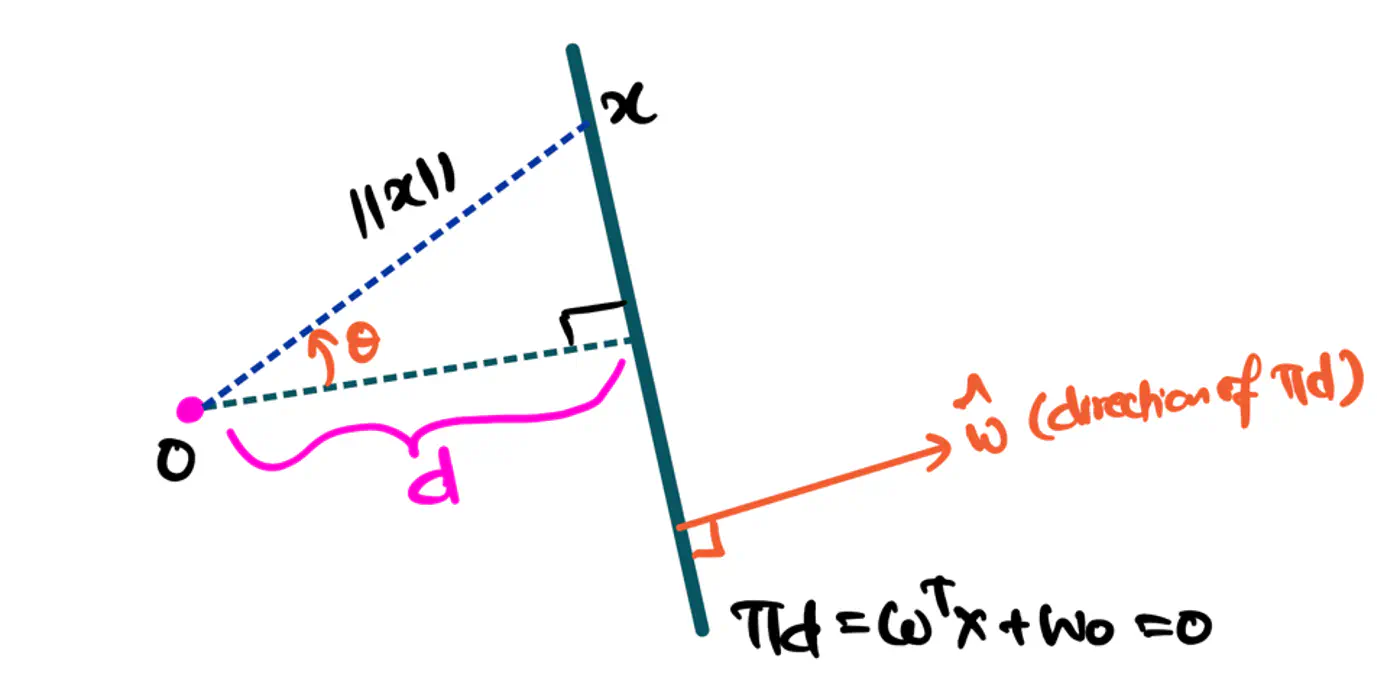
- Separating hyperplane \(\pi\) is exactly equidistant from \(\pi^+\) and \(\pi^-\).
- We want to maximize the margin between +ve(🐶) and -ve (😸) points.
👉Combining above two constraints:
\[y_{i}.(w^{T}x_{i}+w_{0})≥1\]such that, \(y_i.(w^Tx_i + w_0) \ge 1\)
👉To maximize the margin, we must minimize \(\|w\|\).
Since, distance(\(\pi^+, \pi^-\)) = \(\frac{2}{\|w\|}\)
such that, \(y_i.(w^Tx_i + w_0) \ge 1 ~ \forall i = 1,2,\dots, n\)
Note: Hard margin SVM will not work if the data has a single outlier or slight overlap.
End of Section
1.2.5.3 - Soft Margin SVM
💡Imagine the margin is a fence 🌉.
Hard Margin: fence is made of steel.
Nothing can cross it.Soft Margin: fence is made of rubber(porous).
Some points can ‘push’ into the margin or even cross over to the wrong side, but we charge them a penalty 💵 for doing so.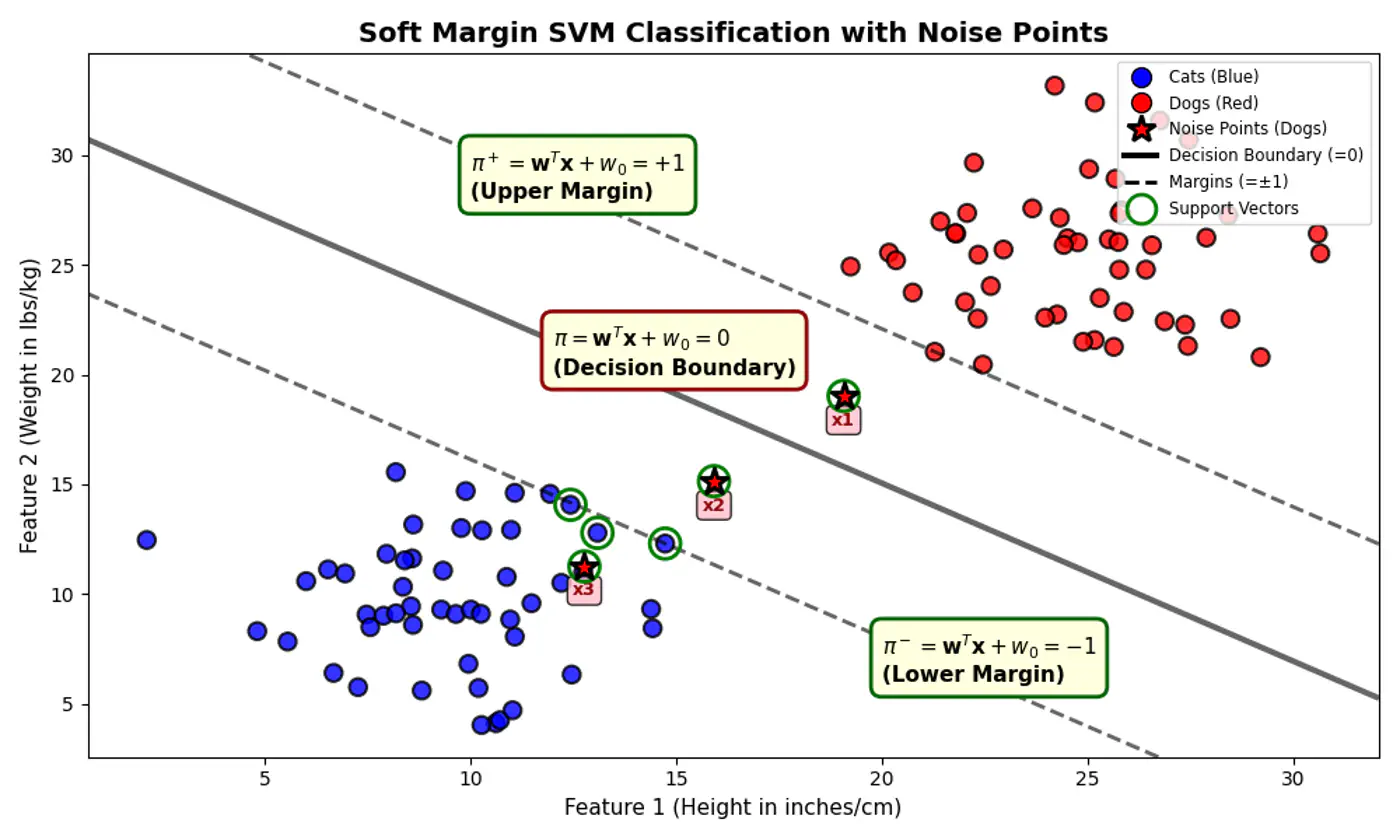
Distance from decision boundary:
- Distance of positive labelled points must be \(\ge 1\)
- But, distance of noise 📢 points (actually positive points) \(x_1, x_2 ~\&~ x_3\) < 1
⚔️ So, we introduce a slack variable or allowance for error term, \(\xi_i\) (pronounced ‘xi’) for every single data point.
\[y_i.(w^Tx_i + w_0) \ge 1 - \xi_i, ~ \forall i = 1,2,\dots, n\]\[ \implies \xi_i \ge 1 - y_i.(w^Tx_i + w_0) \\ also, ~ \xi_i \ge 0 \]\[So, ~ \xi _{i}=\max (0,1-y_{i}\cdot (w^Tx_i + w_0))\]Note: The above error term is also called ‘Hinge Loss’.
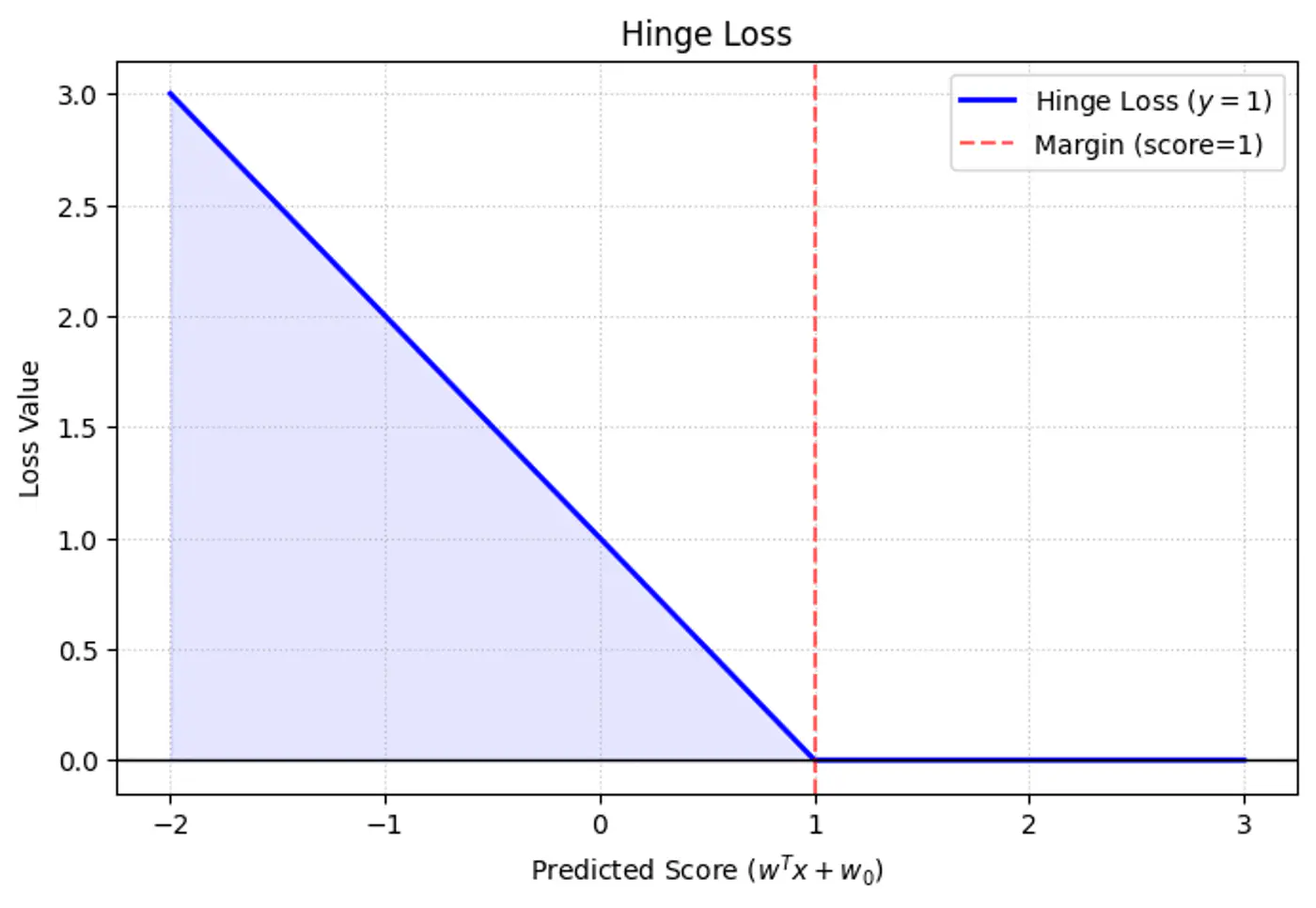
Hinge

- \(\xi_i = 0\) : Correctly classified and outside (or on) the margin.
- \(0 < \xi_i \le 1 \) : Within the margin but on the correct side of the decision boundary.
- \(\xi_i > 0\): On the wrong side of the decision boundary (misclassified).
e.g.: Since, the noise 📢 point are +ve (\(y_i=1\)) labeled:
\[\xi _{i}=\max (0,1-f(x_{i}))\]\(x_1, (d=+0.5)\): \(\xi _{i}=\max (0,1-0.5) = 0.5\)
\(x_2, (d=-0.5)\): \(\xi _{i}=\max (0,1-(-0.5))= 1.5\)
\(x_3, (d=-1.5)\): \(\xi _{i}=\max (0,1-(-1.5)) = 2.5\)
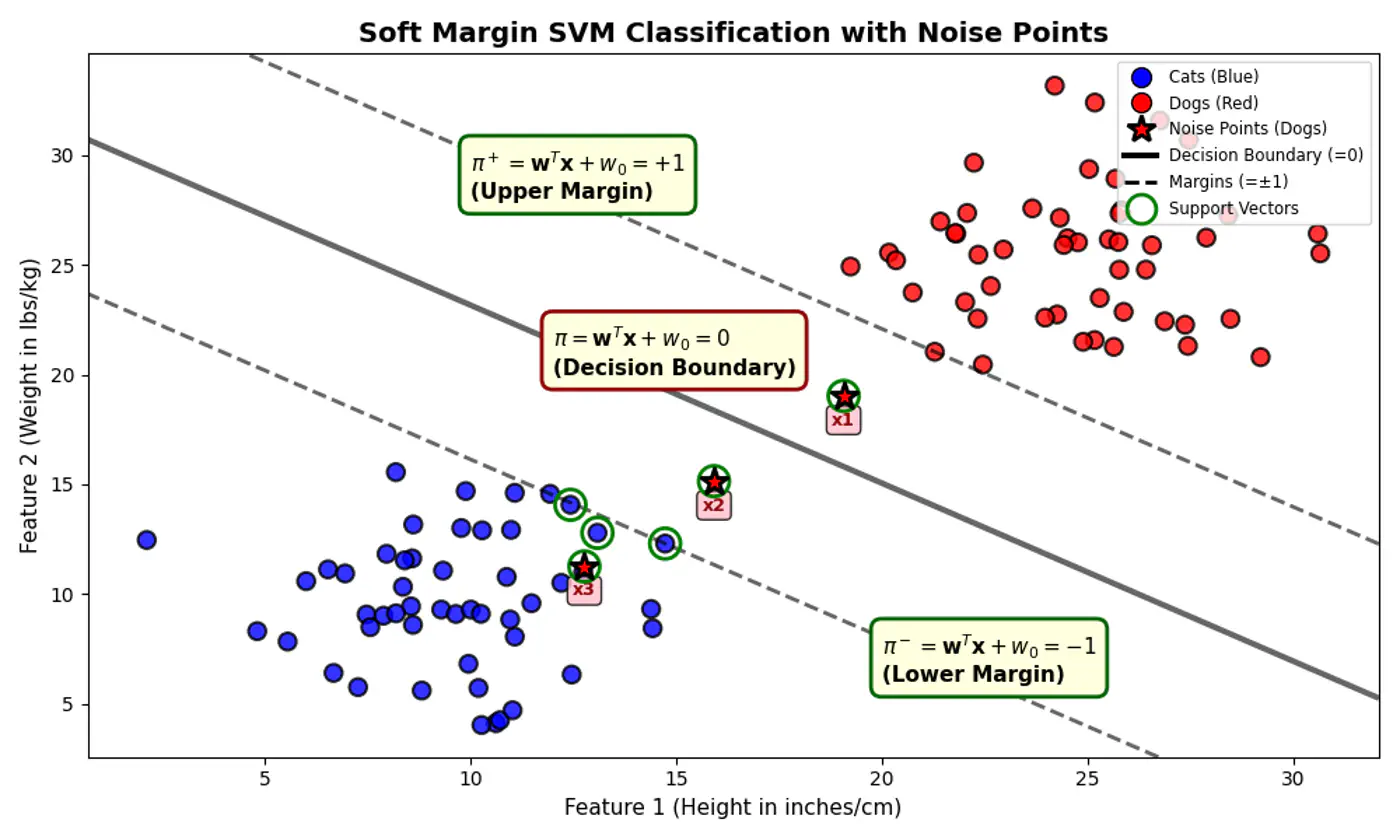
Subject to constraints:
- \(y_i(w^T x_i + b) \geq 1 - \xi_i\): The ‘softened’ margin constraint.
- \(\xi_i \geq 0\): Slack/Error cannot be negative.
Note: We use a hyper-parameter ‘C’ to control the trade-off.
- Large ‘C’: Over-Fitting;
Misclassifications are expensive 💰.
Model tries to keep the errors as low as possible. - Small ‘C’: Under-Fitting;
Margin width is more important than individual errors.
Model will ignore outliers/noise to get a ‘cleaner’(wider) boundary.
Note: SVM is just L2-Regularized Hinge Loss minimization, as Logistic Regression minimizes Log-Loss.
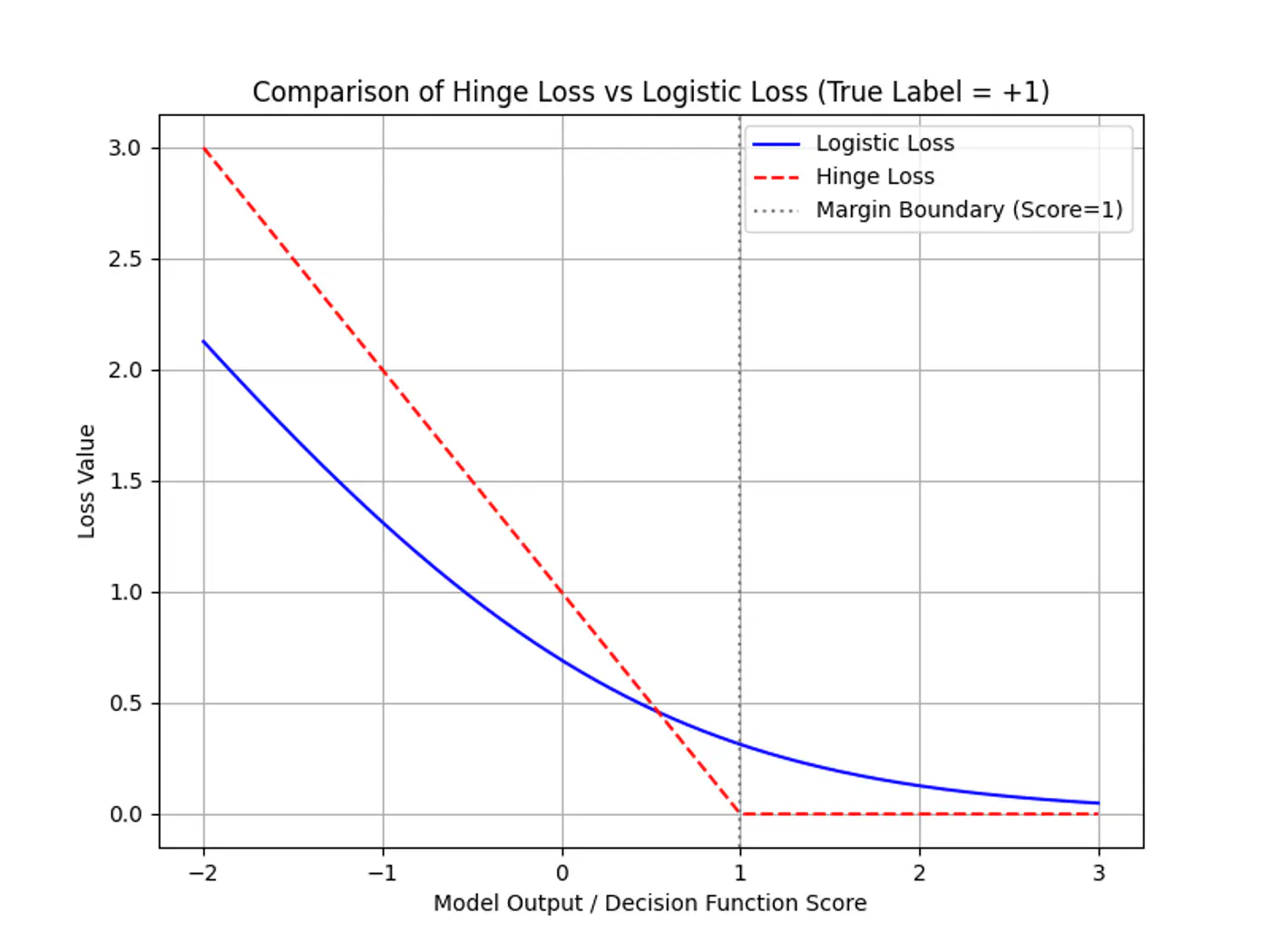
End of Section
1.2.5.4 - Primal Dual Equivalence
The Primal form is intuitive but computationally expensive in high dimensions.
➡️ Number of new features = \({d+p \choose p}\), grows roughly as \(O(d^{p})\)
- d= number of dimensions (features)
- p = degree of polynomial
Note: The Dual form is what enables the Kernel Trick 🪄.
Subject to constraints:
- \(y_i(w^T x_i + b) \geq 1 - \xi_i\): The ‘softened’ margin constraint.
- \(\xi_i \geq 0\): Slack/Error cannot be negative.
⭐️ We start with the Soft-Margin Primal objective and incorporate its constraints using Lagrange Multipliers \((\alpha_i, \mu_i \geq 0)\)
\[L(w, w_0, \xi, \alpha, \mu) = \frac{1}{2}\|w\|^2 + C \sum_{i=1}^n \xi_i - \sum_{i=1}^n \alpha_i \left[y_i(w^T x_i + w_0) - 1 + \xi_i \right] - \sum_{i=1}^n \mu_i \xi_i\]Note: Inequality conditions must be \(\le 0\).
👉The Lagrangian function has two competing objectives:
\[L(w, w_0, \xi, \alpha, \mu) = \frac{1}{2}\|w\|^2 + C \sum_{i=1}^n \xi_i - \sum_{i=1}^n \alpha_i \left[y_i(w^T x_i + w_0) - 1 + \xi_i \right] - \sum_{i=1}^n \mu_i \xi_i\]- Minimization: We want to minimize \(L(w, w_0, \xi, \alpha, \mu)\) w.r.t primal variables (\(w, w_0, \xi_i \) ) to find the hyperplane with the largest possible margin.
- Maximization: We want to maximize \(L(w, w_0, \xi, \alpha, \mu)\) w.r.t dual variables (\(\alpha_i, \mu_i\) ) to ensure all training constraints are satisfied.
Note: A point that is simultaneously a minimum for one set of variables and a maximum for another is, by definition, a saddle point.
👉To find the Dual, we find the saddle point by taking partial derivatives with respect to the primal variables \((w, w_0, \xi)\) and equating them to 0.
\[\frac{\partial L}{\partial w} = 0 \implies \mathbf{w = \sum_{i=1}^n \alpha_i y_i x_i}\]\[\frac{\partial L}{\partial w_0} = 0 \implies \mathbf{\sum_{i=1}^n \alpha_i y_i = 0}\]\[\frac{\partial L}{\partial \xi_i} = 0 \implies C - \alpha_i - \mu_i = 0 \implies \mathbf{0 \leq \alpha_i \leq C}\]subject to: \(0 \leq \alpha_i \leq C\) and \(\sum \alpha_i y_i = 0\)
- \(\alpha_i\)= 0, for non support vectors (correct side)
- \(0 < \alpha_i < C\), for free support vectors (exactly on the margin)
- \(\alpha_i = C\), for bounded support vectors (misclassified or inside margin)
Note: Sequential Minimal Optimization (SMO) algorithm is used to find optimal \(\alpha_i\) values.
🎯To classify unseen point \(x_q\) : \(f(x_q) = \text{sign}(w^T x_q + w_0)\)
✅ From the KKT stationarity condition, we know: \(\mathbf{w} = \sum_{i=1}^n \alpha_i y_i x_i\)
👉 Substituting this into the equation:
\[f(x_q) = \text{sign}\left( \sum_{i=1}^n \alpha_i y_i (x_i^T x_q) + w_0 \right)\]Note: Even if you have 1 million training points, if only 500 are support vectors, the summation only runs for 500 terms.
All other points have \(\alpha_i = 0\) and vanish.
End of Section
1.2.5.5 - Kernel Trick
👉If our data is not linearly separable in its original space , we can map it to a higher-dimensional feature space (where D»d) using a transformation function .
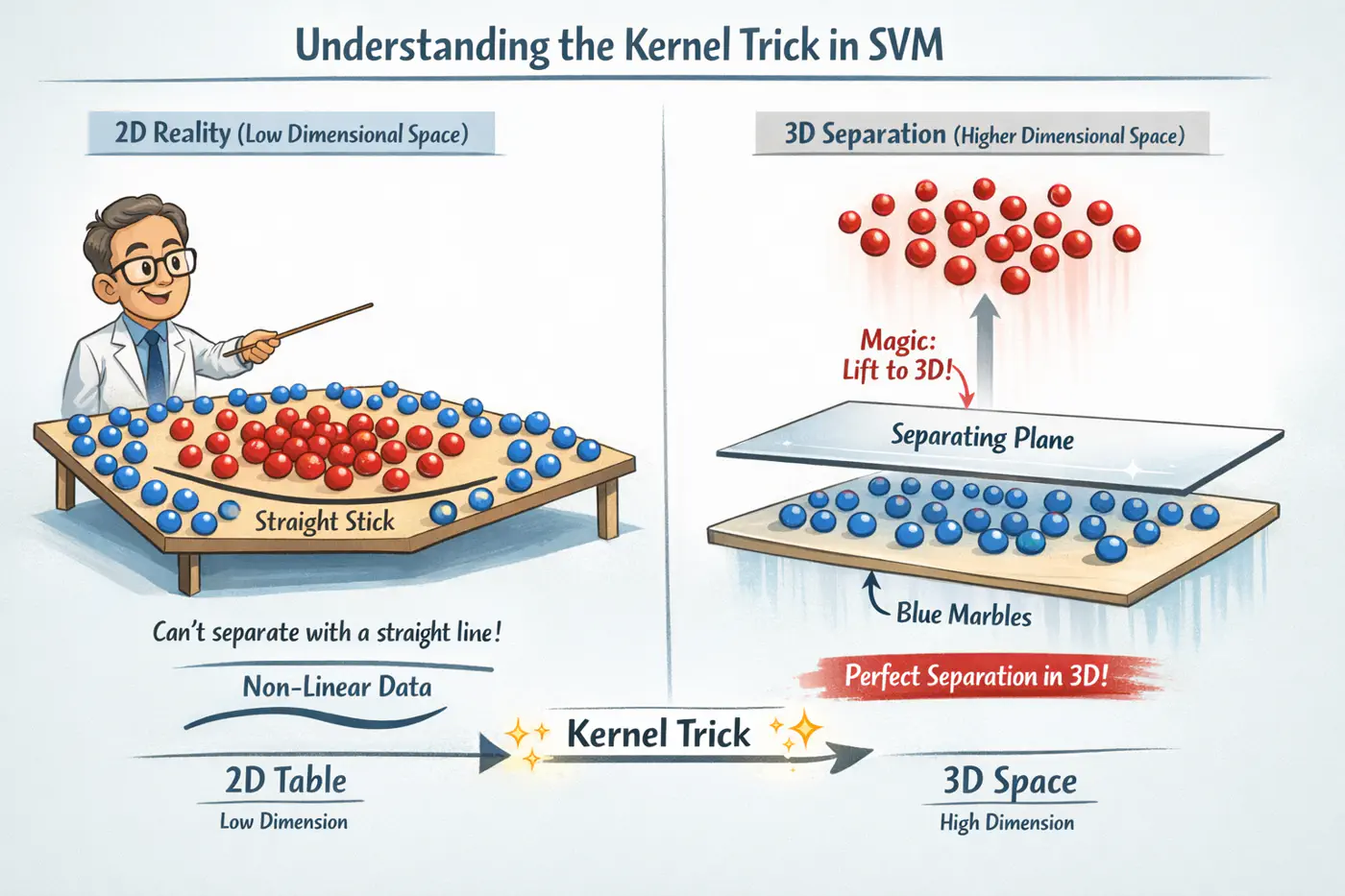
- Bridge between Dual formulation and the geometry of high dimensional spaces.
- It is a way to manipulate inner product spaces without the computational cost 💰 of explicit transformation.
subject to: \(0 \leq \alpha_i \leq C\) and \(\sum \alpha_i y_i = 0\)
💡Actual values of the input vectors \(x_i\) and \(x_j\) never appear in isolation; only appear as inner product.
\[ \max_{\alpha} \sum_{i=1}^n \alpha_i - \frac{1}{2} \sum_{i,j=1}^n \alpha_i \alpha_j y_i y_j \mathbf{(x_i \cdot x_j)}\]\[f(x_q) = \text{sign}\left( \sum_{i=1}^n \alpha_i y_i (x_i^T x_q) + w_0 \right)\]👉The ‘shape’ of the decision boundary is entirely determined by how similar points are to one another, not by their absolute coordinates.
So we define a function called ‘Kernel Function’.
The ‘Kernel Trick’ 🪄 is an optimization that replaces the dot product of a high-dimensional mapping with a function of the dot product in the original space.
\[(x_i, x_j) = \langle \phi(x_i), \phi(x_j) \rangle\]💡How it works ?
Instead of mapping (to higher dimension) \(x_i \rightarrow \phi(x_i)\), \(x_j \rightarrow \phi(x_j)\),
and calculating the dot product.
We simply compute \(K(x_i, x_j)\) directly in the original input space.
👉The ‘Kernel Function’ gives the similar mathematical equivalent of mapping it to a higher dimensions and taking the dot product.
Note: For \(K(x_i, x_j)\) to be a valid kernel, it must satisfy Mercer’s Condition.
Below is an example for a quadratic Kernel function in 2D that is equivalent to mapping the vectors to 3D and taking a dot product in 3D.
\[K(x, z) = (x^T z)^2\]\[(x_1 z_1 + x_2 z_2)^2 = x_1^2 z_1^2 + 2x_1 z_1 x_2 z_2 + x_2^2 z_2^2\]The output of above quadratic kernel function is equivalent to the explicit dot product of two vectors in 3D:
\[\phi(x) = [x_1^2, \sqrt{2}x_1x_2, x_2^2]^T\]\[\phi(z) = [z_1^2, \sqrt{2}z_1z_2, z_2^2]^T\]\[\phi (x)\cdot \phi (z)=x_{1}^{2}z_{1}^{2}+2x_{1}x_{2}z_{1}z_{2}+x_{2}^{2}z_{2}^{2}\]- Computational Efficiency: Avoids the ‘combinatorial blowup’ 💥 of generating thousands of interaction features manually.
- Memory Savings: No need to store 💾 or process high-dimensional coordinates, only the scalar result of the kernel function.
- Designing special purpose domain specific kernel is very hard.
- Basically, we are trying to replace feature engineering with kernel design.
- Note: Deep learning does feature engineering implicitly for us.
- Runtime complexity depends on number of support vectors, whose count is not easy to control.
Note: Runtime Time Complexity ⏰ = \(O(n_{SV}\times d)\) , whereas linear SVM,\(O(d)\) .
End of Section
1.2.5.6 - RBF Kernel
- Unlike the polynomial kernel, which looks at global 🌎 interactions, the RBF kernel acts like a similarity measure.
- If ‘x’ and ‘z’ are identical \(K(x,z)=1\).
- As they move further apart in Euclidean space, the value decays exponentially towards 0.
- If \(x \approx z\) (very close), \(K(x,z)=1\)
- If ‘x’, ‘z’ are far apart, \(K(x,z) \approx 0\)
Note: Kernel function is the measure of similarity or closeness.
Say \(\sigma = 1\), then Euclidean distance: \(\|x - z\|^2 = \|x\|^2 + \|z\|^2 - 2x^Tz\)
\[K(x, z) = \exp(-( \|x\|^2 + \|z\|^2 - 2x^T z )) = \exp(-\|x\|^2) \exp(-\|z\|^2) \exp(2x^T z)\]The Taylor expansion for \(e^u= \sum_{n=0}^{\infty} \frac{u^n}{n!}\)
\[\exp(2x^T z) = \sum_{n=0}^{\infty} \frac{(2x^T z)^n}{n!} = 1 + \frac{2x^T z}{1!} + \frac{(2x^T z)^2}{2!} + \dots + \frac{(2x^T z)^n}{n!} + \dots\]\[K(x, z) = e^{-\|x\|^2} e^{-\|z\|^2} \left( \sum_{n=0}^{\infty} \frac{2^n (x^T z)^n}{n!} \right)\]💡If we expand each \((x^T z)^n\) term, it represents the dot product of all possible n-th order polynomial features.
👉Thus, the implicit feature map is:
\[\phi(x) = e^{-\|x\|^2} \left[ 1, \sqrt{\frac{2}{1!}}x, \sqrt{\frac{2^2}{2!}}(x \otimes x), \dots, \sqrt{\frac{2^n}{n!}}(\underbrace{x \otimes \dots \otimes x}_{n \text{ times}}), \dots \right]^T\]- Important: The tensor product \(x\otimes x\) creates a vector (or matrix) containing all combinations of the features. e.g. if \(x=[x_{1},x_{2}]\), then \(x\otimes x=[x_{1}^{2},x_{1}x_{2},x_{2}x_{1},x_{2}^{2}]\) 
Note: Because the Taylor series has an infinite number of terms, feature map has an infinite number of dimensions.
- High Gamma(low \(\sigma\)): Over-Fitting
- Makes the kernel so ‘peaky’ that each support vector only influences its immediate neighborhood.
- Decision boundary becomes highly irregular, ‘wrapping’ tightly around individual data points to ensure they are classified correctly.
- Low Gamma(high \(\sigma\)): Under-Fitting
- The Gaussian bumps are wide and flat.
- Decision boundary becomes very smooth, essentially behaving more like a linear or low-degree polynomial classifier.
End of Section
1.2.5.7 - Support Vector Regression
👉Imagine a ‘tube’ of radius \(\epsilon\) surrounding the regression line.
- Points inside the tube are considered ‘correct’ and incur zero penalty.
- Points outside the tube are penalized based on their distance from the tube’s boundary.
👉SVR ignores errors as long as they are within a certain distance (\(\epsilon\)) from the true value.
🎯This makes SVR inherently robust to noise and outliers, as it does not try to fit every single point perfectly, only those that ‘matter’ to the structure of the data.
Note: Standard regression (like OLS) tries to minimize the squared error between the prediction and every data point.
Subject to constraints:
- \(y_i - (w^T x_i + w_0) \leq \epsilon + \xi_i\): (Upper boundary)
- \((w^T x_i + w_0) - y_i \leq \epsilon + \xi_i^*\): (Lower boundary)
- \(\xi_i, \xi_i^* \geq 0\): (Slack/Error cannot be negative)
Terms:
- Epsilon(\(\epsilon\)): The width of the tube. Increasing ‘\(\epsilon\)’ results in fewer support vectors and a smoother (flatter) model.
- Slack Variables (\(\xi_i, \xi_i^*\)): How far a point lies outside the upper and lower boundaries of the tube.
- C: The trade-off between the flatness of the model and the extent to which deviations larger than \(\epsilon\) are tolerated.
SVR uses a specific loss function that is 0 when the error<’\(\epsilon\)’.
\[L_\epsilon(y, f(x)) = \max(0, |y - f(x)| - \epsilon)\]- The solution becomes sparse, because the loss is zero for points inside the tube.
- Only the Support Vectors, i.e, points outside or on the boundary of the tube have non-zero Lagrange multipliers (\(\alpha_i\)).
Note: \(\epsilon=0.1\) default value in scikit-learn.
Subject to:
- \(\sum_{i=1}^n (\alpha_i - \alpha_i^*) = 0\)
- \(0 \leq \alpha_i, \alpha_i^* \leq C\)
- \(\alpha_i = \alpha_i^* = 0\): point is inside the tube.
- \(|\alpha_i - \alpha_i^*| > 0\) : support vectors; points on or outside the tube.
Note: \(\alpha_i , \alpha_i^* \) cannot both be non-zero for the same point; a point cannot be simultaneously above and below the tube.
- 👉 For non-linear SVR we replace dot product \(x_i^T x_j\) with kernel function \(K(x_i, x_j)\).
- ✅ Model needs to store only support vectors, i.e, points where \(|\alpha_i - \alpha_i^*| > 0\).
- ⭐️\(\xi_i =0 \) for a point that lies exactly on the boundary, so we can use that to calculate the bias (\(w_0\)):
End of Section
1.2.6.1 - Naive Bayes Intro
📌Let’s understand Naive Bayes through an Email/Text classification example.
- Number of words in an email is not fixed.
- Remove all stop words, such as, the, is , are, if, etc.
- Keep only relevant words, i.e, \(w_1, w_2, \dots , w_d\) words.
- We want to do a binary classification - Spam/Not Spam.
Let’s revise Bayes’ theorem first:
- \(P(S|W)\) is the posterior probability: the probability of the email being spam, given the words inside it.
- \(P(W|S)\) is the likelihood: how likely is this email’s word pattern if it were spam?
- \(P(S)\) is the prior probability: The ‘base rate’ of spam.
- If our dataset has 10,000 emails and 2,000 are spam, then \(P(S)\)=0.2.
- \(P(W)\) is the prior probability of the predictor (evidence): total probability of seeing these words across all emails.
- 👉Since this is the same for both classes, we treat it as a constant and ignore it during comparison.
👉Likelihood = \(P(W|S)\) = \(P(w_1, w_2, \dots w_d | S)\)
➡️ For computing the joint distribution of say d=1000 words, we need to learn from possible \(2^{1000}\) combinations.
- \(2^{1000}\) > the atoms in the observable 🔭 universe 🌌.
- We will never have enough training data to see every possible combination of words even once.
- Most combinations would have a count of zero.
🦉So, how do we solve this ?
💡The ‘Naive’ assumption is a ‘Conditional Independence’ assumption, i.e, we assume each word appears independently of the others, given the class Spam/Not Spam.
- e.g. In a spam email, the likelihood of ‘Free’ and ‘Money’ 💵 appearing are treated as independent events, even though they usually appear together.
Note: The conditional independence assumption makes the probability calculations easier, i.e, the joint probability simply becomes the product of individual probabilities, conditional on the label.
\[P(W|S) = P(w_1|S)\times P(w_2|S)\times \dots P(w_d|S) \]\[\implies P(S|W)=\frac{P(w_1|S)\times P(w_2|S)\times \dots P(w_d|S)\times P(S)}{P(W)}\]\[\implies P(S|W)=\frac{\prod_{i=1}^d P(w_i|S)\times P(S)}{P(W)}\]\[\text{Similarly, } P(NS|W)=\frac{\prod_{i=1}^d P(w_i|NS)\times P(NS)}{P(W)}\]We can generalize it for any number of class labels ‘y’:
Note: We compute the probabilities for both Spam/Not Spam and assign the final label to email, depending upon which probability is higher.
👉Space Complexity: O(d*c)
👉Time Complexity:
- Training: O(n*d*c)
- Inference: O(d*c)
Where,
- d = number of features/dimensions
- c = number of classes
- n = number of training examples
End of Section
1.2.6.2 - Naive Bayes Issues
⭐️Simple, fast, and highly effective probabilistic machine learning classifier based on Bayes’ theorem.
\[P(y|W) \propto \prod_{i=1}^d P(w_i|y)\times P(y)\]\[P(w_i|y) = \frac{count(w_i ~in~ y)}{\text{total words in class y}}\]🦀What if at runtime we encounter a word that was never seen during training ?
e.g. A word ‘crypto’ appears in the test email that was not present in training emails; P(‘crypto’|S) =0.
👉This will force the entire product to zero.
\[P(w_i|S) = \frac{\text{Total count of } w_i \text{ in all Spam emails}}{\text{Total count of all words in all Spam emails}}\]💡Add ‘Laplace smoothing’ to all likelihoods, both during training and test time, so that the probability becomes non-zero.
\[P(x_{i}|y)=\frac{count(x_{i},y)+\alpha }{count(y)+\alpha \cdot |V|}\]- \(count(x_{i},y)\) : number of times word appears in documents of class ‘y'.
- \(count(y)\): The total count of all words in documents of class ‘y'.
- \(|V|\)(or \(N_{features}\)):Vocabulary size or total number of unique possible words.
Let’s understand this by the examples below:
\[P(w_{i}|S)=\frac{count(w_{i},S)+\alpha }{count(S)+\alpha \cdot |V|}\]- \(count(w_{i},S) = 0 \), \(count(S) = 100\), \(|V|\)(or \(N_{features}) =2, \alpha = 1\) \[P(w_{i}|S)=\frac{ 0+1 }{100 +1 \cdot 2} = \frac{1}{102}\]
- \(count(w_{i},S) = 0 \), \(count(S) = 100\), \(|V|\)(or \(N_{features}) =2, \alpha = 10,000\) \[P(w_{i}|S)=\frac{ 0+10,000 }{100 +10,000 \cdot 2} = \frac{10,000}{20,100} \approx \frac{1}{2}\]
Note: High alpha value may lead to under-fitting; \(\alpha = 1\) recommended.
🦀What happens if the number of words ‘d’ is very large ?
👉Multiplying 500 times will result in a number so small a computer 💻 cannot store it (underflow).
Note: Computers have a limit to store floating point numbers, e.g., 32 bit: \(1.175 x 10^{-38}\)
💡Take ‘Logarithm’ that will convert the product to sum.
\[P(y|W) \propto \prod_{i=1}^d P(w_i|y)\times P(y)\]\[\log(P(y| W)) \propto \sum_{i=1}^d \log(P(w_i|y)) + \log(P(y))\]Note: In the next section we will solve a problem covering all the concepts discussed in this section.
End of Section
1.2.6.3 - Naive Bayes Example
⭐️Simple, fast, and highly effective probabilistic machine learning classifier based on Bayes’ theorem.
\[\log(P(y| W)) \propto \sum_{i=1}^d \log(P(w_i|y)) + \log(P(y))\]\[P(w_{i}|y)=\frac{count(w_{i},y)+\alpha }{count(y)+\alpha \cdot |V|}\]Let’s solve an email classification problem, below we have list of emails (tokenized) and labelled as Spam/Not Spam for training.
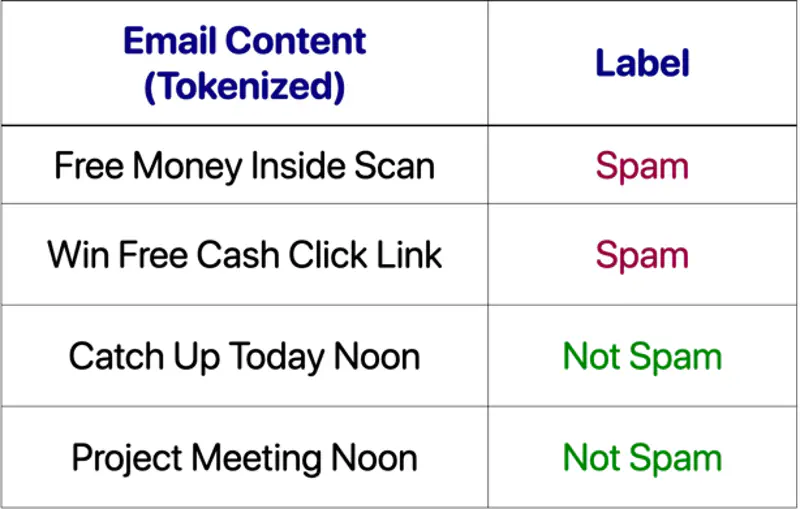
🏛️Class Priors:
- P(Spam) = 2/4 =0.5
- P(Not Spam) = 2/4 = 0.5
📕 Vocabulary = { Free, Money, Inside, Scan, Win, Cash, Click, Link, Catch, Up Today, Noon, Project, Meeting }
- |V| = Total unique word count = 14
🧮 Class count:
- count(Spam) = 9
- count(Not Spam) = 7
✅ Laplace smoothing: \(\alpha = 1\)
- P(‘free’| Spam) = \(\frac{2+1}{9+14} = \frac{3}{23} = 0.13\)
- P(‘free’| Not Spam) = \(\frac{0+1}{7+14} = \frac{1}{21} = 0.047\)
👉Say a new email 📧 arrives - “Free money today”; lets classify it as Spam/Not Spam.
Spam:
- P(‘free’| Spam) = \(\frac{2+1}{9+14} = \frac{3}{23} = 0.13\)
- P(‘money’| Spam) = \(\frac{1+1}{9+14} = \frac{2}{23} = 0.087\)
- P(‘today’| Spam) = \(\frac{0+1}{9+14} = \frac{1}{23} = 0.043\)
Not Spam:
- P(‘free’| Not Spam) = \(\frac{0+1}{7+14} = \frac{1}{21} = 0.047\)
- P(‘money’| Not Spam) = \(\frac{0+1}{7+14} = \frac{1}{21} = 0.047\)
- P(‘today’| Not Spam) = \(\frac{1+1}{7+14} = \frac{2}{21} = 0.095\)
- Score(Spam) = log(P(Spam)) + log(P(‘free’|S)) + log(P(‘money’|S)) + log(P(‘today’|S))
= log(0.5) + log(0.13) + log(0.087) + log(0.043) = -0.301 -0.886 -1.06 -1.366 = -3.614 - Score(Not Spam) = log(P(Not Spam)) + log(P(‘free’|NS)) + log(P(‘money’|NS)) + log(P(‘today’|NS))
= log(0.5) + log(0.047) + log(0.047) + log(0.095) = -0.301 -1.328 -1.328 -1.022 = -3.979
👉Since, Score(Spam) (-3.614 )> Score(Not Spam) (-3.979) , the model chooses ‘Spam’ as the label for the email.
End of Section
1.3 - Unsupervised Learning
End of Section
1.3.1.1 - K Means
🌍In real-world systems, labeled data is scarce and expensive 💰.
💡Unsupervised learning discovers inherent structure without human annotation.
👉Clustering answers: “Given a set of points, what natural groupings exist?”
- Customer Segmentation: Group users by behavior without predefined categories.
- Image Compression: Reduce color palette by clustering pixel colors.
- Anomaly Detection: Points far from any cluster are outliers.
- Data Exploration: Understand structure before building supervised models.
- Preprocessing: Create features from cluster assignments.
💡Clustering assumes that ‘similar’ points should be grouped together.
👉But what is ‘similar’? This assumption drives everything.
Given:
- Dataset X = {x₁, x₂, …, xₙ} where xᵢ ∈ ℝᵈ.
- Desired number of clusters ‘k'.
Find:
Cluster assignments C = {C₁, C₂, …, Cₖ}.
Such that points within clusters are ‘similar’.
And points across clusters are ‘dissimilar’.
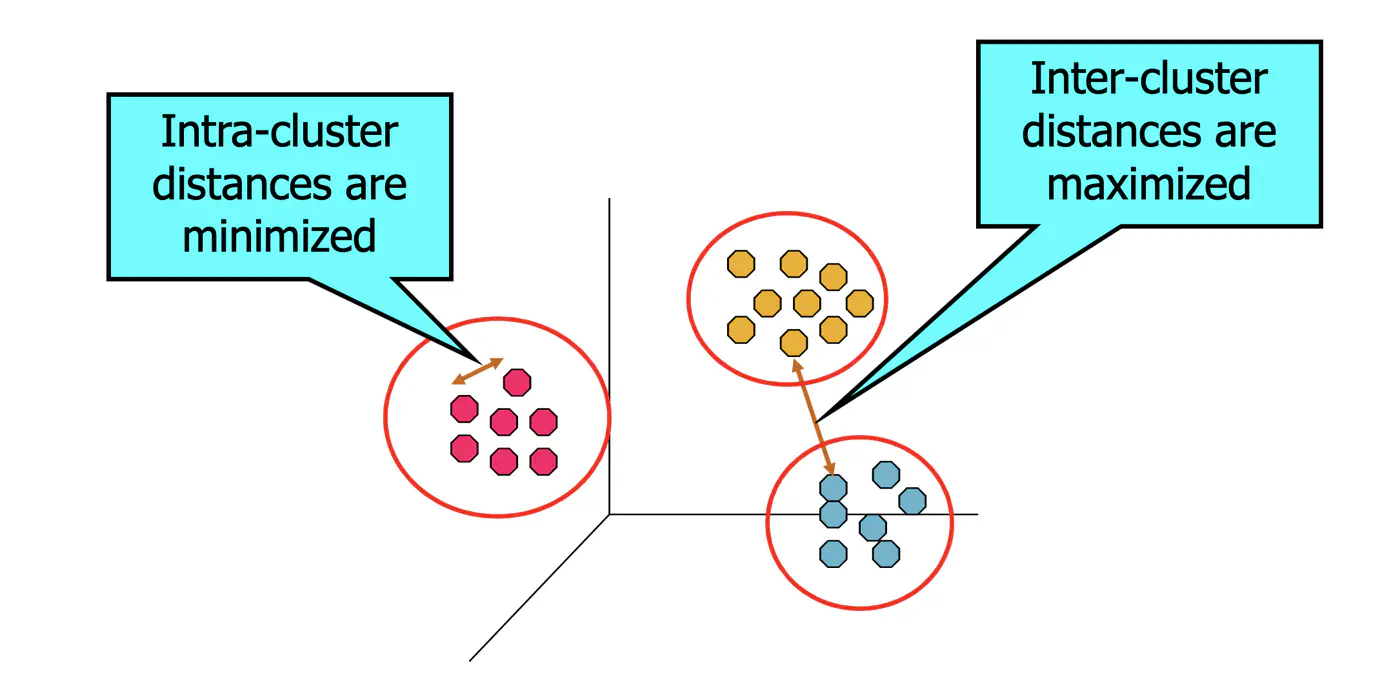
This is fundamentally an optimization problem, i.e, find parameters such that the value is minimum/maximum. We need:
- An objective function
- what makes a clustering ‘good’?
- An algorithm to optimize it
- how do we find good clusters?
- Evaluation metrics
- how do we measure quality?
Objective function:
👉Minimize the within-cluster sum of squares (WCSS).
- Where:
- C = {C₁, …, Cₖ} are cluster assignments.
- μⱼ is the centroid (mean) of cluster Cₖ.
- ||·||² is squared Euclidean distance.
Note: Every point belongs to one and only one cluster.
💡Within-Cluster Sum of Squares (WCSS) is nothing but variance.
⭐️ Total Variance = Within-Cluster Variance + Between-Cluster Variance
👉K-Means minimizes within-Cluster variance, which implicitly maximizes between-cluster separation.
Geometric Interpretation:
- Each point is ‘pulled’ toward its cluster center.
- The objective measures total squared distance of all points to their centers.
- Lower J(C, μ) means tighter, more compact clusters.
Note: K-Means works best when clusters are roughly spherical, similarly sized, and well-separated.
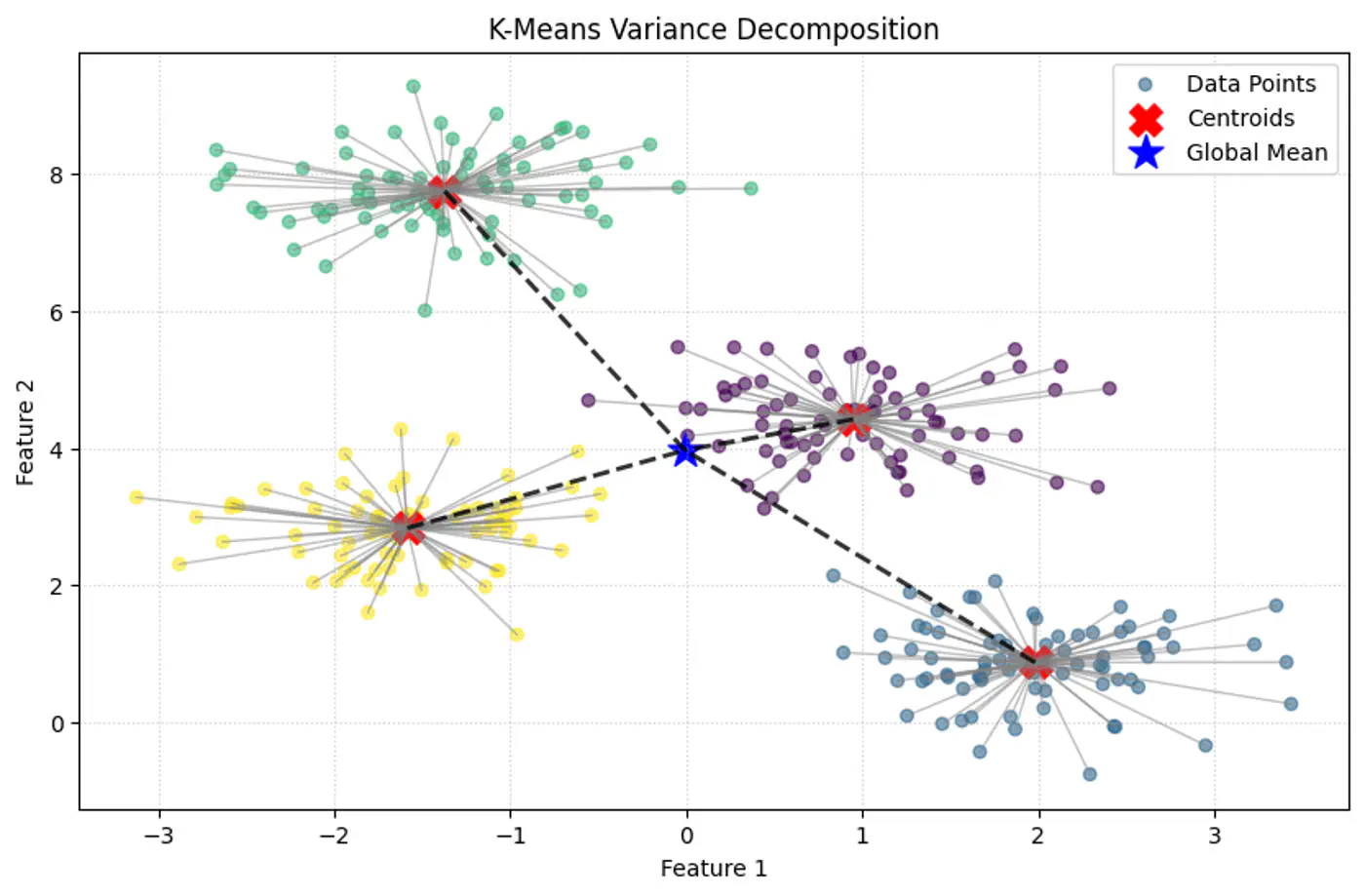
⭐️The problem requires partitioning ’n’ observations into ‘k’ distinct, non-overlapping clusters, which is given by the Stirling number of the second kind, which grows at a rate roughly equal to \(k^n/k!\).
\[S(n,k)=\frac{1}{k!}\sum _{j=0}^{k}(-1)^{k-j}{k \choose j}j^{n}\]\[S(100,2)=2^{100-1}-1=2^{99}-1\]\[2^{99}\approx 6.338\times 10^{29}\]👉This large number of possible combinations makes the problem NP-Hard.
🦉The k-means optimization problem is NP-hard because it belongs to a class of problems for which no efficient (polynomial-time ⏰) algorithm is known to exist.
End of Section
1.3.1.2 - Lloyds Algorithm
Iterative method for partitioning ’n’ data points into ‘k’ groups by repeatedly assigning data points to the nearest centroid (mean) and then recalculating centroids until assignments stabilize, aiming to minimize within-cluster variance.
📥Input: X = {x₁, …, xₙ}, ‘k’ (number of clusters)
📤Output: ‘C’ (clusters), ‘μ’ (centroids)
👉Steps:
- Initialize: Randomly choose ‘k’ cluster centroids μ₁, …, μₖ.
- Repeat until convergence, i.e, until cluster assignments and centroids no longer change significantly.
- a) Assignment: Assign each data point to the cluster whose centroid is closest (usually using Euclidean distance).
- For each point xᵢ: cᵢ = argminⱼ ||xᵢ - μⱼ||²
- b) Update: Recalculate the centroid (mean) of each cluster.
- For each cluster j: μⱼ = (1/|Cⱼ|) Σₓᵢ∈Cⱼ xᵢ
- Initialization sensitive, different initialization may lead to different clusters.
- Tries to make each cluster of same size that may not be the case in real world.
- Tries to make each cluster with same density(variance)
- Does not work well with non-globular(spherical) data.
👉See how 2 different runs of K-Means algorithm gives totally different clusters.
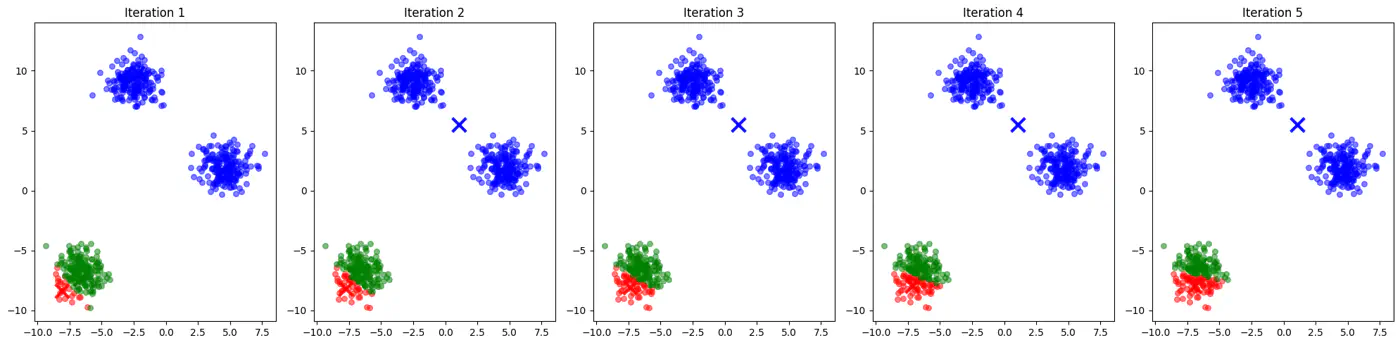
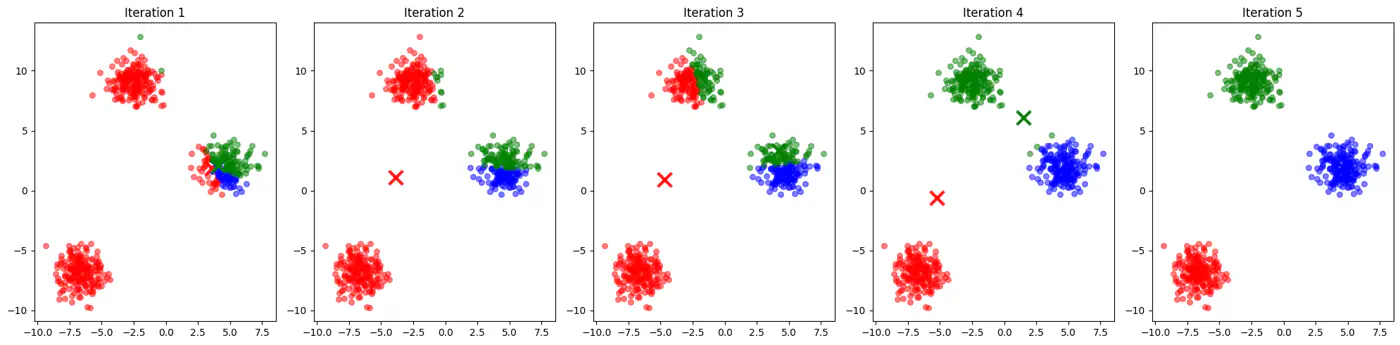
👉Also, K-Means does not work well with non-spherical clusters, or clusters with different densities and sizes.
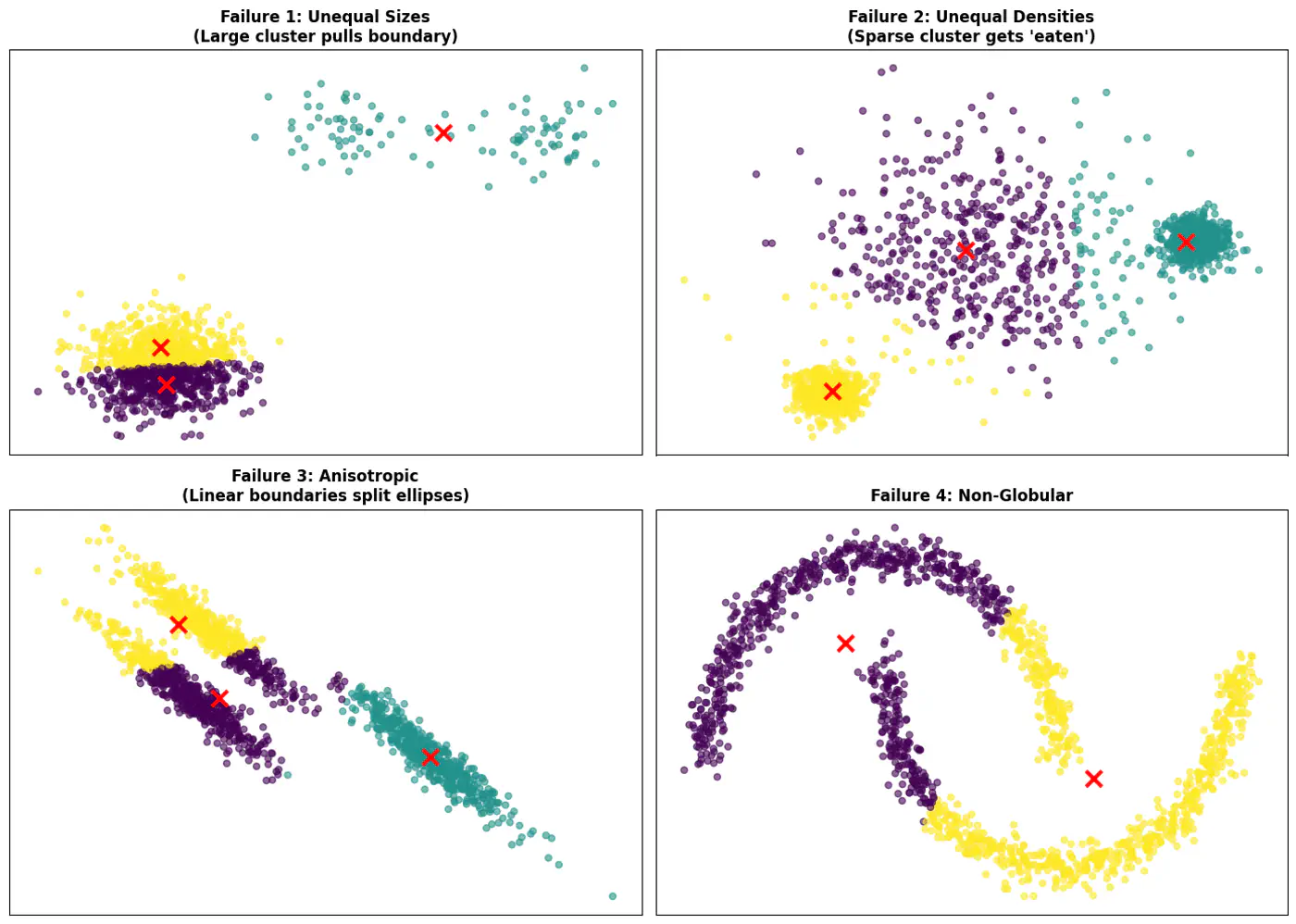
✅ Do not select initial points randomly, but some logic, such as, K-means++ algorithm.
✅ Use hierarchical clustering or density based clustering DBSCAN.
End of Section
1.3.1.3 - K Means++
- If two initial centroids belong to the same natural cluster, the algorithm will likely split that natural cluster in half and be forced to merge two other distinct clusters elsewhere to compensate.
- Inconsistent; different runs may lead to different clusters.
- Slow convergence; Centroids may need to travel much farther across the feature space, requiring more iterations.
👉Example for different K-Means algorithm runs give different clusters
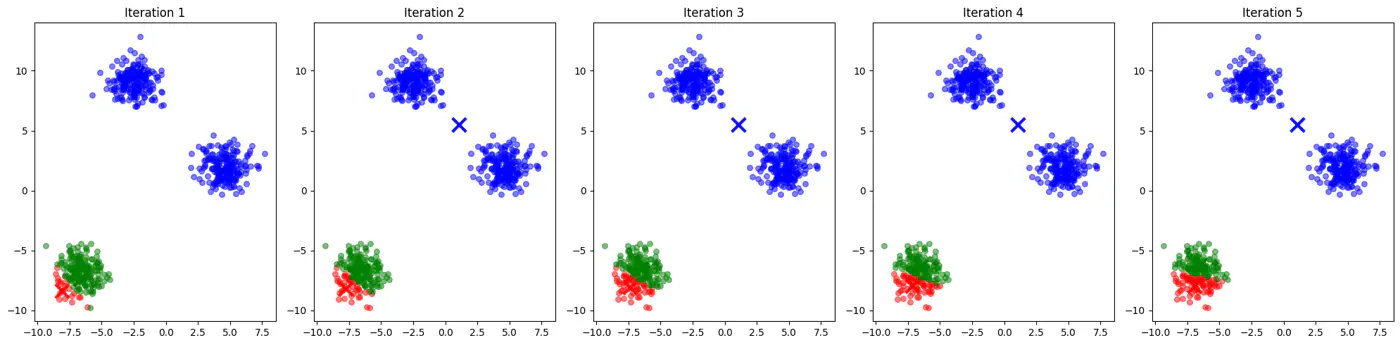
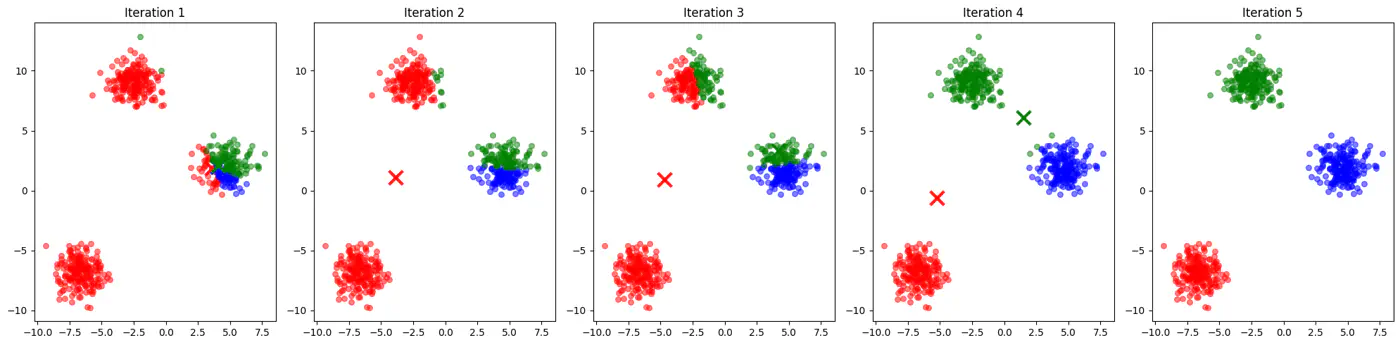
💡Addresses the issue due to random initialization by aiming to spread out the initial centroids across the data points.
Steps:
- Select the first centroid: Choose one data point randomly from the dataset to be the first centroid.
- Calculate distances: For every data point x not yet selected as a centroid, calculate the distance, D(x), between x and the nearest centroid chosen so far.
- Select the next centroid: Choose the next centroid from the remaining data points with a probability
proportional to D(x)^2.
This makes it more likely that a point far from existing centroids is selected, ensuring the initial centroids are well-dispersed. - Repeat: Repeat steps 2 and 3 until ‘k’ number of centroids are selected.
- Run standard K-means: Once the initial centroids are chosen, the standard k-means algorithm proceeds with assigning data points to the nearest centroid and iteratively updating the centroids until convergence.
End of Section
1.3.1.4 - K Medoid
- In K-Means, the centroid is the arithmetic mean of the cluster. The mean is very sensitive to outliers.
- Not interpretable; centroid is the mean of cluster data points and may not be an actual data point, hence not representative.
⭐️Medoid is a specific data point from a dataset that acts as the ‘center’ or most representative member of its cluster.
👉It is defined as the object within a cluster whose average dissimilarity (distance) to all other members in that same cluster is the smallest.
💡Selects actual data points from the dataset as cluster representatives, called medoids (most centrally located).
👉a.k.a Partitioning Around Medoids(PAM).
Steps:
- Initialization: Select ‘k’ data points from the dataset as the initial medoids using K-Means++ algorithm.
- Assignment: Calculate the distance (e.g., Euclidean or Manhattan) from each non-medoid point to all medoids and assign each point to the cluster of its nearest medoid.
- Update (Swap):
- For each cluster, swap current medoid with a non-medoid point from the same cluster.
- For each swap, calculate the total cost 💰(sum of distances from medoid).
- Pick the medoid with minimum cost 💰.
- Repeat🔁: Repeat the assignment and update steps until (convergence), i.e, medoids no longer change or a maximum number of iterations is reached.
Note: Kind of brute-force algorithm, computationally expensive for large dataset.
✅ Flexible Distance Metrics: It can work with any dissimilarity measure (Manhattan, Cosine similarity), making it ideal for categorical or non-Euclidean data.
✅ Interpretable Results: Cluster centers are real observations (e.g., a ‘typical’ customer profile), which makes the output easier to explain to stakeholders.
End of Section
1.3.1.5 - Clustering Quality Metrics
- 👉 Elbow Method: Quickest to compute; good for initial EDA (Exploratory Data Analysis).
- 👉 Dunn Index: Focuses on the ‘gap’ between the closest clusters.
- 👉 Silhouette Score: Balances compactness and separation.
- 👉 Domain specific knowledge and system constraints.
⭐️Heuristic used to determine the optimal number of clusters (k) for clustering by visualizing how the quality of clustering improves as ‘k’ increases.
🎯The goal is to find a value of ‘k’ where adding more clusters provides a diminishing return in terms of variance reduction.
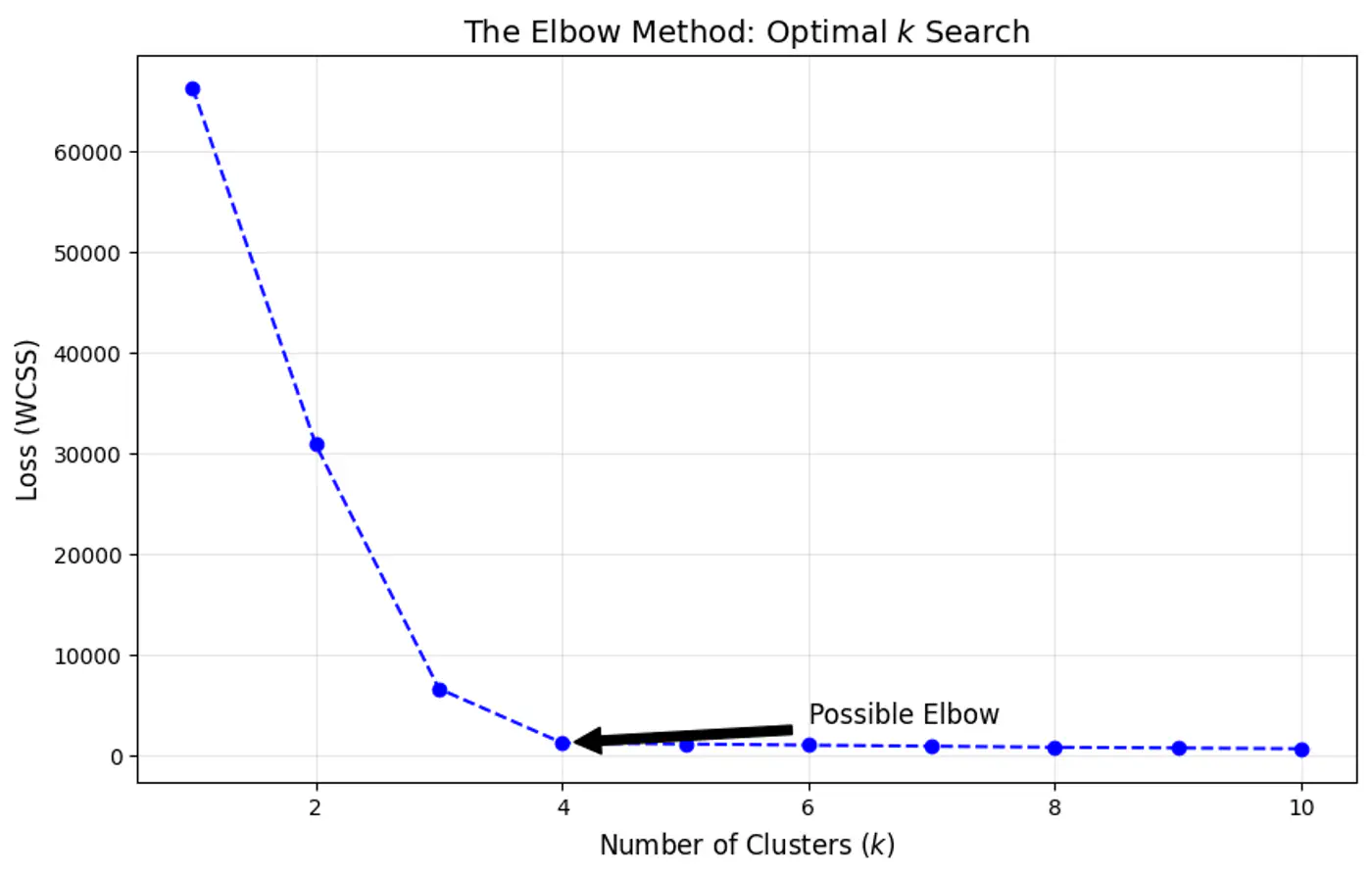
⭐️Clustering quality evaluation metric that measures: separation (between clusters) and compactness (within clusters)
Note: A higher Dunn Index value indicates better clustering, meaning clusters are well-separated from each other and compact.
👉Dunn Index Formula:
\[DI = \frac{\text{Minimum Inter-Cluster Distance(between different clusters)}}{\text{Maximum Intra-Cluster Distance(within a cluster)}}\]\[DI = \frac{\min_{1 \le i < j \le k} \delta(C_i, C_j)}{\max_{1 \le l \le k} \Delta(C_l)}\]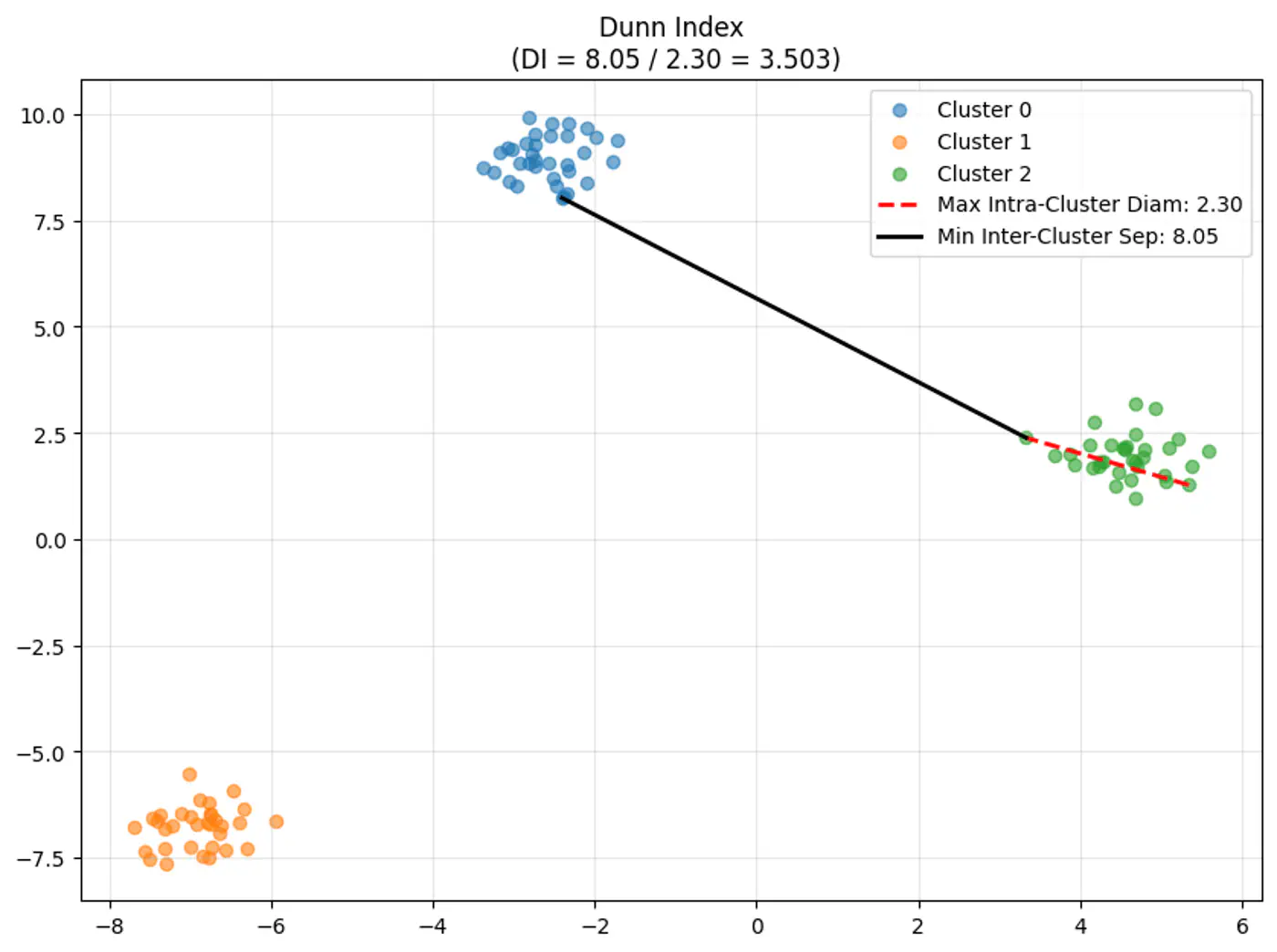
👉Let’s understand the terms in the above formula:
\(\delta(C_i, C_j)\) (Inter-Cluster Distance):
- Measures how ‘far apart’ the clusters are.
- Distance between the two closest points of different clusters (Single-Linkage distance). \[\delta(C_i, C_j) = \min_{x \in C_i, y \in C_j} d(x, y)\]
\(\Delta(C_l)\) (Intra-Cluster Diameter):
- Measures how ‘spread out’ a cluster is.
- Distance between the two furthest points within the same cluster (Complete-Linkage distance). \[\Delta(C_l) = \max_{x, y \in C_l} d(x, y)\]
- Single Linkage (MIN): Uses the minimum distance between any two points in different clusters.
- Complete Linkage (MAX): Uses the maximum distance between any two points in same cluster.
End of Section
1.3.1.6 - Silhouette Score
- ✅ Elbow Method: Quickest to compute; good for initial EDA.
- ✅ Dunn Index: Focuses on the ‘gap’ between the closest clusters.
——- We have seen the above 2 methods in the previous section ———- - 👉 Silhouette Score: Balances compactness and separation.
- 👉 Domain specific knowledge and system constraints.
⭐️Clustering quality evaluation metric that measures how similar a data point is to its own cluster (cohesion) compared to other clusters (separation).
Note: Higher scores (closer to 1) indicate better-defined, distinct clusters, while scores near 0 suggest overlapping clusters, and negative scores mean points might be in the wrong cluster.
Silhouette score for point ‘i’ is the difference between separation b(i) and cohesion a(i), normalized by the larger of the two.
\[ s(i) = \frac{b(i) - a(i)}{\max(a(i), b(i))} \]Note: The Global Silhouette Score is simply the mean of s(i) for all points in the dataset.
👉Example for Silhouette Score:
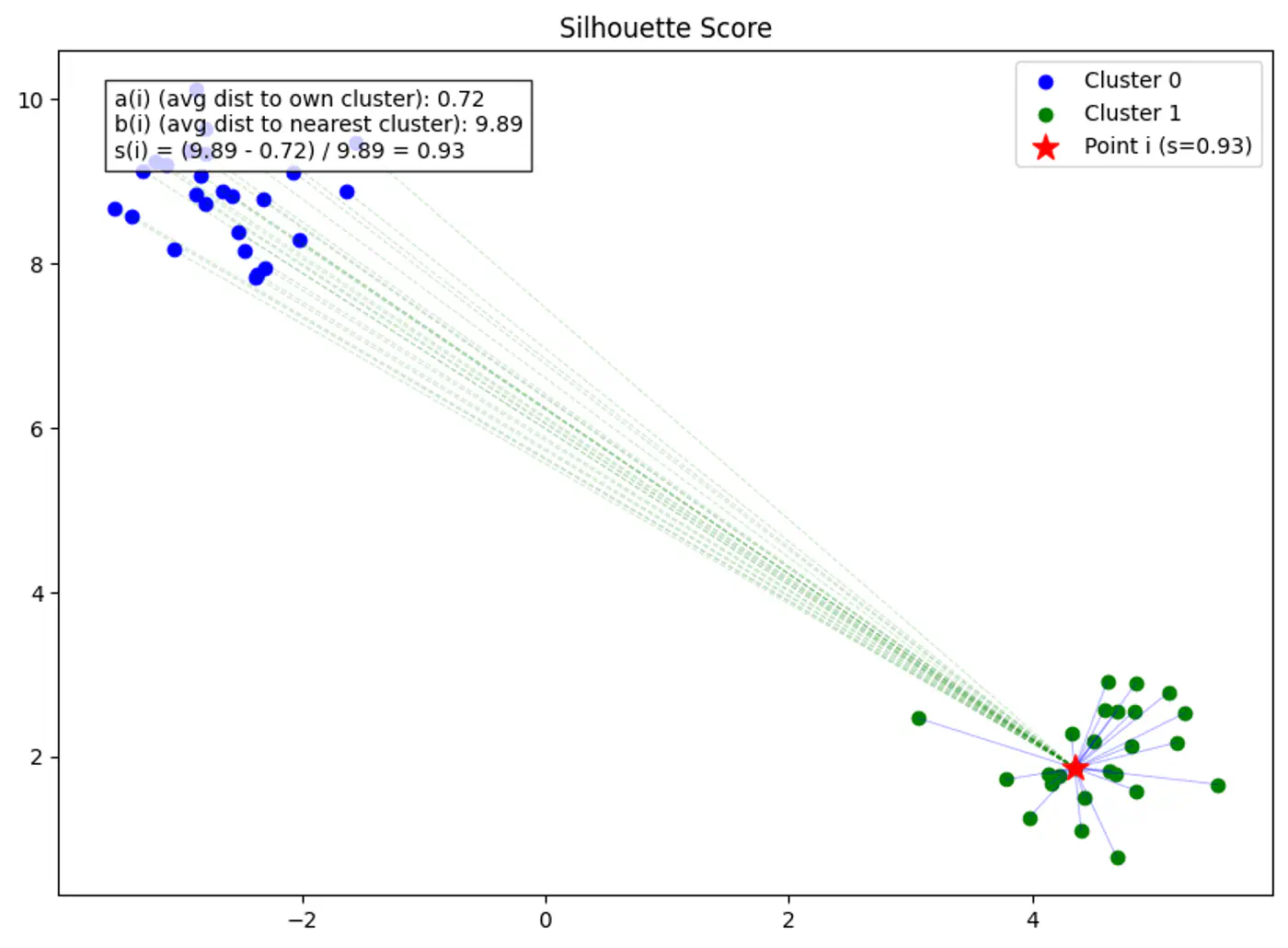
👉Example for Silhouette Score of 0(Border Point) and negative(Wrong Cluster).
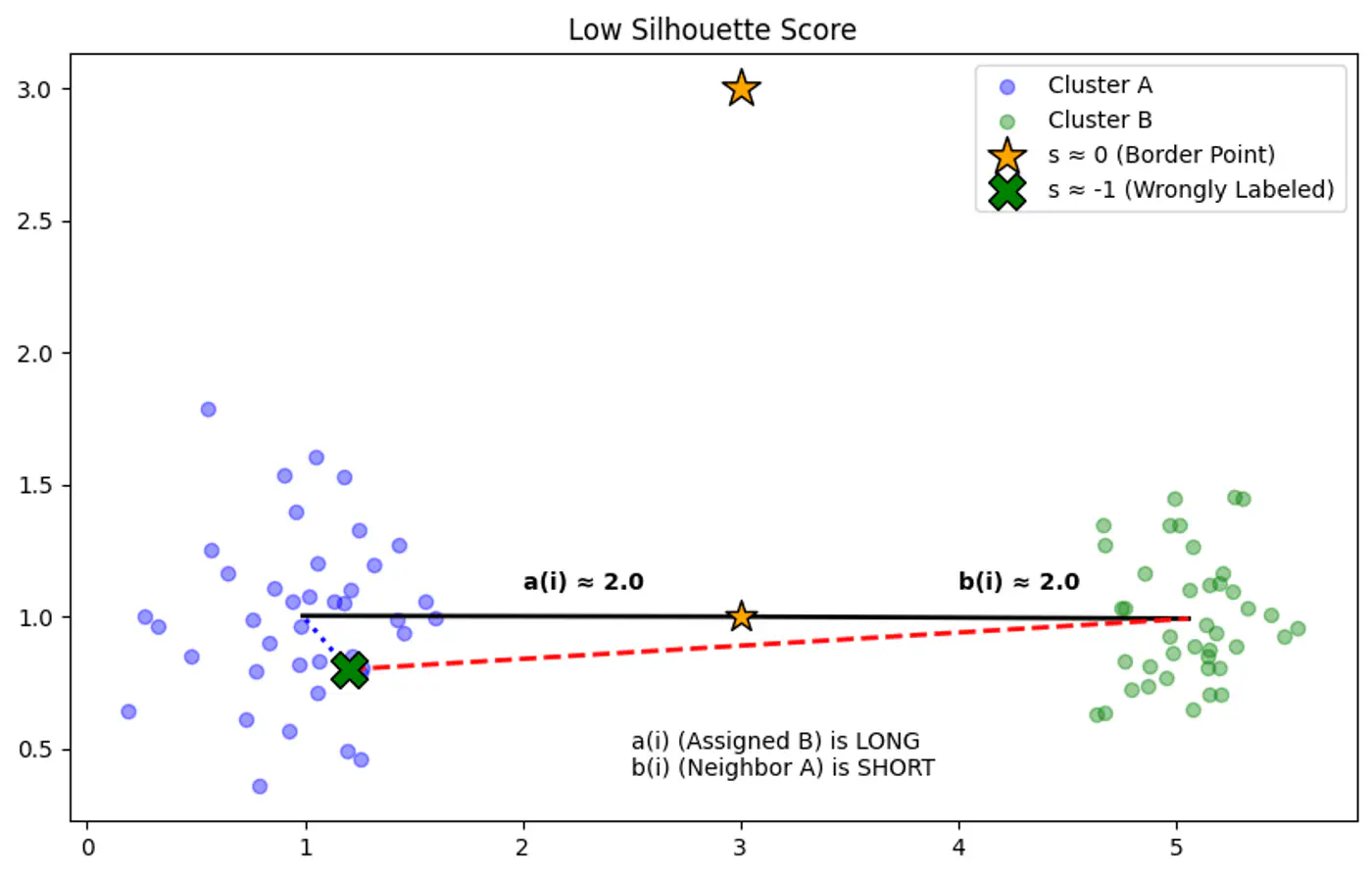
🦉Now let’s understand the terms in Silhouette Score in detail.
Average distance between point ‘i’ and all other points in the same cluster.
\[a(i) = \frac{1}{|C_A| - 1} \sum_{j \in C_A, i \neq j} d(i, j)\]Note: Lower a(i) means the point is well-matched to its own cluster.
Average distance between point ‘i’ and all points in the nearest neighboring cluster (the cluster that ‘i’ is not a part of, but is closest to).
\[b(i) = \min_{C_B \neq C_A} \frac{1}{|C_B|} \sum_{j \in C_B} d(i, j)\]Note: Higher b(i) means the point is very far from the next closest cluster.
⭐️A silhouette plot is a graphical tool used to evaluate the quality of clustering algorithms (like K-Means), showing how well each data point fits within its cluster.
👉Each bar gives the average silhouette score of the points assigned to that cluster.
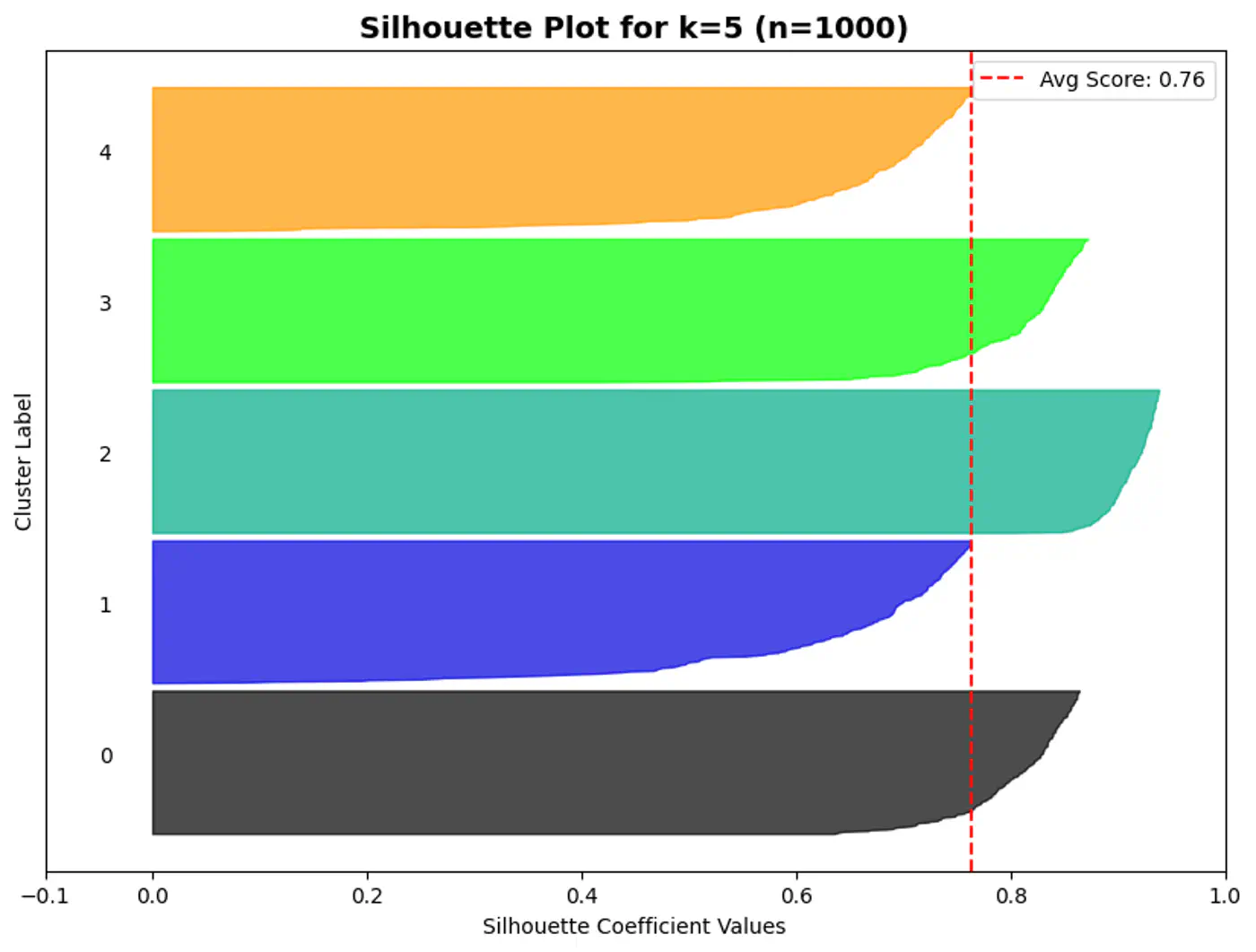
⛳️ Like K-Means, the Silhouette Score (when using Euclidean distance) assumes convex clusters.
🌘 If we use it on ‘Moon’ shaped clusters, it will give a low score even if the clusters are perfectly separated, because the ‘average distance’ to a neighbor might be small due to the curvature of the manifold.
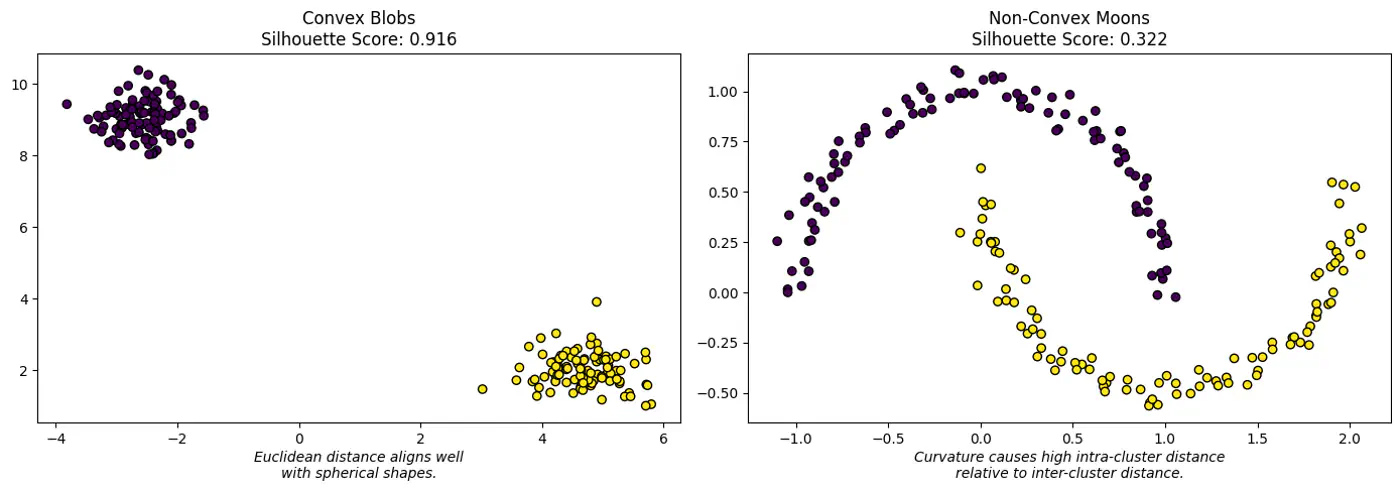
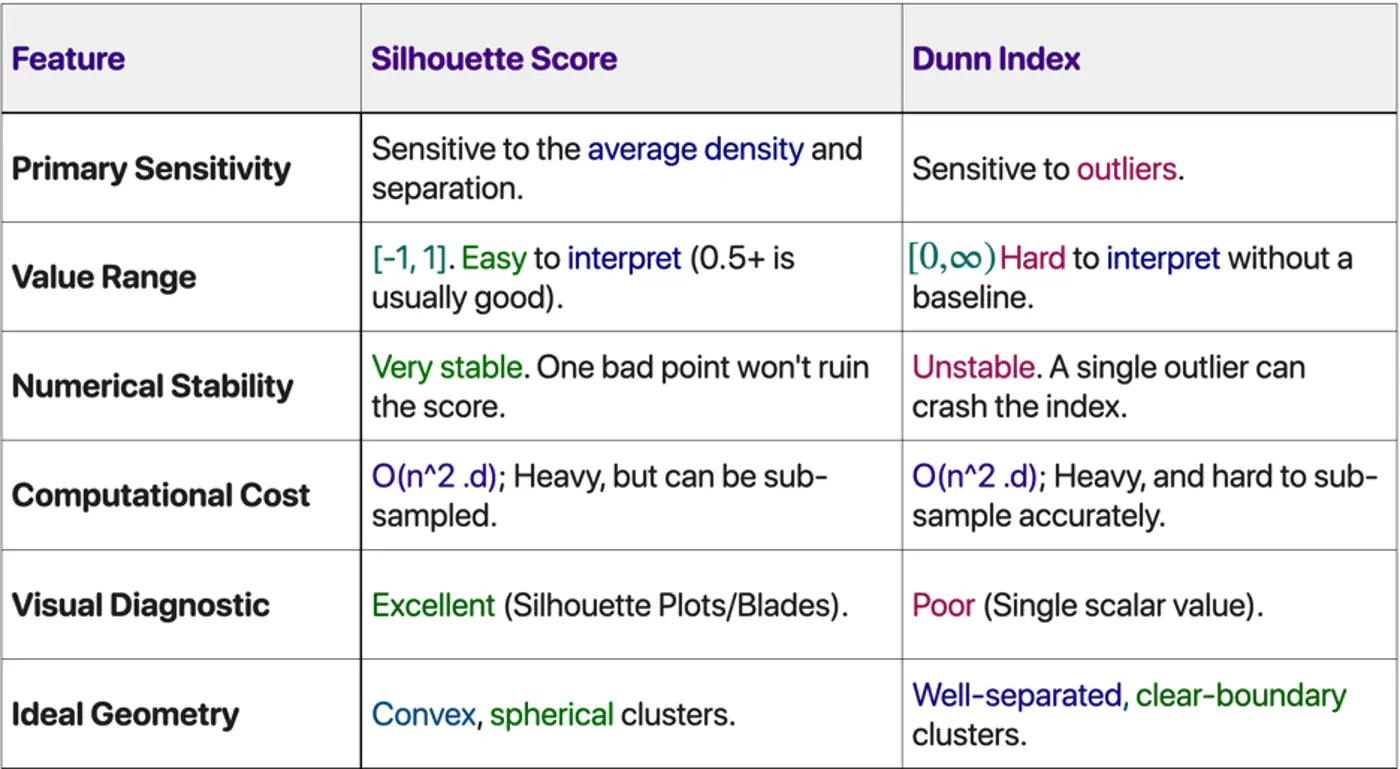
Choose Silhouette Score if:
✅ Need a human-interpretable metric to present to stakeholders.
✅ Dealing with real-world noise and overlapping ‘fuzzy’ boundaries.
✅ Want to see which specific clusters are weak (using the plot).
Choose Dunn Index if:
✅ Performing ‘Hard Clustering’ where separation is a safety or business requirement.
✅ Data is pre-cleaned of outliers (e.g., in a curated embedding space).
✅ Need to compare different clustering algorithms (e.g., K-Means vs. DBSCAN) on a high-integrity task.
End of Section
1.3.2 - Hierarchical Clustering
End of Section
1.3.2.1 - Hierarchical Clustering
- 🤷 We might not know in advance the number of distinct clusters ‘k’ in the dataset.
- 🕸️ Also, sometimes the dataset may contain a nested structure or some inherent hierarchy, such as, file system, organizational chart, biological lineages, etc.
⭐️ Method of cluster analysis that seeks to build a hierarchy of clusters, resulting in a tree like structure called dendrogram.
👉Hierarchical clustering allows us to explore different possibilities (of ‘k’) by cutting the dendrogram at various levels.
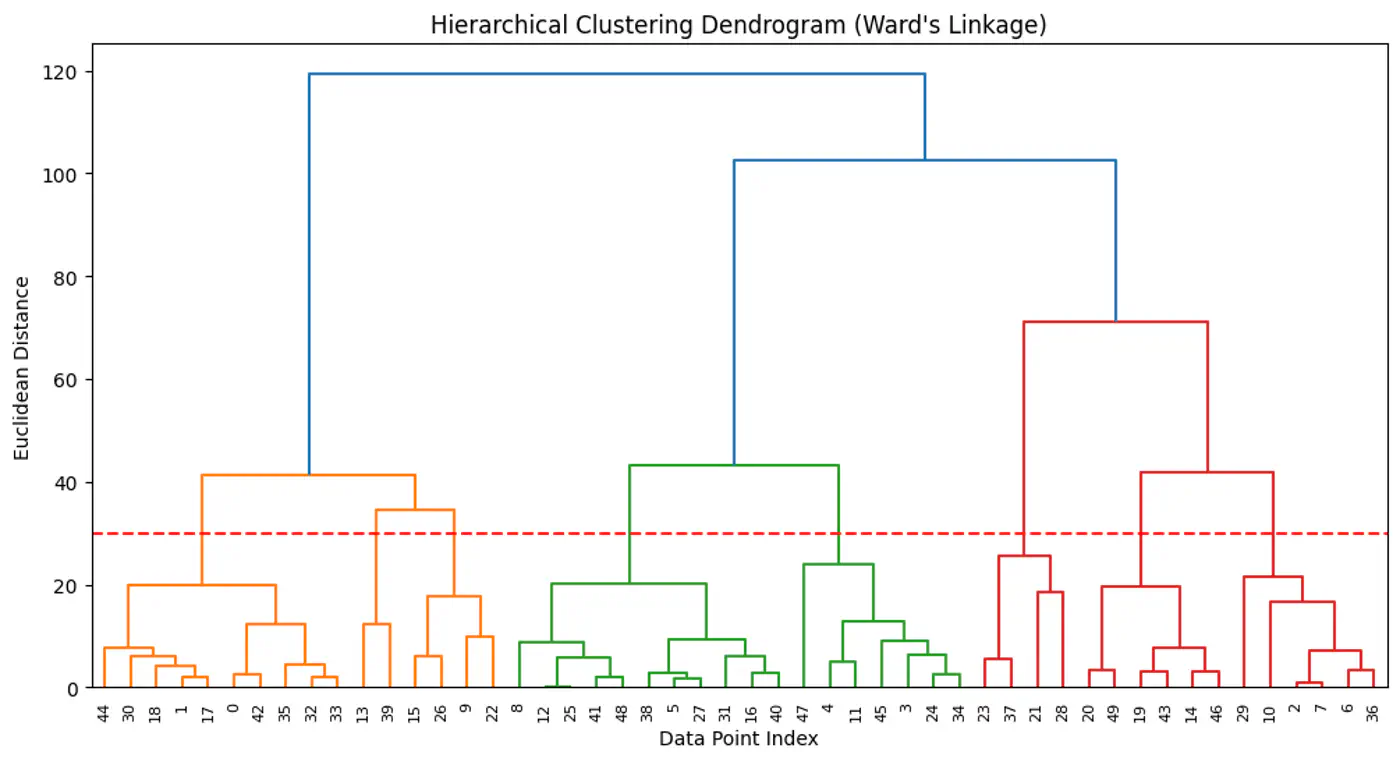
Agglomerative (Bottom-Up):
Most common, also known as Agglomerative Nesting (AgNes).
- Every data point starts as its own cluster.
- In each step, merge the two ‘closest’ clusters.
- Repeat step 2, until all points are merged in a single cluster.
Divisive (Top-Down):
- All data points start in one large cluster.
- In each step, divide the cluster into two halves.
- Repeat step 2, until every point is its own cluster.
Agglomerative Clustering Example:
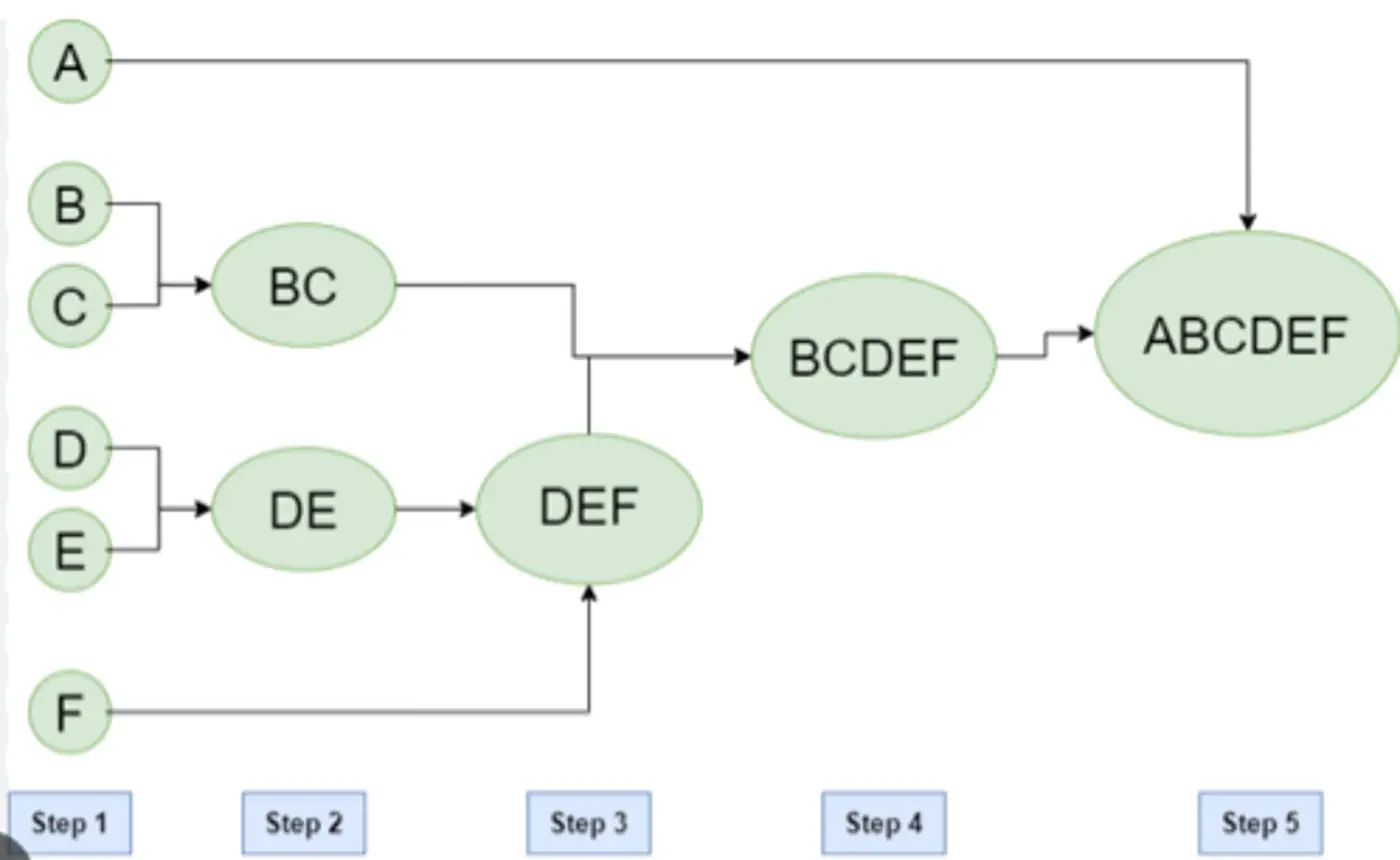
- Ward’s Method:
- Merges clusters to minimize the increase in the total within-cluster variance (sum of squared errors), resulting in compact, equally sized clusters.
- Single Linkage (MIN):
- Uses the minimum distance between any two points in different clusters.
- Prone to creating long, ‘chain-like’ 🔗 clusters, sensitive to outliers.
- Complete Linkage (MAX):
- Uses the maximum distance between any two points in different clusters.
- Forms tighter, more spherical clusters, less sensitive to outliers than single linkage.
- Average Linkage:
- Combines clusters by the average distance between all points in two clusters, offering a compromise between single and complete linkage.
- A good middle ground, often overcoming limitations of single and complete linkage.
- Centroid Method:
- Merges clusters based on the distance between their centroids (mean points).
👉Single Linkage is more sensitive to outlier than Complete Linkage, as Single Linkage can keep linking to the closest point forming a bridge to outlier.
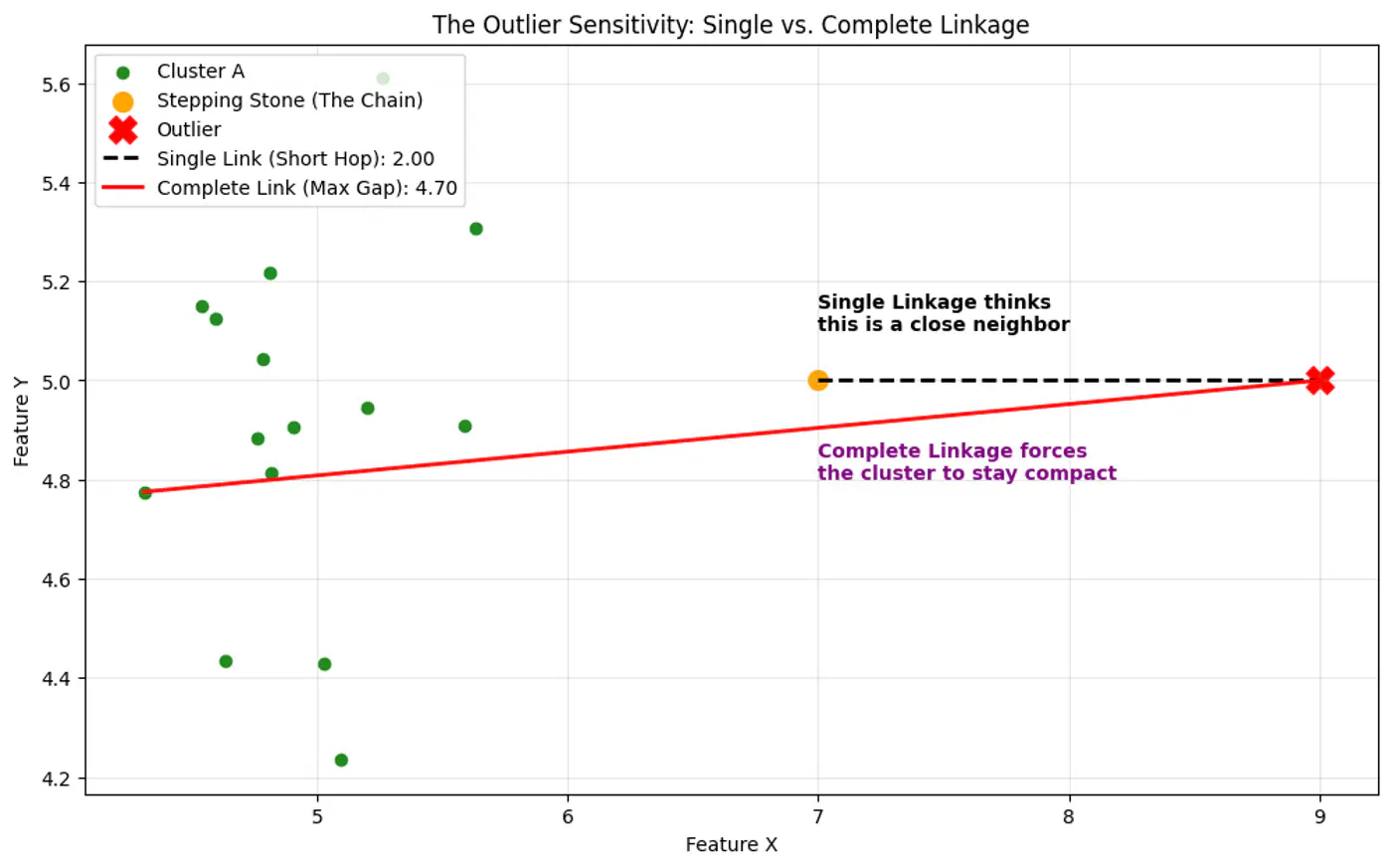
👉All cluster linkage distances.
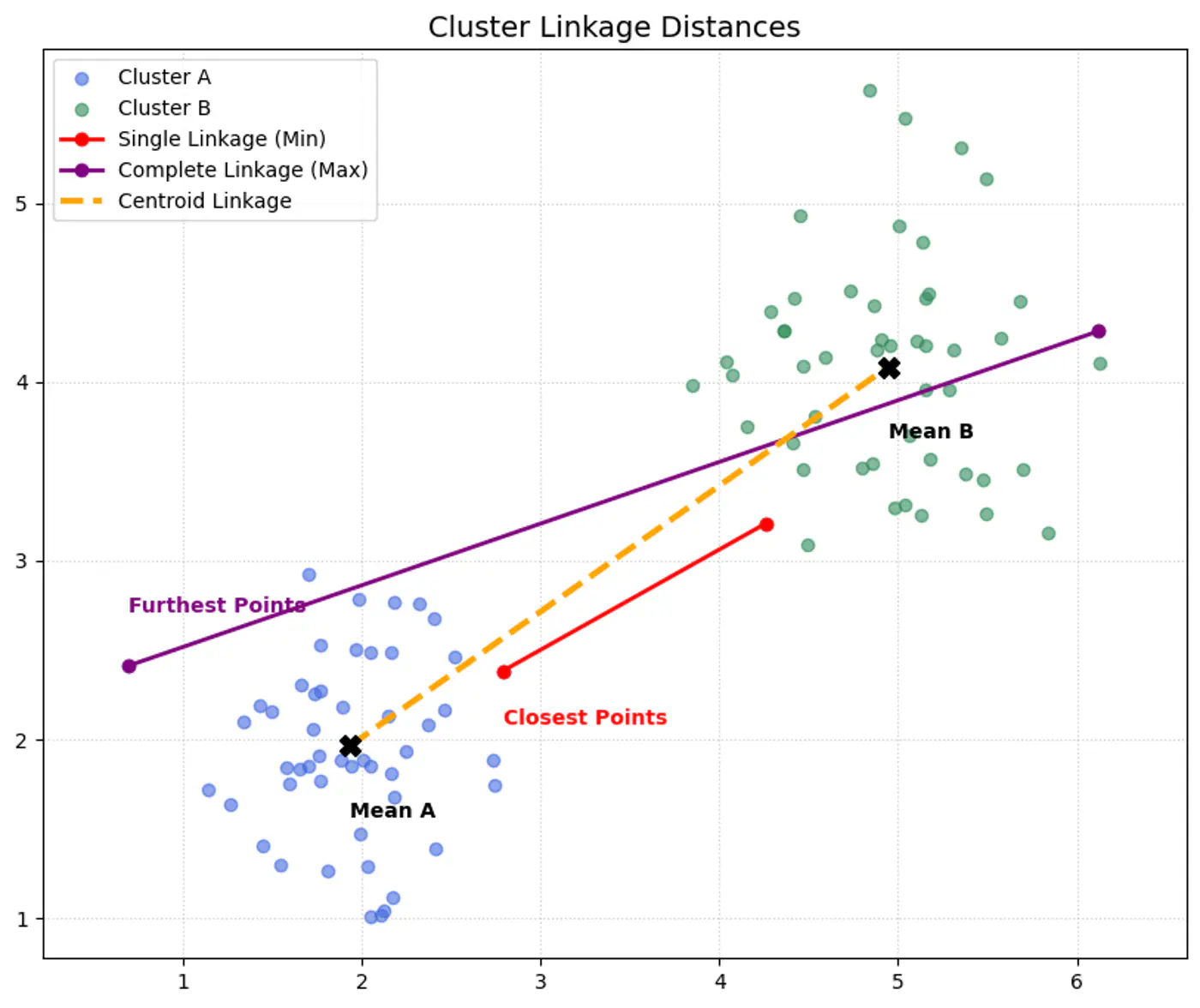
👉We get different clustering using different linkages.
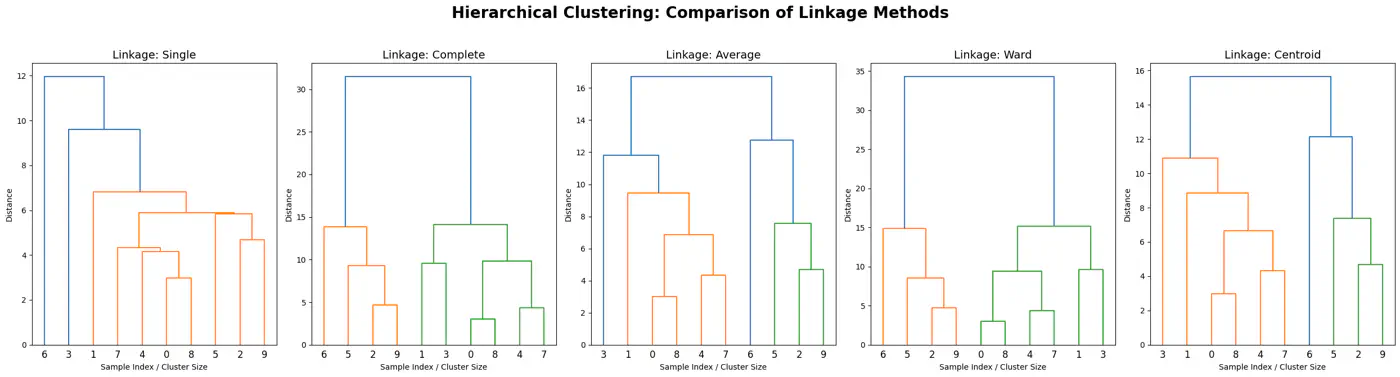
End of Section
1.3.3 - DBSCAN
End of Section
1.3.3.1 - DBSCAN
- Non-Convex Shapes: K-Means can not find ‘crescent’ or ‘ring’ shape clusters.
- Noise: K-Means forces every point into a cluster, even outliers.
👉K-Means asks:
- “Which center is closest to this point?”
👉DBSCAN asks:
- “Is this point part of a dense neighborhood?”
⭐️Groups closely packed data points into clusters based on their density, and marks points that lie alone in low-density regions as outliers or noise.
Note: Unlike K-means, DBSCAN can find arbitrarily shaped clusters and does not require the number of clusters to be specified beforehand.
- Epsilon (eps or \(\epsilon\)):
- Radius that defines the neighborhood around a data point.
- If it’s too small, many points will be noise, and if too large, distinct clusters may merge.
- Minimum Points(minPts or min_samples):
- Minimum number of data points required within a point’s -neighborhood for that point to be considered a dense region (a core point).
- Defines threshold for ‘density’.
- Rule of thumb: minPts dimensions + 1; use larger value for noisy data (minPts 2*dimensions).
- Core Point:
- If it has at least minPts (including itself) within its -neighborhood.
- Forms the dense backbone of the clusters and can expand them.
- Border Point:
- If it has at fewer than minPts within its -neighborhood but falls within the -neighborhood of a core point.
- Border points belong to a cluster but cannot expand it further.
- Noise Point (Outlier):
- If it is neither a core point nor a border point, i.e., it is not density-reachable from any core point.
- Not assigned to any cluster.
- Random Start:
- Mark all points as unvisited; pick an arbitrary unvisited point ‘P’ from the dataset.
- Density Check:
- Check the point P’s ϵ-neighborhood.
- If ‘P’ has at least minPts, it is identified as a core point, and a new cluster is started.
- If it has fewer than minPts, the point is temporarily labeled as noise (it might become a border point later).
- Cluster Expansion:
- Recursively visit all points in P’s ϵ-neighborhood.
- If they are also core points, their neighbors are added to the cluster.
- Iteration 🔁:
- This ‘density-reachable’ logic continues until the cluster is fully expanded.
- The algorithm then picks another unvisited point and repeats the process, discovering new clusters or marking more points as noise until all points are processed.
👉DBSCAN can correctly detect non-spherical clusters.
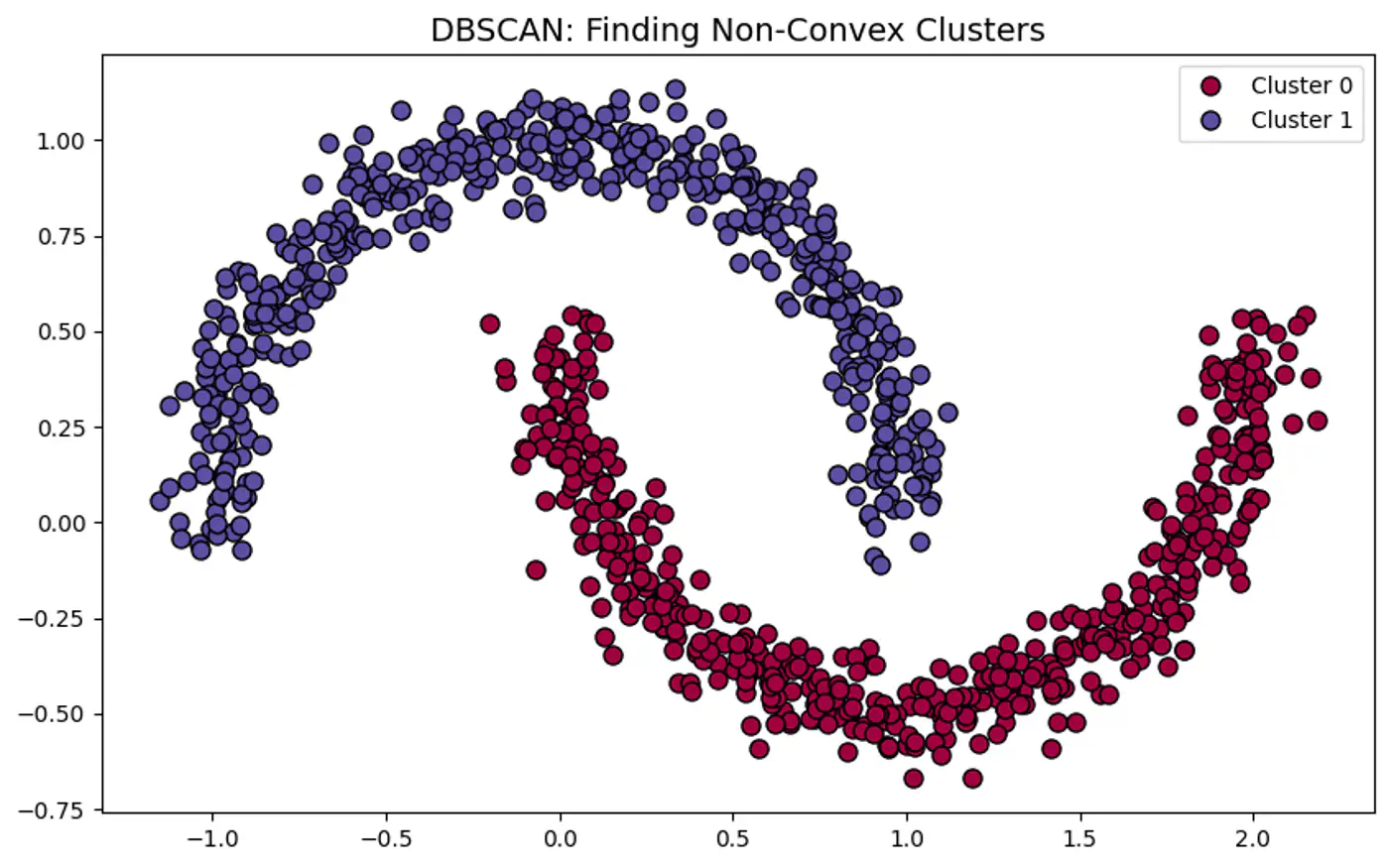
👉DBSCAN Points and Epsilon-Neighborhood.
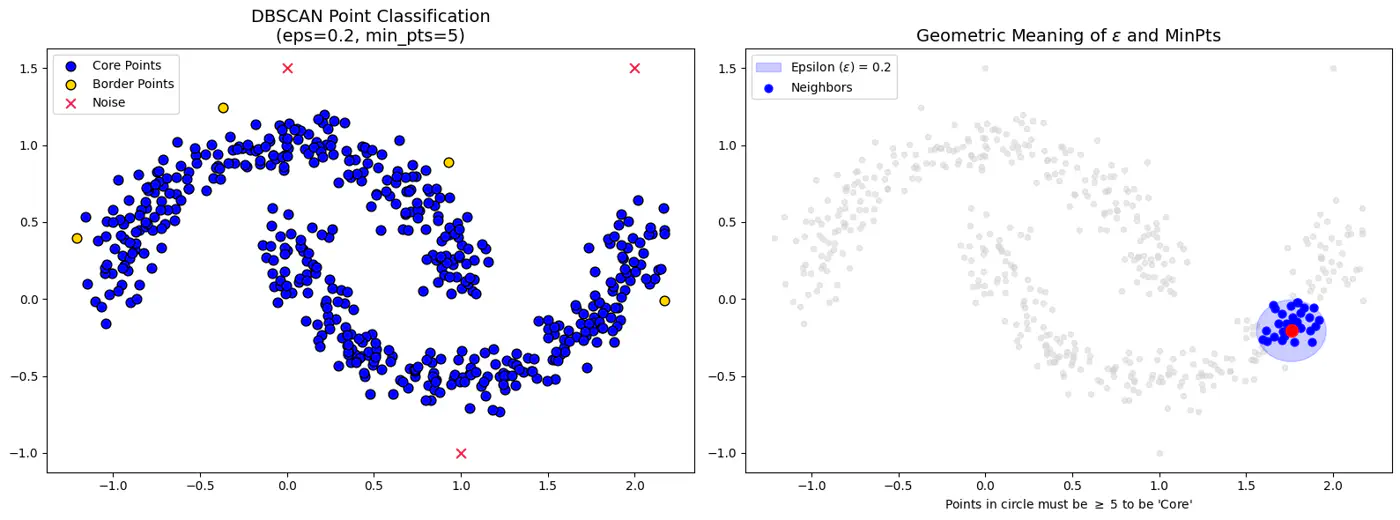
- Varying Density Clusters:
- Say A cluster is very dense and B is sparse, a single cannot satisfy both clusters.
- High Dimensional Data:
- ‘Curse of Dimensionality’ - In high-dimensional space, the distance between any two points converge.
Note: Sensitive to parameter eps and minPts; tricky to get it work.
👉DBSCAN Failure and Epsilon ((\epsilon)
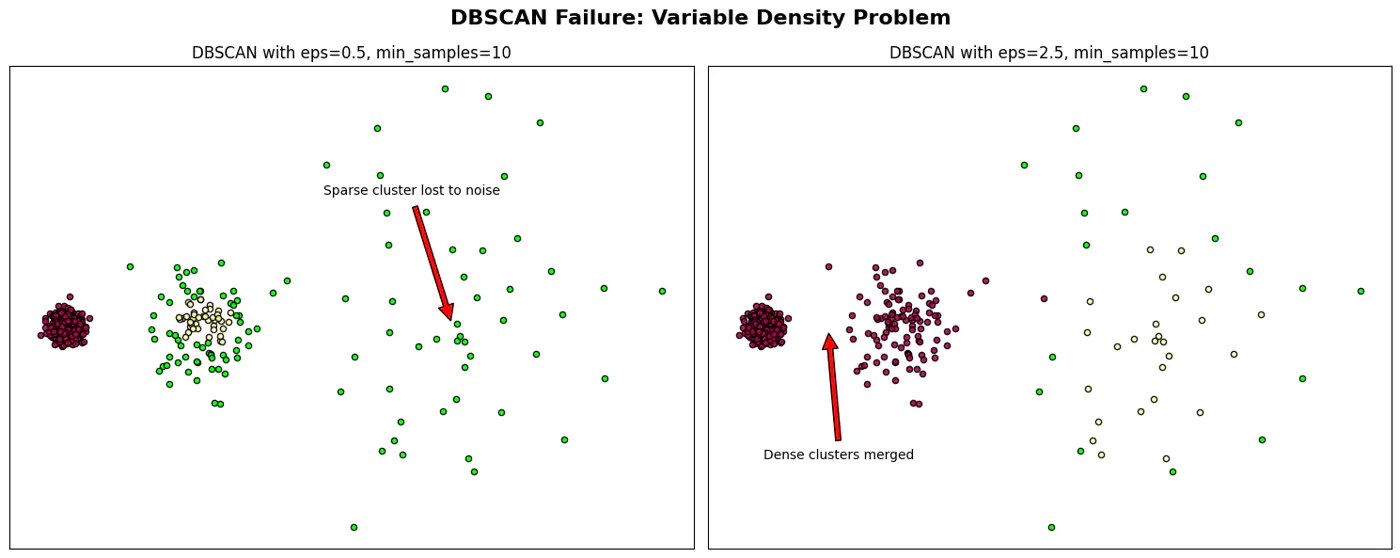
Arbitrary Cluster Shapes:
- When clusters are intertwined, nested, or ‘moon-shaped’; where K-Means would fail by splitting them.
Presence of Noise and Outliers:
- Robust to noise and outliers because it explicitly identifies low-density points as noise (labeled as -1) rather than forcing them into a cluster.
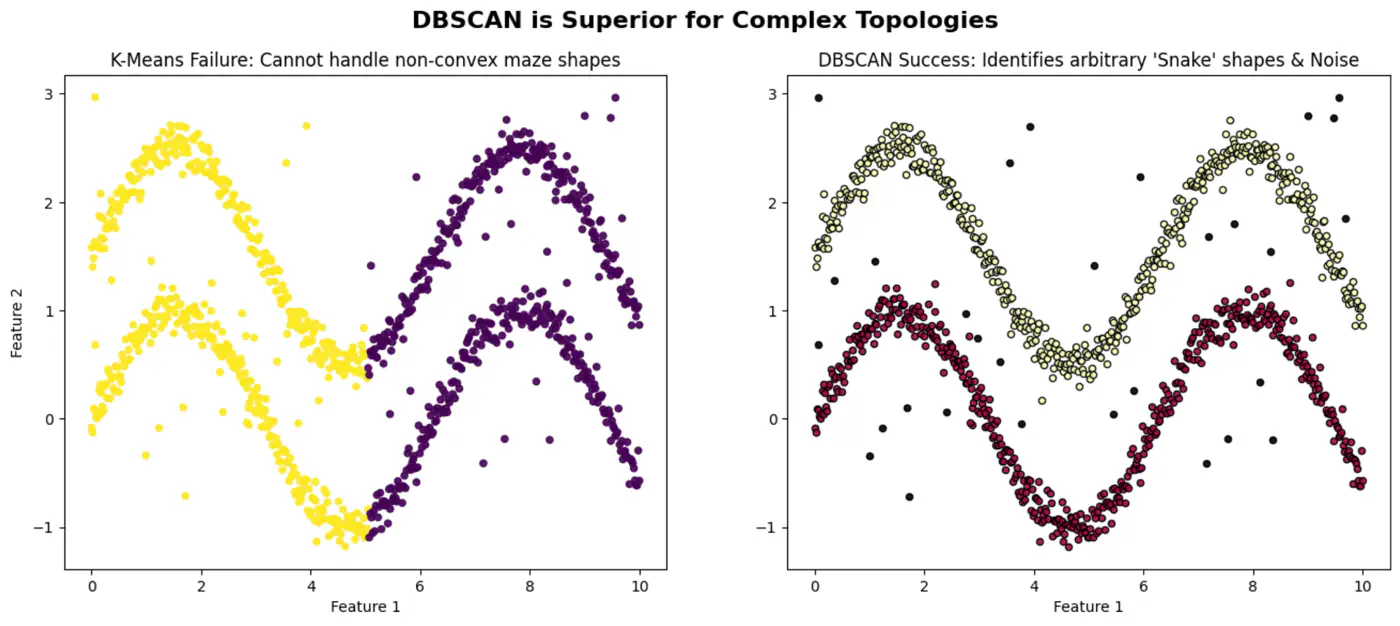
End of Section
1.3.4 - Gaussian Mixture Model
End of Section
1.3.4.1 - Gaussian Mixture Models
- K-means uses Euclidean distance and assumes that clusters are spherical (isotropic) and have the same variance across all dimensions.
- Places a circle or sphere around each centroid.
- What if the clusters are elliptical ? 🤔
👉K-Means Fails with Elliptical Clusters.
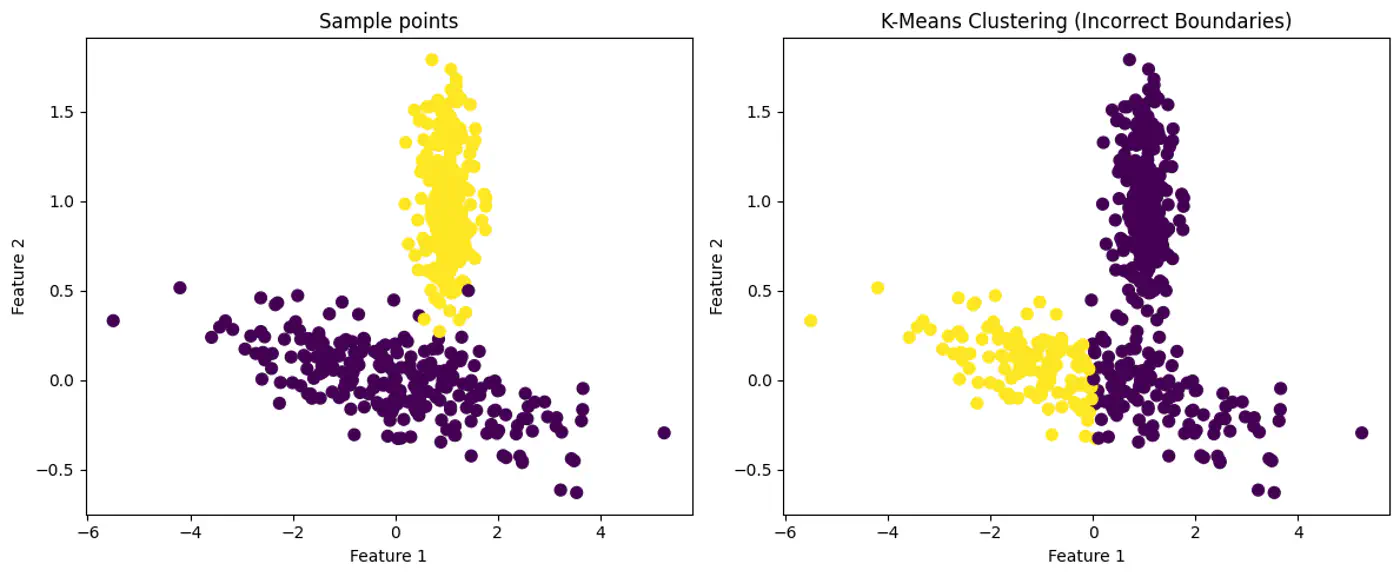
💡GMM: ‘Probabilistic evolution’ of K-Means
⭐️ GMM provides soft assignments and can model elliptical clusters by accounting for variance and correlation between features.
Note: GMM assumes that all data points are generated from a mixture of a finite number of Gaussian Distributions with unknown parameters.
👉GMM can Model Elliptical Clusters.
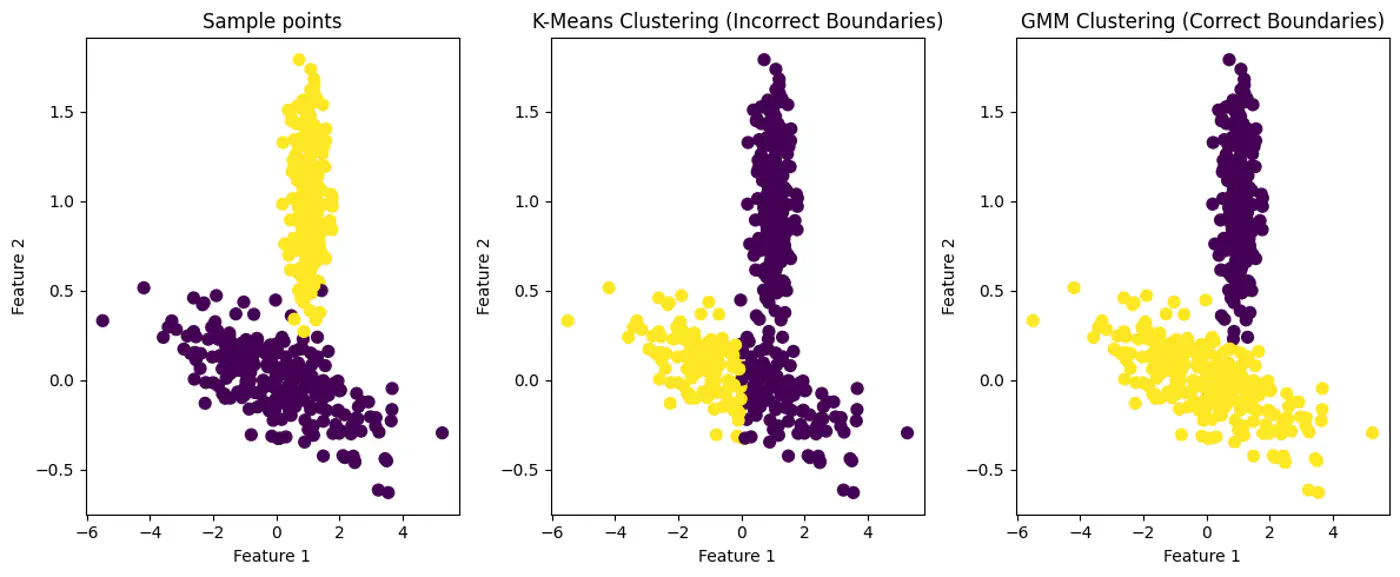
💡‘Combination of probability distributions’.
👉Soft Assignment: Instead of a simple ‘yes’ or ’no’ for cluster membership, a data point is assigned a set of probabilities, one for each cluster.
e.g: A data point might have a 60% probability of belonging to cluster ‘A’, 30% probability for cluster ‘B’, and 10% probability for cluster ‘C’.
👉Gaussian Mixture Model Example:
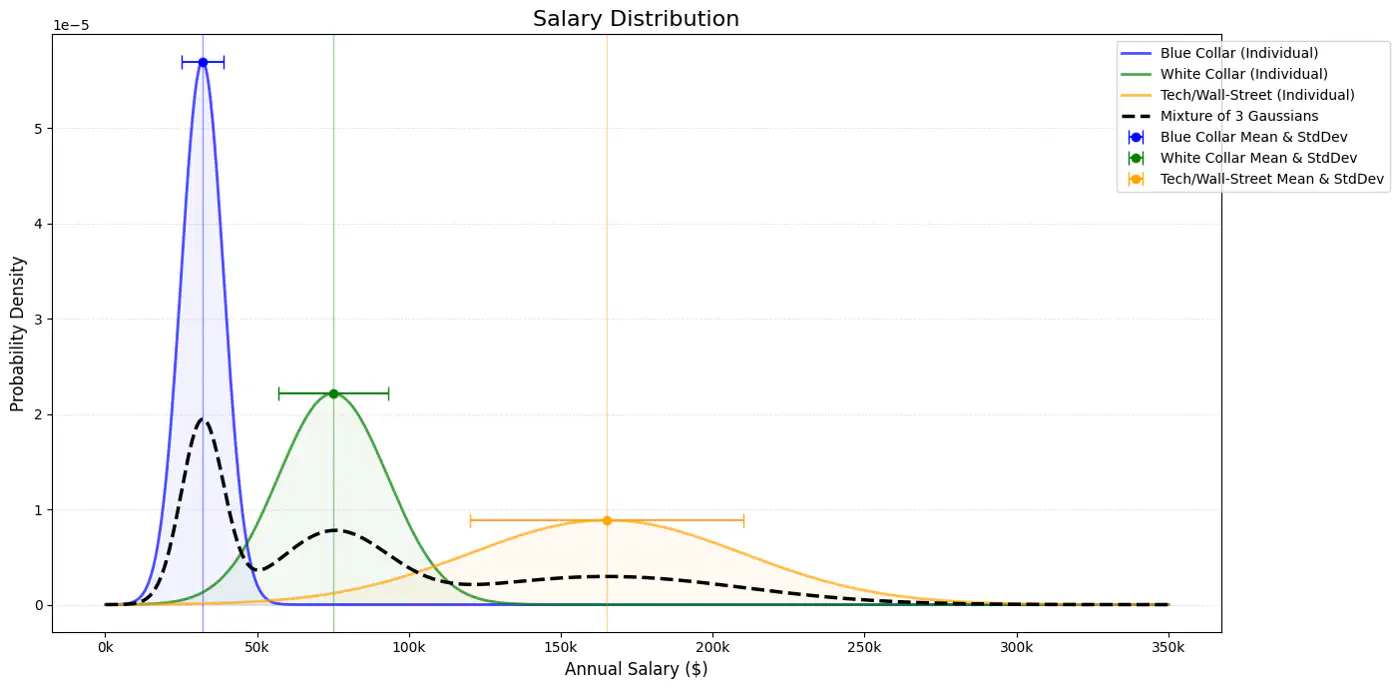
Note: The term \(1/(\sqrt{2\pi \sigma ^{2}})\) is a normalization constant to ensure the total area under the curve = 1.
👉Multivariate Gaussian Example:
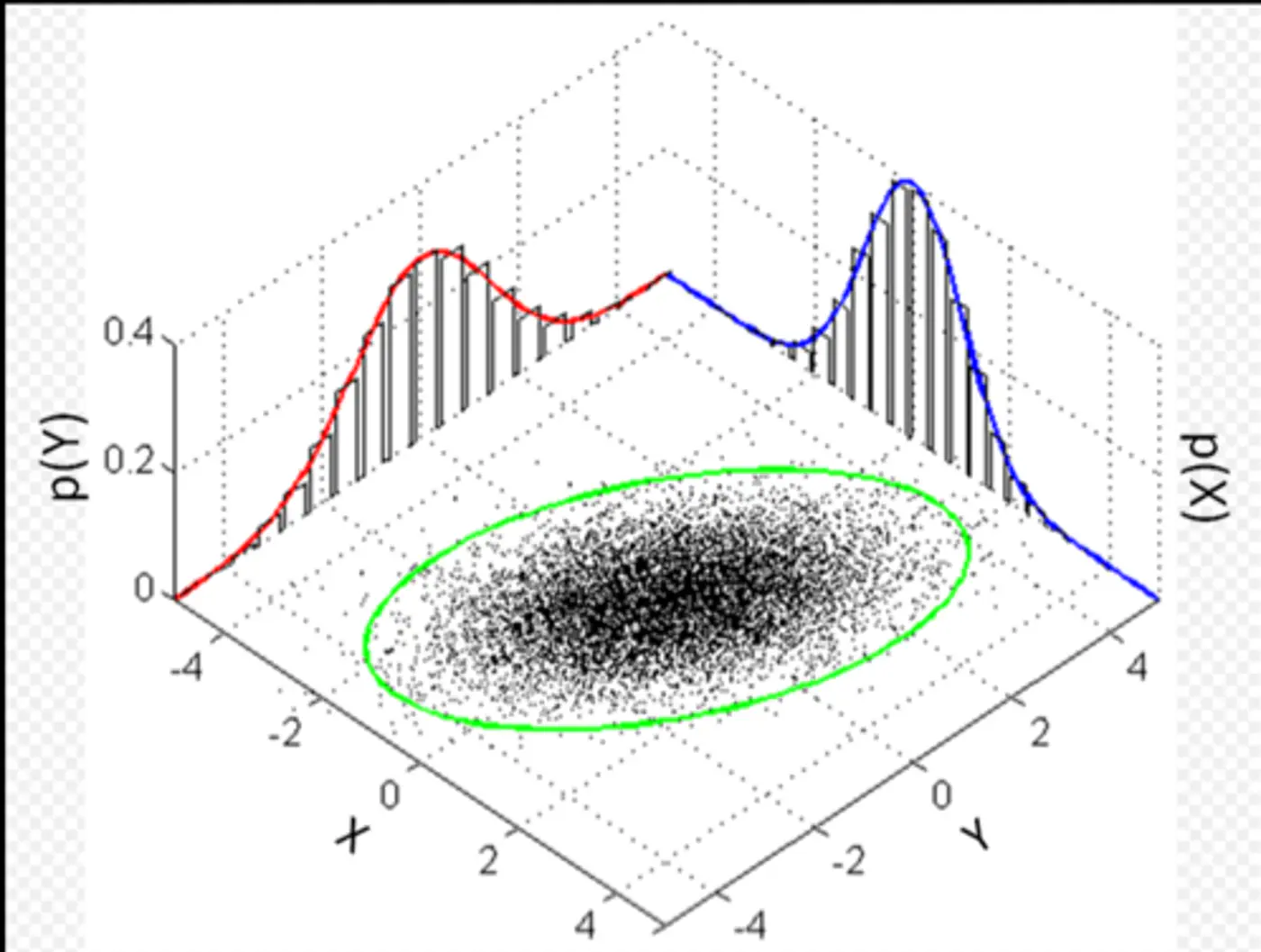
Whenever we have multivariate Gaussian, then the variables may be independent or correlated.
👉Feature Covariance:
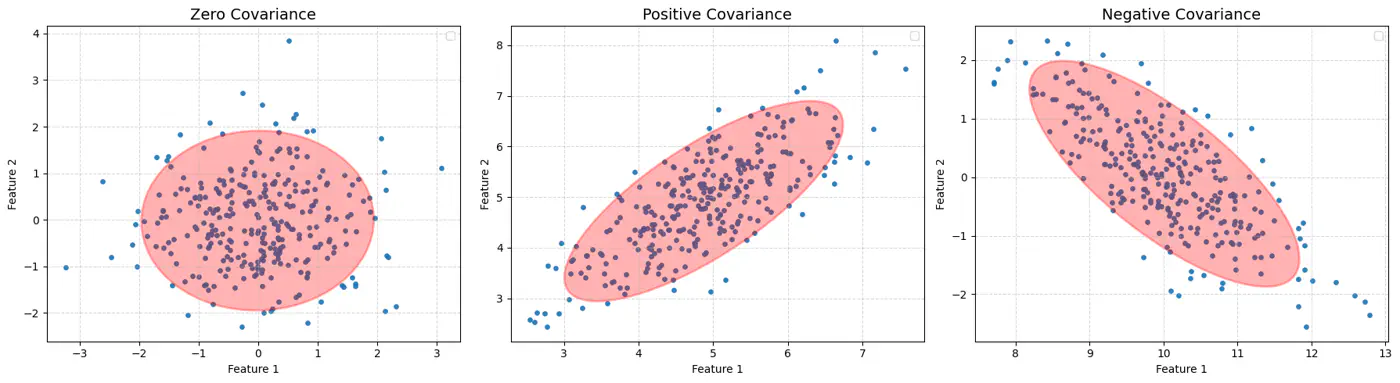
👉Gaussian Mixture with PDF
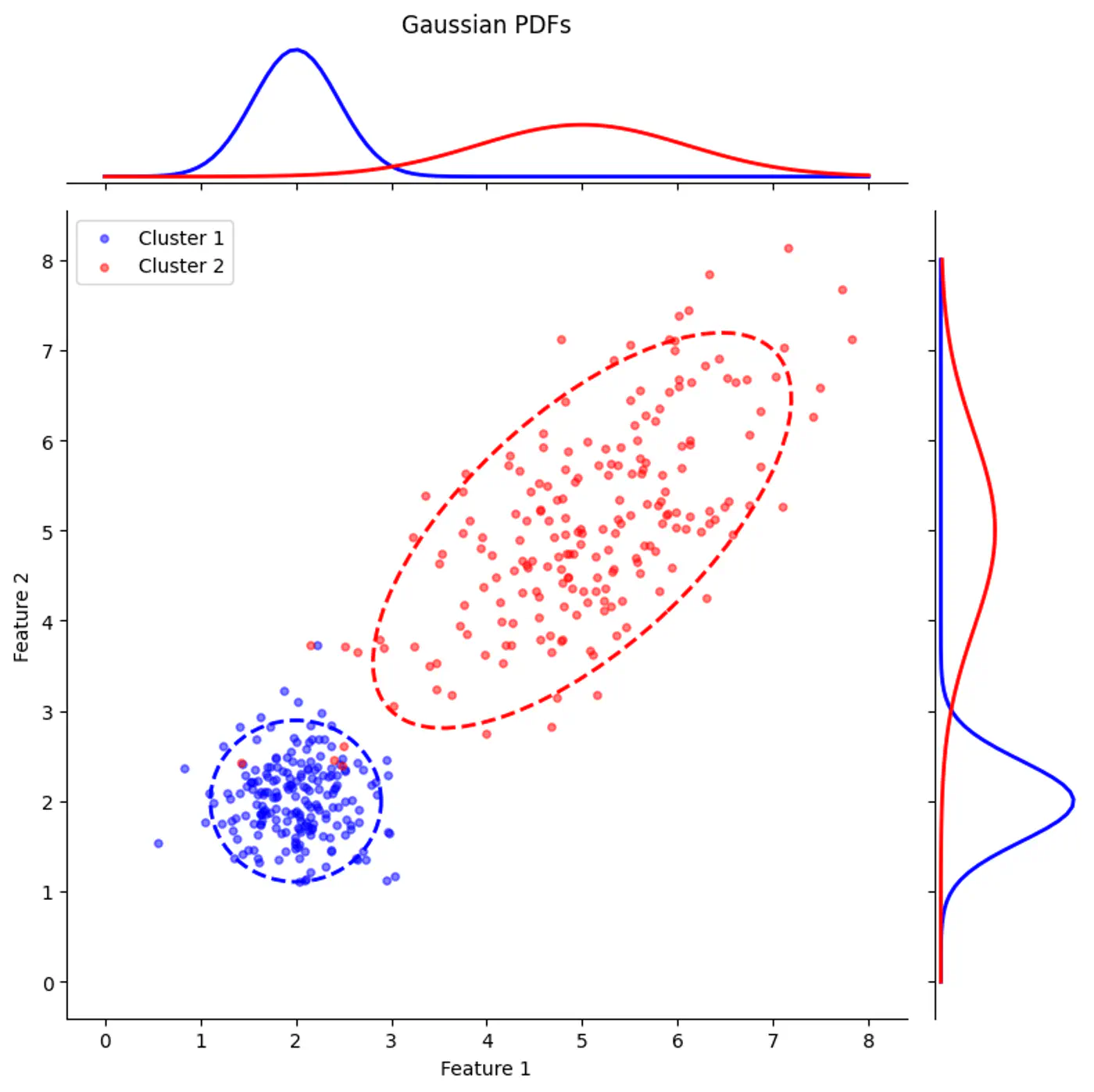
👉Gaussian Mixture (2D)
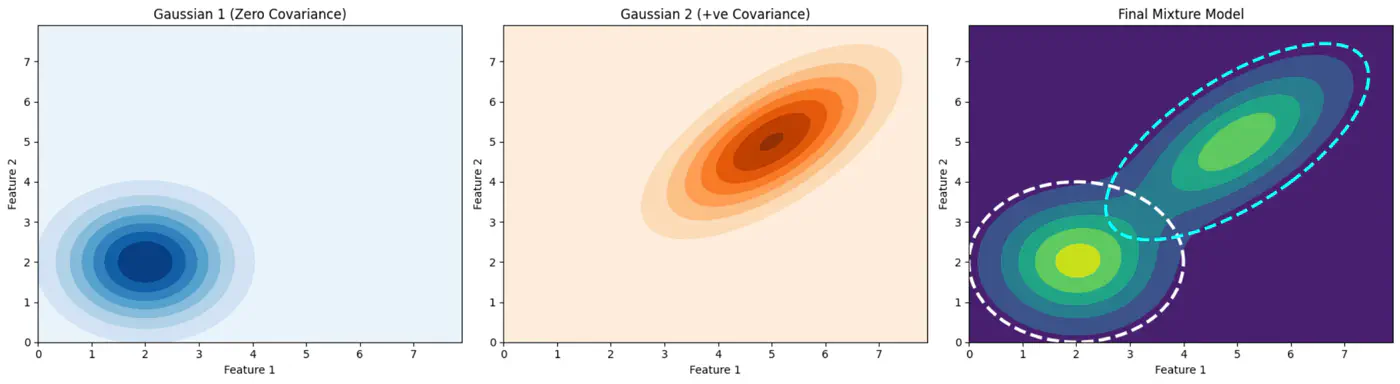
End of Section
1.3.4.2 - Latent Variable Model
⭐️Probabilistic model that assumes data is generated from a mixture of several Gaussian (normal) distributions with unknown parameters.
🎯GMM represents the probability density function of the data as a weighted sum of ‘K’ component Gaussian densities.
👉Below plot shows the probability of a point being generated by 3 different Gaussians.
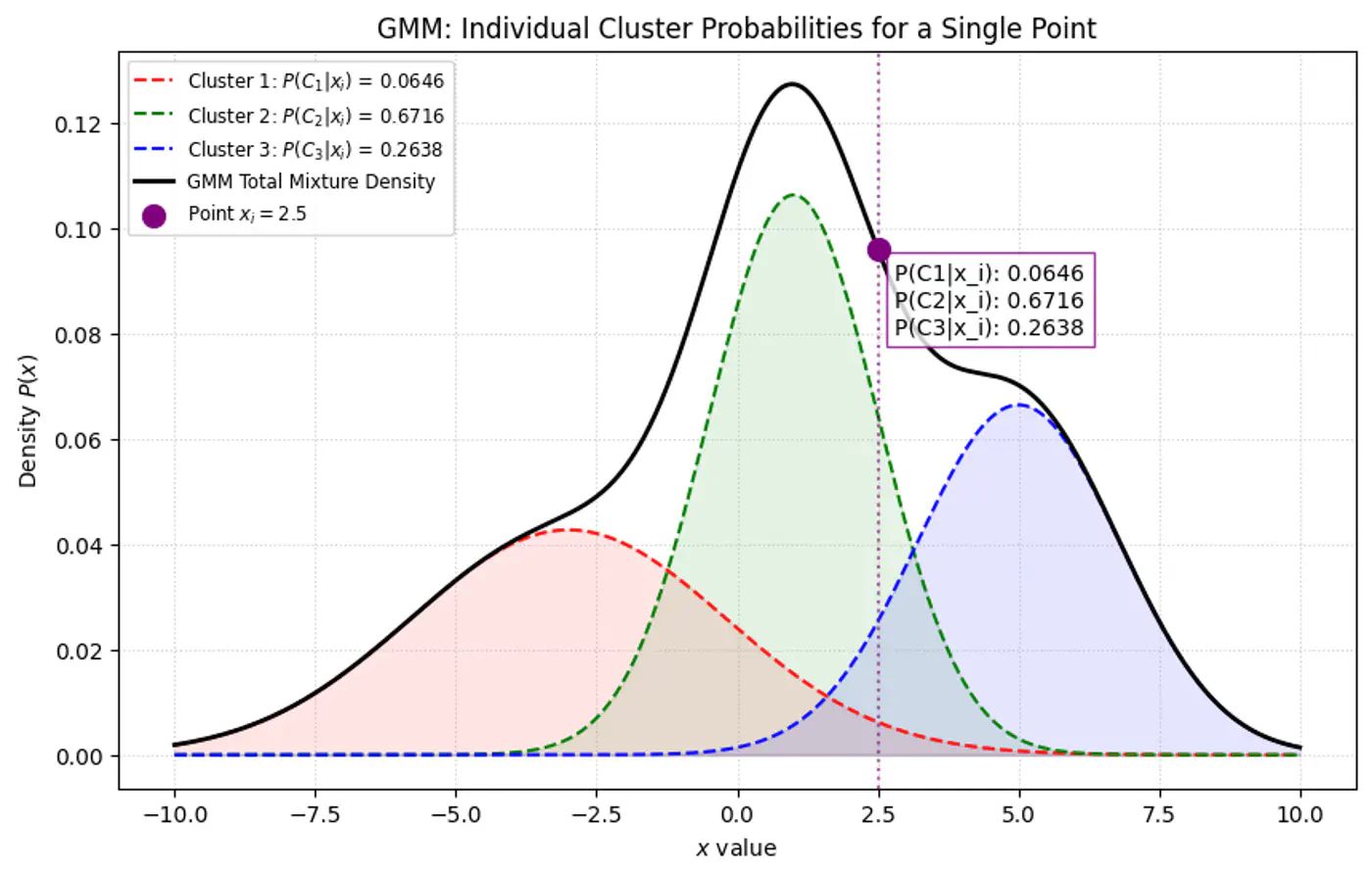
Overall density \(p(x_i|\mathbf{\theta })\) for a data point ‘\(x_i\)’:
\[p(x_i|\mathbf{\mu},\mathbf{\Sigma} )=\sum _{k=1}^{K}\pi _{k}\mathcal{N}(x_i|\mathbf{\mu }_{k},\mathbf{\Sigma }_{k})\]- K: number of component Gaussians.
- \(\pi_k\): mixing coefficient (weight) of the k-th component, such that, \(\pi_k \ge 0\) and \(\sum _{k=1}^{K}\pi _{k}=1\).
- \(\mathcal{N}(x_i|\mathbf{\mu }_{k},\mathbf{\Sigma }_{k})\): probability density function of the k-th Gaussian component with mean \(\mu_k\) and covariance matrix \(\Sigma_k\).
- \(\mathbf{\theta }=\{(\pi _{k},\mathbf{\mu }_{k},\mathbf{\Sigma }_{k})\}_{k=1}^{K}\): complete set of parameters to be estimated.
Note: \(\pi _{k}\approx \frac{\text{Number\ of\ points\ in\ cluster\ }k}{\text{Total\ number\ of\ points\ }(N)}\)
👉 Weight of the cluster is proportional to the number of points in the cluster.
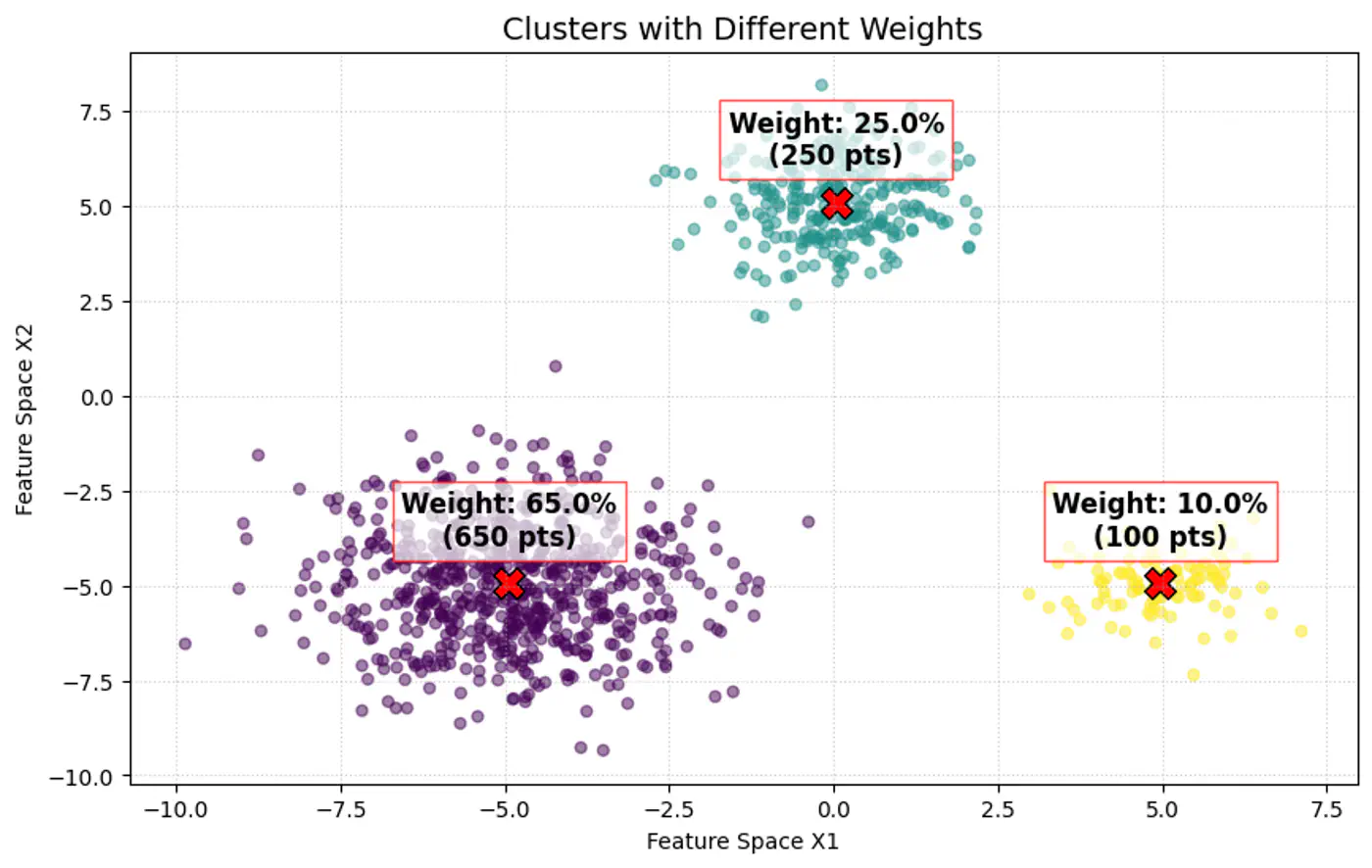
👉Below image shows the weighted Gaussian PDF, given the weights of clusters.
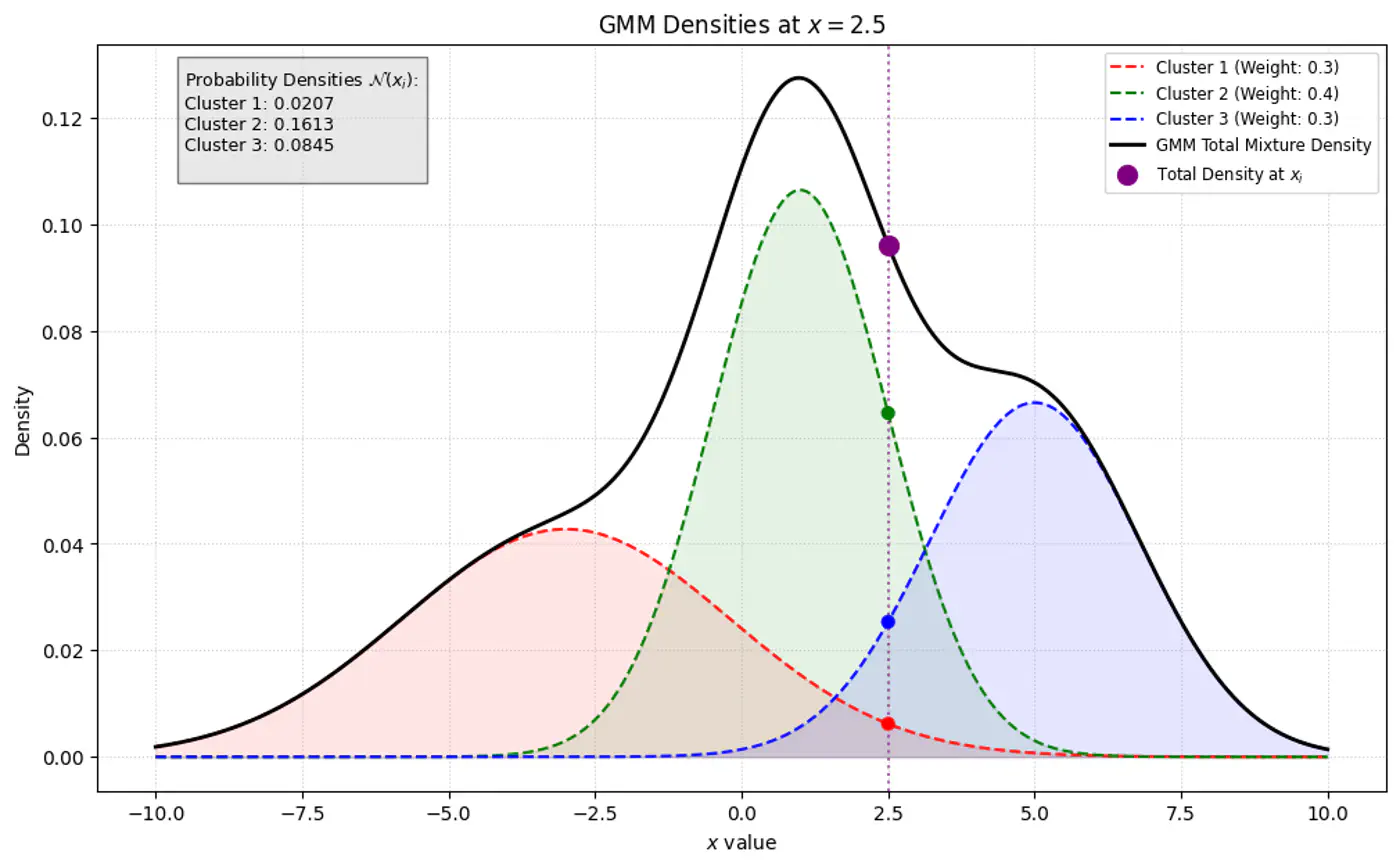
🎯 Goal of a GMM optimization is to find the set of parameters \(\Theta =\{(\pi _{k},\mu _{k},\Sigma _{k})\mid k=1,\dots ,K\}\) that maximize the likelihood of observing the given data.
\[L(\Theta |X)=\sum _{i=1}^{N}\log P(x_i|\Theta )=\sum _{i=1}^{N}\log \left(\sum _{k=1}^{K}\pi _{k}\mathcal{N}(x_i|\mu _{k},\Sigma _{k})\right)\]- 🦀 \(\log (A+B)\) cannot be simplified.
- 🦉So, is there any other way ?
⭐️Imagine we are measuring the heights of people in a college.
- We see a distribution with two peaks (bimodal).
- We suspect there are two underlying groups:
- Group A (Men) and Group B (Women).
Observation:
- Observed Variable (X): Actual height measurements.
- Latent Variable (Z): The ‘label’ (Man or Woman) for each person.
Note: We did not record gender, so it is ‘hidden’ or ‘latent’.
⭐️GMM is a latent variable model, meaning each data point \(\mathbf{x}_{i}\) is assumed to have an associated unobserved (latent) variable \(z_{i}\in \{1,\dots ,K\}\) indicating which component generated it.
Note: We observe the data point, but we do not observe which cluster it belongs to (\(z_i\)).
👉If we knew the value of \(z_i\) (component indicator) for every point, estimating the parameters of each Gaussian component would be straightforward.
Note: The challenge lies in estimating both the parameters of the Gaussians and the values of the latent variables simultaneously.
- With ‘z’ unknown:
- maximize: \[ \log \sum _{k}\pi _{k}\mathcal{N}(x_{i}|\mu _{k},\Sigma _{k}) = \log \Big(\pi _{1}\mathcal{N}(x_{i}\mid \mu _{1},\Sigma _{1})+\pi _{2}\mathcal{N}(x_{i}\mid \mu _{2},\Sigma _{2})+ \dots + \pi _{k}\mathcal{N}(x_{i}\mid \mu _{k},\Sigma _{k})\Big)\]
- \(\log (A+B)\) cannot be simplified.
- maximize: \[ \log \sum _{k}\pi _{k}\mathcal{N}(x_{i}|\mu _{k},\Sigma _{k}) = \log \Big(\pi _{1}\mathcal{N}(x_{i}\mid \mu _{1},\Sigma _{1})+\pi _{2}\mathcal{N}(x_{i}\mid \mu _{2},\Sigma _{2})+ \dots + \pi _{k}\mathcal{N}(x_{i}\mid \mu _{k},\Sigma _{k})\Big)\]
- With ‘z’ known:
- The log-likelihood of the ‘complete data’ simplifies into a sum of logarithms:
\[\sum _{i}\log (\pi _{z_{i}}\mathcal{N}(x_{i}|\mu _{z_{i}},\Sigma _{z_{i}}))\]
- Every point is assigned to exactly one cluster, so the sum disappears because there is only one cluster responsible for that point.
- The log-likelihood of the ‘complete data’ simplifies into a sum of logarithms:
\[\sum _{i}\log (\pi _{z_{i}}\mathcal{N}(x_{i}|\mu _{z_{i}},\Sigma _{z_{i}}))\]
Note: This allows the logarithm to act directly on the exponential term of the Gaussian, leading to simple linear equations.
👉When ‘z’ is known, every data point is ‘labeled’ with its parent component.
To estimate the parameters (mean \(\mu_k\) and covariance \(\Sigma_k\)) for a specific component ‘k’ :
- Gather all data points \(x_i\), where \(z_i\)= k.
- Calculate the standard Maximum Likelihood Estimate.(MLE) for that single Gaussian using only those points.
⭐️ Knowing ‘z’ provides exact counts and component assignments, leading to direct formulae for the parameters:
- Mean (\(\mu_k\)): Arithmetic average of all points assigned to component ‘k’.
- Covariance (\(\Sigma_k\)): Sample covariance of all points assigned to component ‘k’.
- Mixing Weight (\(\pi_k\)): Fraction of total points assigned to component ‘k’.
End of Section
1.3.4.3 - Expectation Maximization
- If we knew the parameters (\(\mu, \Sigma, \pi\)) we could easily calculate which cluster ‘z’ each point belongs to (using probability).
- If we knew the cluster assignments ‘z’ of each point, we could easily calculate the parameters for each cluster (using simple averages).
🦉But we do not know either of them, as the parameters of the Gaussians - we aim to find and cluster indicator latent variable is hidden.
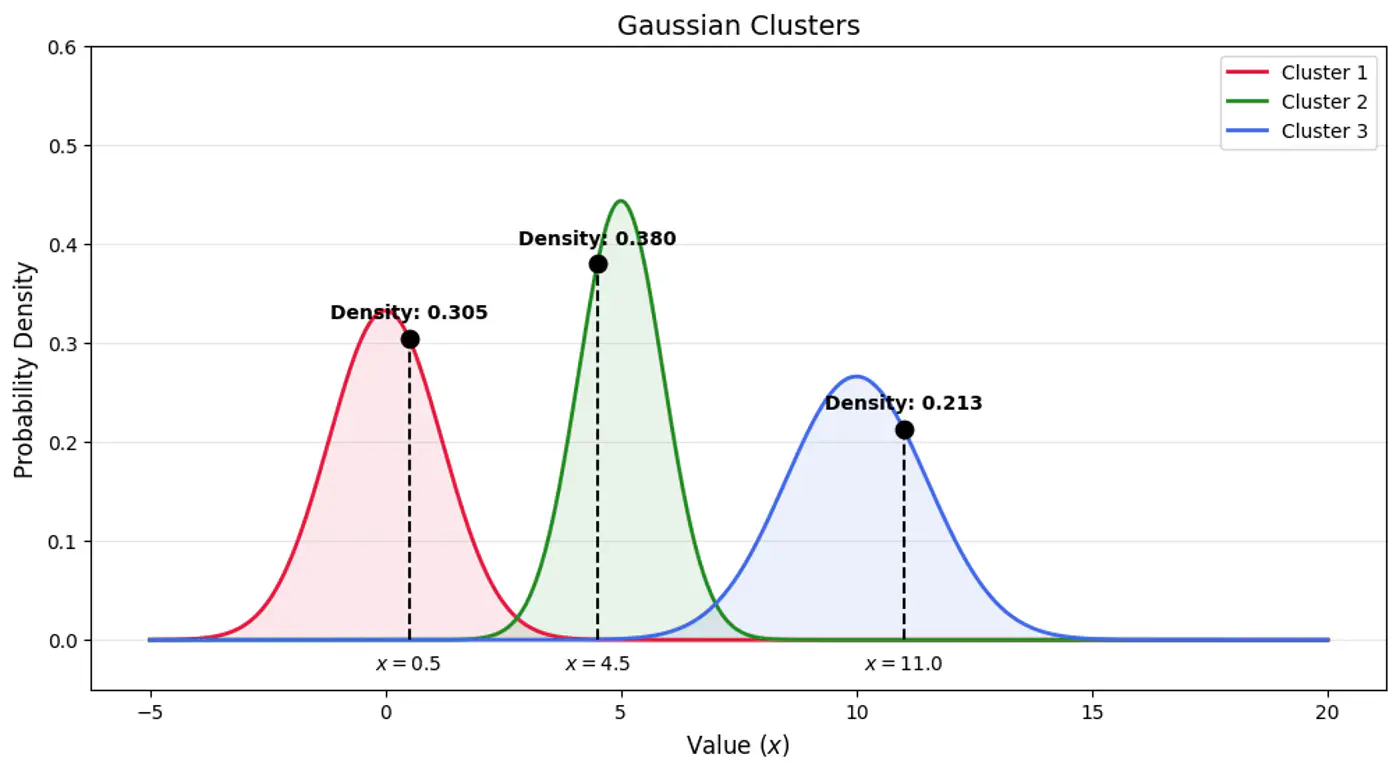
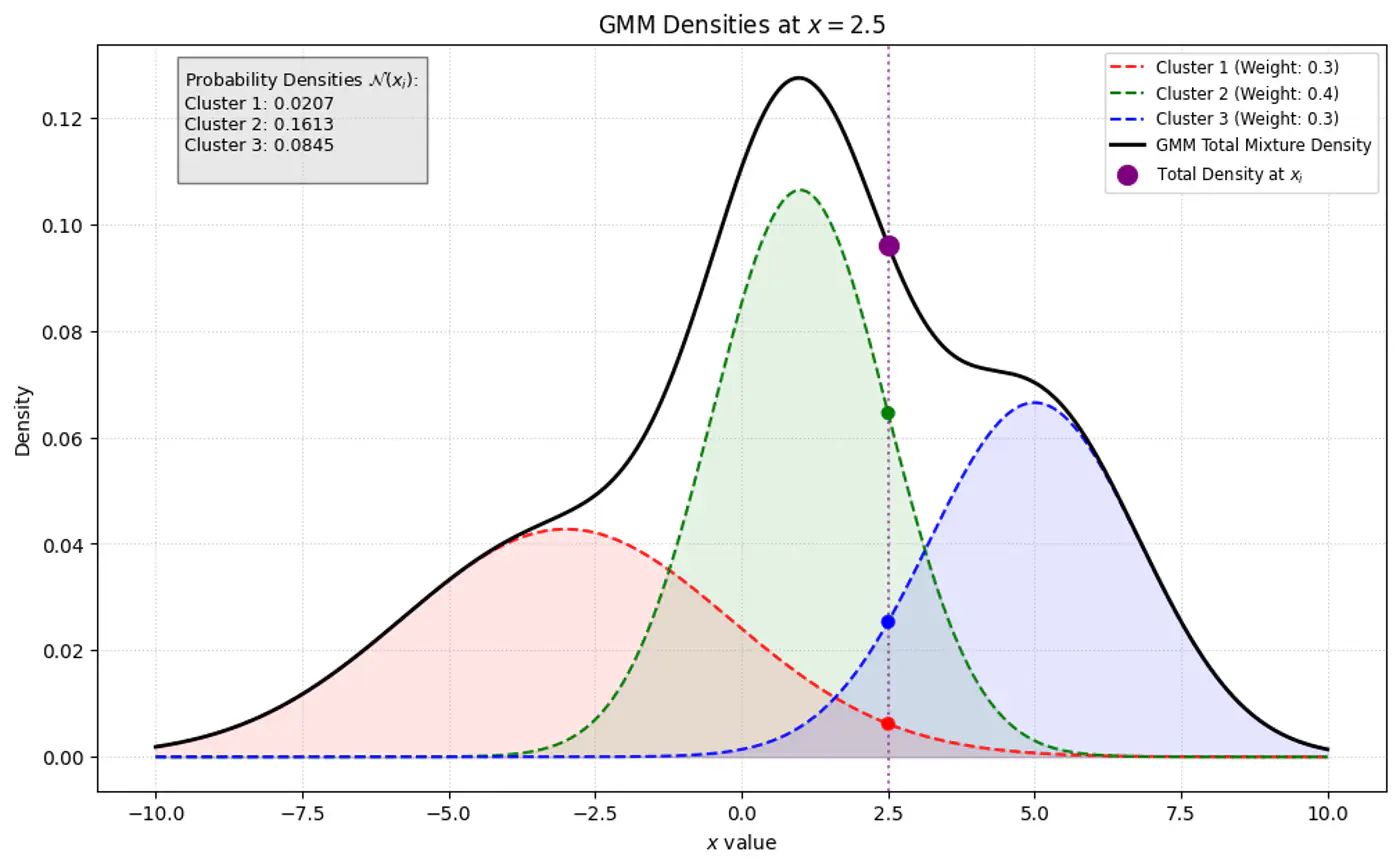
⛳️ Find latent cluster indicator variable \(z_{ik}\).
But \(z_{ik}\) is a ‘hard’ assignment’ (either ‘0’ or ‘1’).
- 🦆 Because we do not observe ‘z’, we use another variable ‘Responsibility’ (\(\gamma_{ik}\)) as a ‘soft’ assignment (value between 0 and 1).
- 🐣 \(\gamma_{ik}\) is the expected value of the latent variable \(z_{ik}\), given the observed data \(x_{i}\) and parameters \(\Theta\). \[\gamma _{ik}=E[z_{ik}\mid x_{i},\theta ]=P(z_{ik}=1\mid x_{i},\theta )\]
Note: \(\gamma_{ik}\) is the posterior probability (or ‘responsibility’) that cluster ‘k’ takes for explaining data point \(x_{i}\).
⭐️Using Bayes’ Theorem, we derive responsibility (posterior probability that component ‘k’ generated data point \(x_i\)) by combining the prior/weights (\(\pi_k\)) and the likelihood (\(\mathcal{N}(x_{i}\mid \mu _{k},\Sigma _{k})\)).
\[\gamma _{ik}= P(z_{ik}=1\mid x_{i},\theta ) = \frac{P(z_{ik}=1)P(x_{i}\mid z_{ik}=1)}{P(x_{i})}\]\[\gamma _{ik}=\frac{\pi _{k}\mathcal{N}(x_{i}\mid \mu _{k},\Sigma _{k})}{\sum _{j=1}^{K}\pi _{j}\mathcal{N}(x_{i}\mid \mu _{j},\Sigma _{j})}\]👉Bayes’ Theorem: \(P(A|B)=\frac{P(B|A)\cdot P(A)}{P(B)}\)
👉 GMM Densities at point
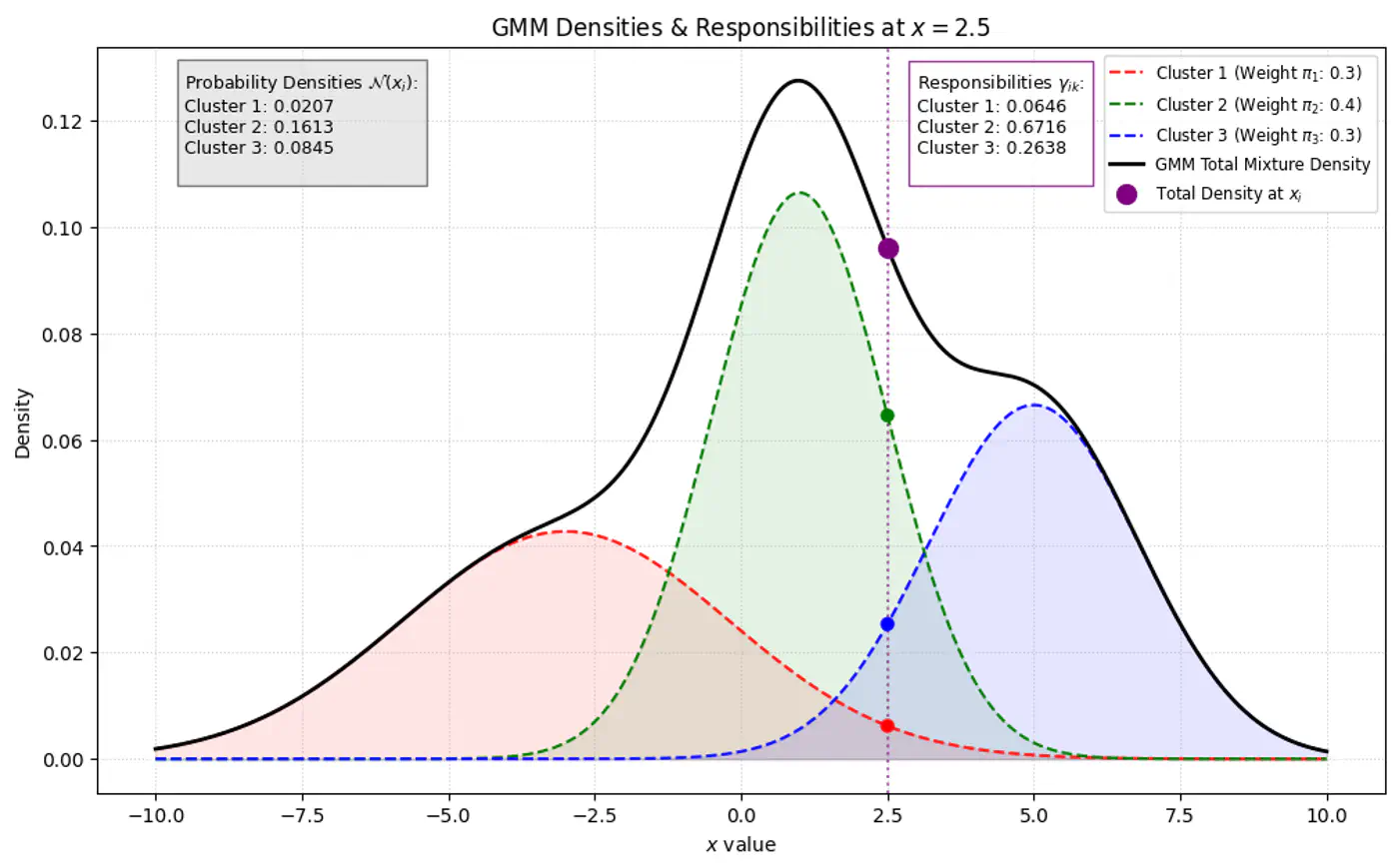
👉GMM Densities at point (different cluster weights)
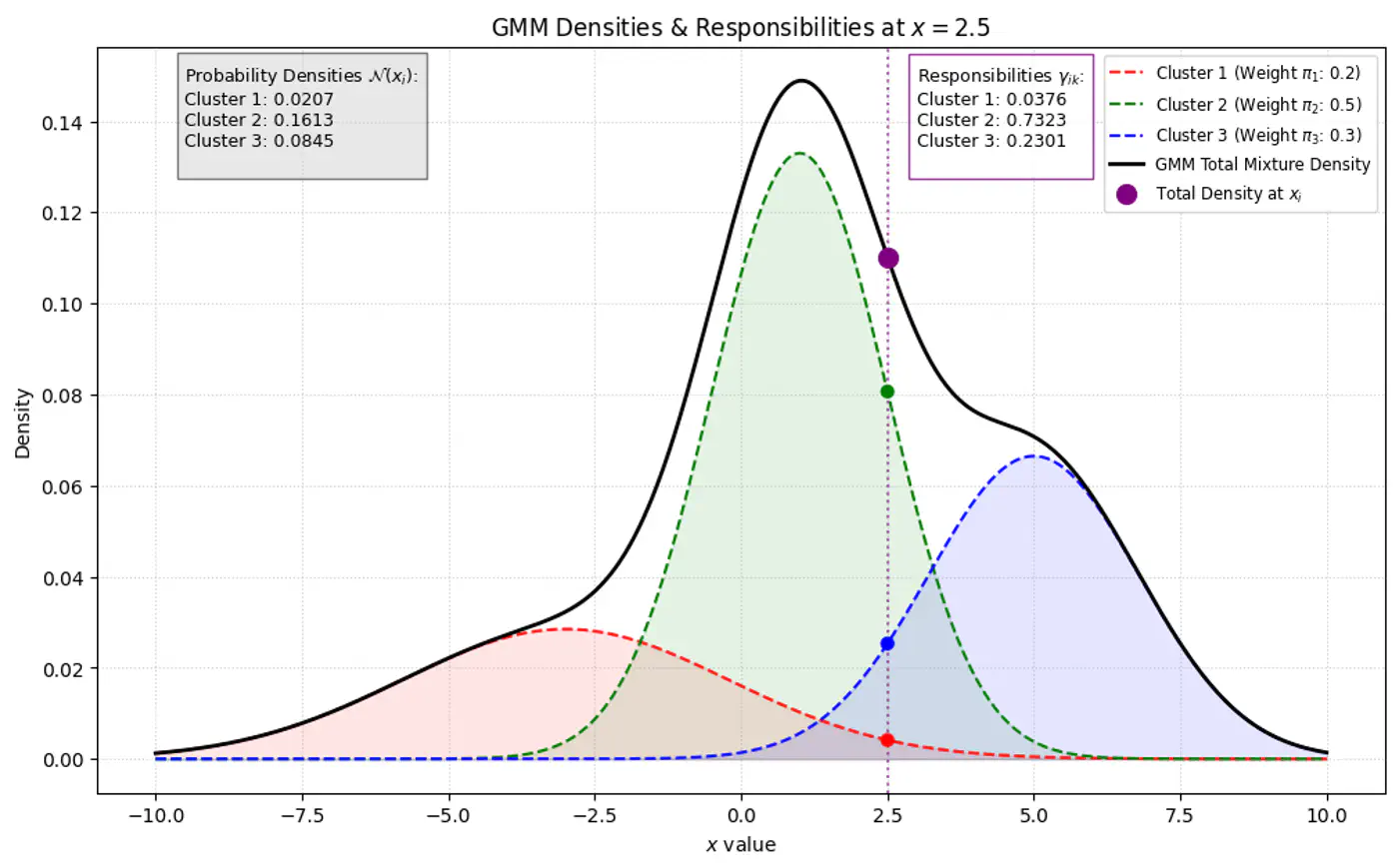
- Initialization: Assign initial values to parameters (\(\mu, \Sigma, \pi\)), often using K-Means results.
- Expectation Step (E): Calculate responsibilities; provides ‘soft’ assignments of points to clusters.
- Maximization Step (M): Update parameters using responsibilities as weights to maximize the expected log-likelihood.
- Convergence: Repeat ‘E’ and ‘M’ steps until the change in log-likelihood falls below a threshold.
👉Given our current guess of the clusters, what is the probability that point \(x_i\) came from cluster ‘k’ ?
\[\gamma (z_{ik})=p(z_{i}=k|\mathbf{x}_{i},\mathbf{\theta }^{(\text{old})})=\frac{\pi _{k}^{(\text{old})}\mathcal{N}(\mathbf{x}_{i}|\mathbf{\mu }_{k}^{(\text{old})},\mathbf{\Sigma }_{k}^{(\text{old})})}{\sum _{j=1}^{K}\pi _{j}^{(\text{old})}\mathcal{N}(\mathbf{x}_{i}|\mathbf{\mu }_{j}^{(\text{old})},\mathbf{\Sigma }_{j}^{(\text{old})})}\]\(\pi_k\) : Probability that a randomly selected data point \(x_i\) belongs to the k-th component before we even look at the specific value of \(x_i\), such that \(\pi_k \ge 0\) and \(\sum _{k=1}^{K}\pi _{k}=1\).
👉Update the parameters (\(\mu, \Sigma, \pi\)) by calculating weighted versions of the standard MLE formulas using responsibilities as weight 🏋️♀️.
\[ \begin{align*} &\bullet \mathbf{\mu }_{k}^{(\text{new})}=\frac{1}{N_{k}}\sum _{i=1}^{N}\gamma (z_{ik})\mathbf{x}_{i} \\ &\bullet \mathbf{\Sigma }_{k}^{(\text{new})}=\frac{1}{N_{k}}\sum _{i=1}^{N}\gamma (z_{ik})(\mathbf{x}_{i}-\mathbf{\mu }_{k}^{(\text{new})})(\mathbf{x}_{i}-\mathbf{\mu }_{k}^{(\text{new})})^{\top } \\ &\bullet \pi _{k}^{(\text{new})}=\frac{N_{k}}{N} \end{align*} \]- where, \(N_{k}=\sum _{i=1}^{N}\gamma (z_{ik})\) is the effective number of points assigned to component ‘k'.
End of Section
1.3.5 - Anomaly Detection
End of Section
1.3.5.1 - Anomaly Detection
🦄 Anomaly is a rare item, event or observation which deviates significantly from the majority of the data and does not conform to a well-defined notion of normal behavior.
Note: Such examples may arouse suspicions of being generated by a different mechanism, or appear inconsistent with the remainder of that set of data.
❌ Remove Outliers:
- Rejection or omission of outliers from the data to aid statistical analysis, for example to compute the mean or standard deviation of the dataset.
- Remove outliers for better predictions from models, such as linear regression.
🔦Focus on Outliers:
- Fraud detection in banking and financial services.
- Cyber-security: intrusion detection, malware, or unusual user access patterns.
- Supervised
- Semi-Supervised
- Unsupervised (most common) ✅
Note: Labeled anomaly data is often unavailable in real-world scenarios.
- Statistical Methods: Z-Score, large value means outlier, IQR, point beyond fences (Q1 - 1.5*IQR or Q3 + 1.5*IQR) is flagged as an outlier.
- Distance Based: KNN, points far from their neighbors as potential anomalies.
- Density Based: DBSCAN, points in low density regions are considered outliers.
- Clustering Based: K-Means, points far from cluster centroids that do not fit any cluster are anomalies.
- Elliptic Envelope (MCD - Minimum Covariance Determinant)
- One-Class SVM (OC-SVM)
- Local Outlier Factor (LOF)
- Isolation Forest (iForest)
- RANSAC (Random Sample Consensus)
End of Section
1.3.5.2 - Elliptic Envelope
Detect anomalies in multivariate Gaussian data, such as, biometric data (height/weight) where features are normally distributed and correlated.
Assumption: The data can be modeled by a Gaussian distribution.
In a normal distribution, most data points cluster around the mean, and the probability density decreases as we move farther away from the center.
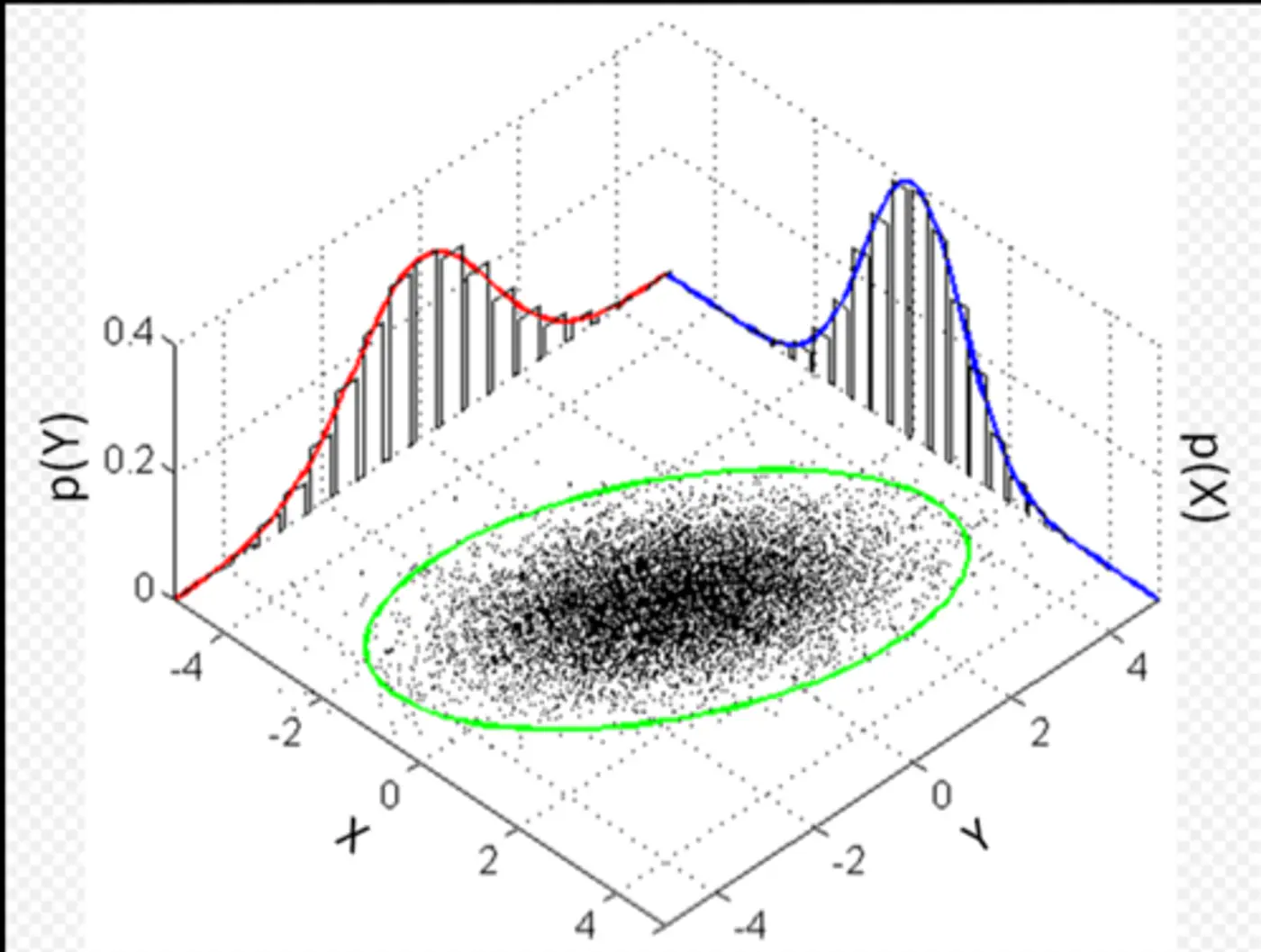
🌍 Euclidean distance measures the simple straight-line distance from the center of the cloud.
👉If the data is spherical, this works fine.
🦕 However, real-world data is often stretched or skewed (e.g., taller people are generally heavier), due to correlations between variables, forming an elliptical shape.
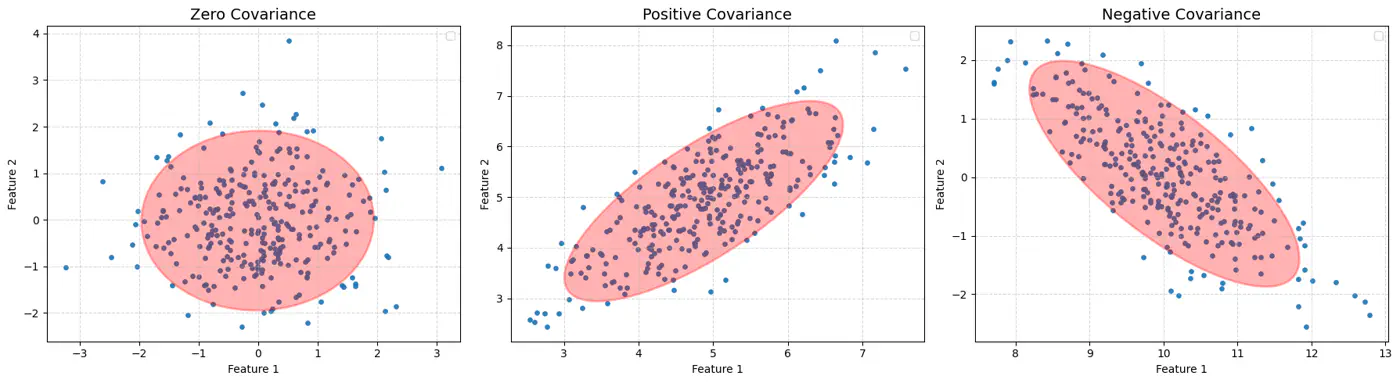
⭐️Mahalanobis distance essentially re-scales the data so that the elliptical distribution appears spherical, and then measures the Euclidean distance in that transformed space.
👉This way, it measures how many standard deviations(\(\sigma\)) away a point is from the mean, considering the data’s spread and correlation (covariance).
\[D_M(x) = \sqrt{(x - \mu)^T \Sigma^{-1} (x - \mu)}\]👉Find the most dense core of the data.
\[D_M(x) = \sqrt{(x - \mu)^T \Sigma^{-1} (x - \mu)}\]🦣 Determinant of covariance matrix \(\Sigma\) represents the volume of the ellipsoid.
⏲️ Therefore, minimize \(|\Sigma|\) to find the tight core.
👉 \(\text {Small} ~ \Sigma \rightarrow \text {Large} ~\Sigma ^{-1}\rightarrow \text {Large} ~ D_{M} ~\text {for outliers}\)
MCD algorithm is used to find the covariance matrix \(\Sigma\) with minimum determinant, so that the volume of the ellipsoid is minimized.
- Initialization: Select several random subsets of size h < n (default h = \(\frac{n+d+1}{2}\), d = # dimensions), representing ‘robust’ majority of the data.
- Calculate preliminary mean (\(\mu\)) and covariance (\(\Sigma\)) for each random subset.
- Concentration Step: Iterative core of the algorithm designed to ‘tighten’ the ellipsoid.
- Calculate Distances: Compute the Mahalanobis distance of all ’n’ points in the dataset from the current subset’s mean (\(\mu\)) and covariance (\(\Sigma\)).
- Select New Subset: Identify the ‘h’ points with the smallest Mahalanobis distances.
- These are the points most centrally located relative to the current ellipsoid.
- Update Estimates: Calculate a new and based only on these ‘h’ most central points.
- Repeat 🔁: The steps repeat until the determinant stops shrinking.
Note: Select the best subset that achieved the absolute minimum determinant.
- Assumptions
- Gaussian data.
- Unimodal data (single center).
- Cost 💰of covariance matrix \(\Sigma^{-1}\) inversion is O(d^3).
End of Section
1.3.5.3 - One Class SVM
⭐️Only one class of data (normal, non-outlier) is available for training, making standard supervised learning models impossible.
e.g. Only normal observations are available for fraud detection, cyber attack, fault detection etc.

🦂 The core problem is to build a model that can distinguish between ‘normal’ and ‘anomalous’ data when we only have examples of the ‘normal’ class during training.
🦖 We need to find a decision boundary that is as compact as possible while still encompassing the bulk of the training data.
🦍 Define a boundary for a single class in high-dimensional space where data might be non-linearly distributed (e.g.‘U’ shape).
🦧 Use the Kernel Trick to project data into a higher-dimensional space and find a hyperplane that separates the data from the origin with the maximum margin.
⭐️OC-SVM, as introduced by Bernhard Schölkopf et al., uses a hyperplane ‘H’ defined by a weight vector \(\mathbf{w}\) and a bias term \(\rho\).
👉Solve the following optimization problem:
\[\min _{\mathbf{w},\xi _{i},\rho }\frac{1}{2}||\mathbf{w}||^{2}+\frac{1}{\nu N}\sum _{i=1}^{N}\xi _{i}-\rho \]Subject to constraints:
\[\mathbf{w}\cdot \phi (\mathbf{x}_{i})\ge \rho -\xi _{i}\quad \text{and}\quad \xi _{i}\ge 0,\quad \text{for\ }i=1,\dots ,N\]- \(\mathbf{x}_{i}\): i-th training data point.
- \(\phi (\mathbf{x}_{i})\): RBF kernel function \(K(x, y) = \exp(-\gamma \|x-y\|^2)\) that maps the data into a higher-dimensional feature space, making it easier to separate from the origin.
- \(\mathbf{w}\): normal vector to the separating hyperplane.
- \(\rho\): scalar bias term that determines the offset of the hyperplane from the origin.
- \(\xi_i\): Slack variables that allow some data points to fall on the ‘wrong’ side of the hyperplane (inside the anomalous region) to prevent overfitting.
- N: total number of training points.
- \(\nu\): hyper-parameter between 0 and 1. It acts as an upper bound on the fraction of outliers (training data points outside the boundary) and a lower bound on the fraction of support vectors.
- \(\frac{1}{2}\|\mathbf{w}\|^{2}\): aims to maximize the margin/compactness of the region.
- \(\frac{1}{\nu N}\sum _{i=1}^{N}\xi _{i}-\rho\): penalizes points (outliers) that violate the boundary constraints.
After solving the optimization problem using standard quadratic programming techniques, we obtain the optimal \(\mathbf{w}^{*}\) and \(\rho ^{*}\).
For a new data point \(x_{new}\), decision function is:
\[f(\mathbf{x}_{\text{new}})=\text{sign}(\mathbf{w}^{*}\cdot \phi (\mathbf{x}_{\text{new}})-\rho ^{*})\]\(f(\mathbf{x}_{\text{new}})\ge 0\): normal point.
\(f(\mathbf{x}_{\text{new}})< 0\): anomalous point (outlier).
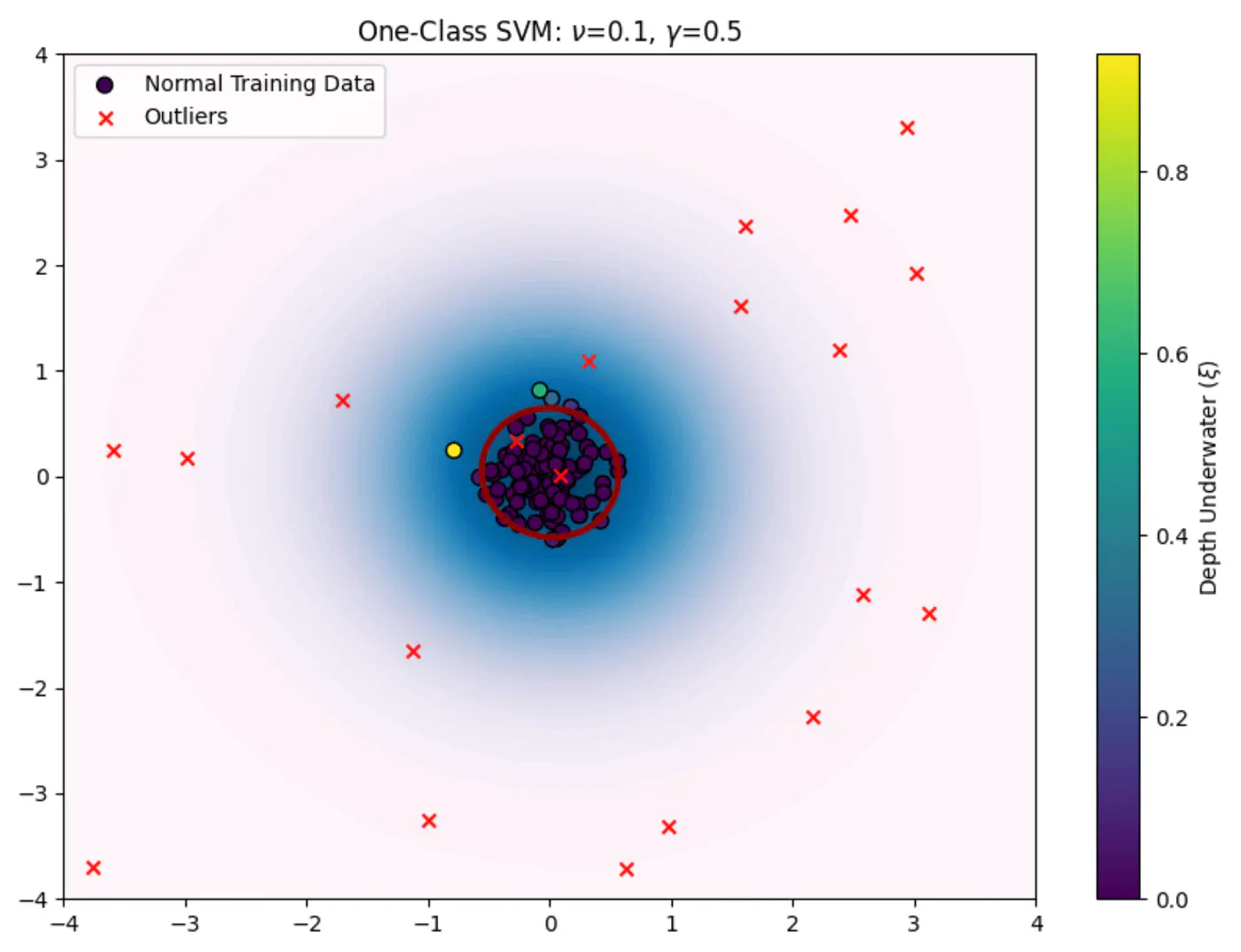
End of Section
1.3.5.4 - Local Outlier Factor
A $100 transaction might be ‘normal’ in New York but an ‘outlier’ in a small rural village.
‘Local context matters.’
Global distance metrics fail when density is non-uniform.
🦄 An outlier is a point that is ‘unusual’ relative to its immediate neighbors, regardless of how far it is from the center of the entire dataset.
💡Traditional distance-based outlier detection methods, such as, KNN, often struggle with datasets where data is clustered at varying densities.
- A point in a sparse region might be considered an outlier by a global method, even if it is a normal part of that sparse cluster.
- Conversely, a point very close to a dense cluster might be an outlier relative to that specific neighborhood.
👉Calculate the relative density of a point compared to its immediate neighborhood.
e.g. If the neighbors are in a dense crowd and the point is not, it is an outlier.
Local Outlier Factor (LOF) is a density-based algorithm designed to detect anomalies by measuring the local deviation of a data point relative to its neighbors.
👉Size of the red circle represents the LOF score.
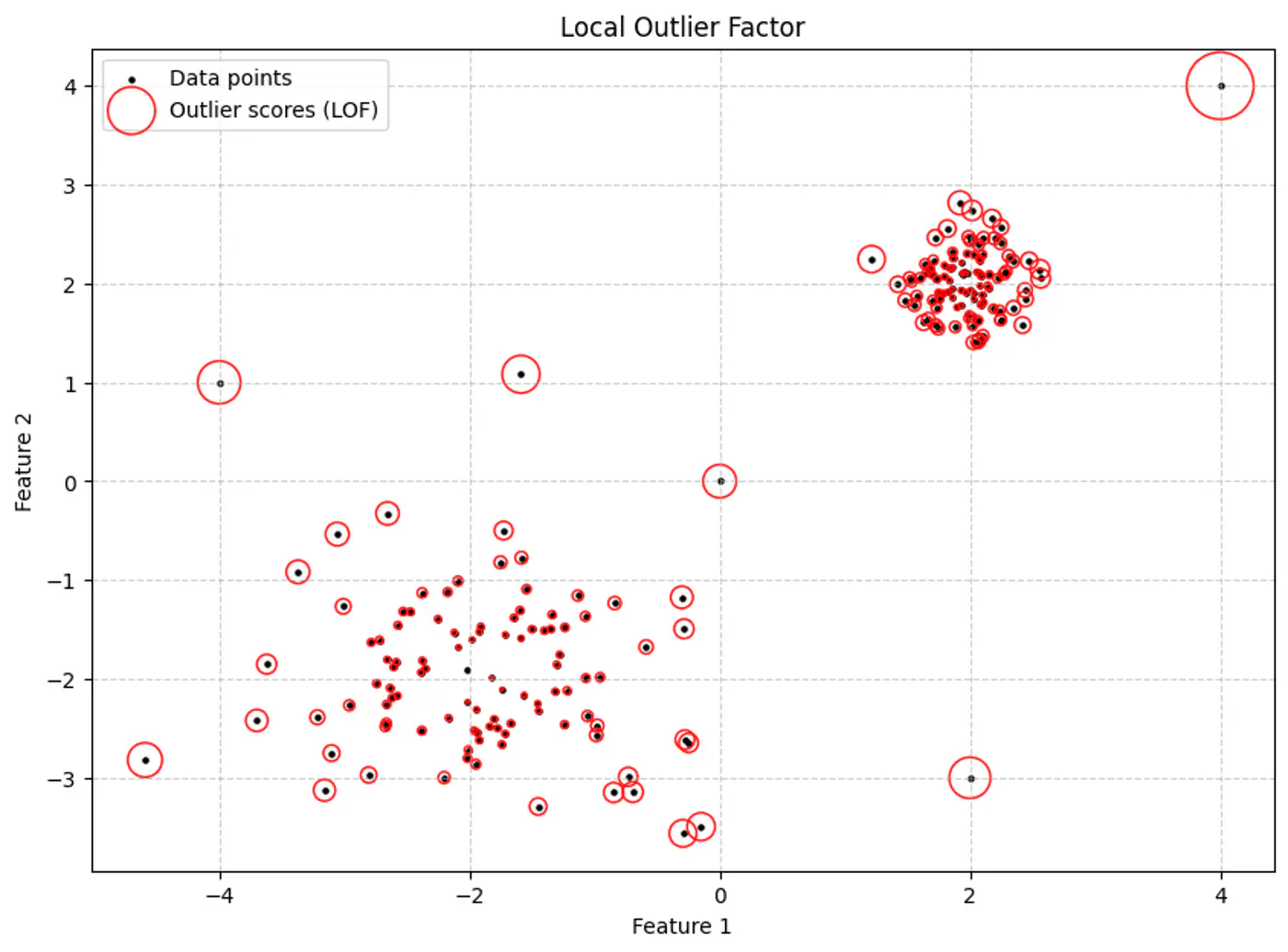
- K-Distance (\(k\text{-dist}(p)\)):
- The distance from point ‘p’ to its k-th nearest neighbor.
- Reachability Distance (\(\text{reach-dist}_{k}(p,o)\)):
\[\text{reach-dist}_{k}(p,o)=\max \{k\text{-dist}(o),\text{dist}(p,o)\}\]
- \(\text{dist}(p,o)\): actual Euclidean distance between ‘p’ and ‘o’.
- This acts as ‘smoothing’ factor.
- If point ‘p’ is very close to ‘o’ (inside o’s k-neighborhood), round up distance to \(k\text{-dist}(o)\).
- \(\text{dist}(p,o)\): actual Euclidean distance between ‘p’ and ‘o’.
- Local Reachability Density (\(\text{lrd}_{k}(p)\)):
- The inverse of the average reachability distance from ‘p’ to its k-neighbors (\(N_{k}(p)\)).
\[\text{lrd}_{k}(p)=\left[\frac{1}{|N_{k}(p)|}\sum _{o\in N_{k}(p)}\text{reach-dist}_{k}(p,o)\right]^{-1}\]
- High LRD: Neighbors are very close; the point is in a dense region.
- Low LRD: Neighbors are far away; the point is in a sparse region.
- The inverse of the average reachability distance from ‘p’ to its k-neighbors (\(N_{k}(p)\)).
\[\text{lrd}_{k}(p)=\left[\frac{1}{|N_{k}(p)|}\sum _{o\in N_{k}(p)}\text{reach-dist}_{k}(p,o)\right]^{-1}\]
- Local Outlier Factor (\(\text{LOF}_{k}(p)\)):
- The ratio of the average ‘lrd’ of p’s neighbors to p’s own ‘lrd’. \[\text{LOF}_{k}(p)=\frac{1}{|N_{k}(p)|}\sum _{o\in N_{k}(p)}\frac{\text{lrd}_{k}(o)}{\text{lrd}_{k}(p)}\]
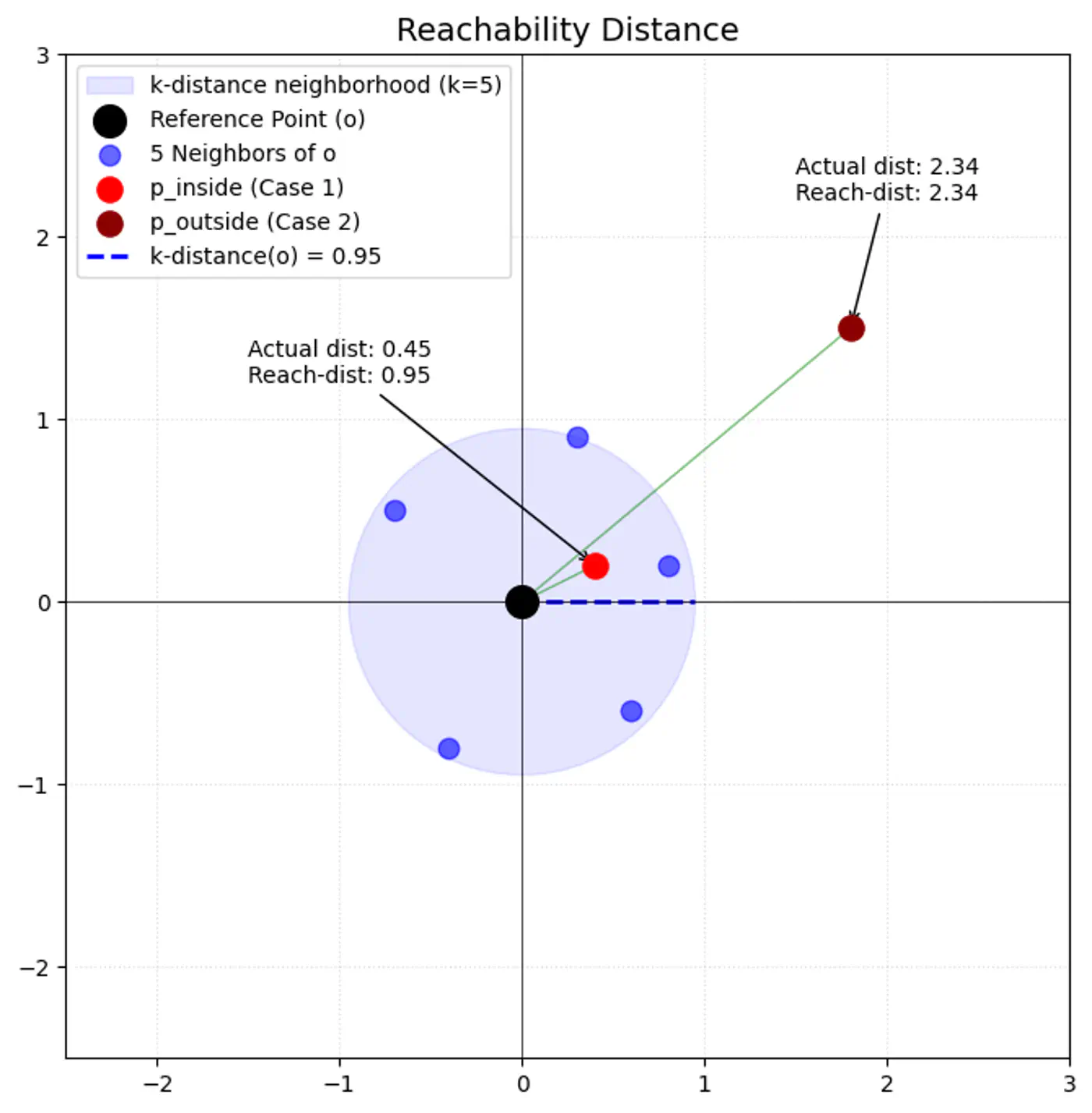
- LOF ≈ 1: Point ‘p’ has similar density to its neighbors (inlier).
- LOF > 1: Point p’s density is much lower than its neighbors’ density (outlier).
End of Section
1.3.5.5 - Isolation Forest
‘Large scale tabular data.’
Credit card fraud detection in datasets with millions of rows and hundreds of features.
Note: Supervised learning requires balanced, labeled datasets (normal vs. anomaly), which are rarely available in real-world scenarios like fraud or cyber-attacks.
🦥 ‘Flip the logic.’
🦄 ‘Anomalies’ are few and different, so they are much easier to isolate from the rest of the data than normal points.
🐦🔥 ‘Curse of dimensionality.’
🐎 Distance based (K-NN), and density based (LOF) algorithms require calculation of distance between all pair of points.
🐙 As the number of dimensions and data points grows, these calculations become exponentially more expensive 💰 and less effective.
‘Randomly partition the data.’
🦄 If a point is an outlier, it will take fewer partitions (splits) to isolate it into a leaf 🍃 node compared to a point that is buried deep within a dense cluster of normal data.
🌳Isolation Forest uses an ensemble of ‘Isolation Trees’ (iTrees) 🌲.
👉iTree (Isolation Tree) 🌲 is a proper binary tree structure specifically designed to separate individual data points through random recursive partitioning.
- Sub-sampling:
- Select a random subset of data (typically 256 instances) to build an iTree.
- Tree Construction: Randomly select a feature.
- Randomly select a split value between the minimum and maximum values of that feature.
- Divide the data into two branches based on this split.
- Repeat recursively until the point is isolated or a height limit is reached.
- Forest Creation:
- Repeat 🔁 the process to create ‘N’ trees (typically 100).
- Inference:
- Pass a new data point through all trees, calculate the average path length, and compute the anomaly score.
⭐️🦄 Assign an anomaly score based on the path length h(x) required to isolate a point ‘x’.
- Path Length (h(x)): The number of edges ‘x’ traverses from the root node to a leaf node.
- Average Path Length (c(n)): Since iTrees are structurally similar to Binary Search Trees (BST), the average path length for a dataset of size ’n’ is given by: \[c(n)=2H(n-1)-\frac{2(n-1)}{n}\]
where, H(i) is the harmonic number, estimated as \(\ln (i)+0.5772156649\) (Euler’s constant).
To normalize the score between 0 and 1, we define it as:
\[s(x,n)=2^{-\frac{E(h(x))}{c(n)}}\]👉E(H(x)): is the average path length of across a forest of trees 🌲.
- \(s\rightarrow 1\): Point is an anomaly; Path length is very short.
- \(s\approx 0.5\): Point is normal, path length approximately equal to c(n).
- \(s\rightarrow 0\): Point is normal; deeply buried point, path length is much larger than c(n).
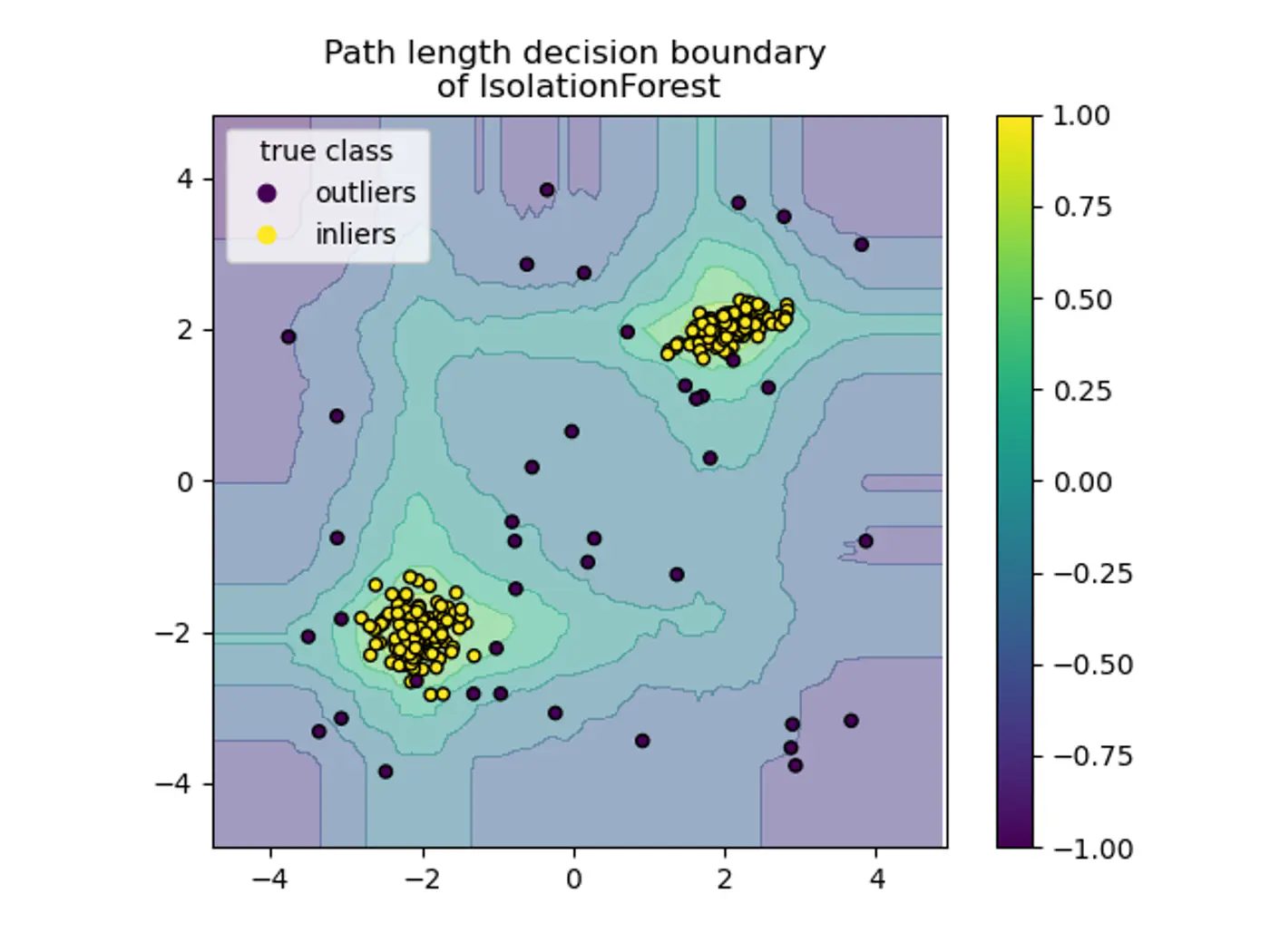
- Axis-Parallel Splits:
- Standard iTrees 🌲 split only on one feature at a time, so:
- We do not get a smooth decision boundary.
- Anything off-axis has a higher probability of being marked as an outlier.
- Note: Extended Isolation Forest fixes this by using random slopes.
- Standard iTrees 🌲 split only on one feature at a time, so:
- Score Sensitivity: The threshold for what constitutes an ‘anomaly’ often requires manual tuning or domain knowledge.
End of Section
1.3.5.6 - RANSAC
👉Ordinary Least Squares use all data points to find a fit.
- However, a single outlier can ‘pull’ the resulting line significantly, leading to a poor representative model.
💡If we pick a very small random subset of points, there is a higher probability that this small subset contains only good data (inliers) compared to a large set.
Ordinary Least Squares (OLS) minimizes the Sum of Errors.
- A huge outlier has an exponentially large impact on the final line.
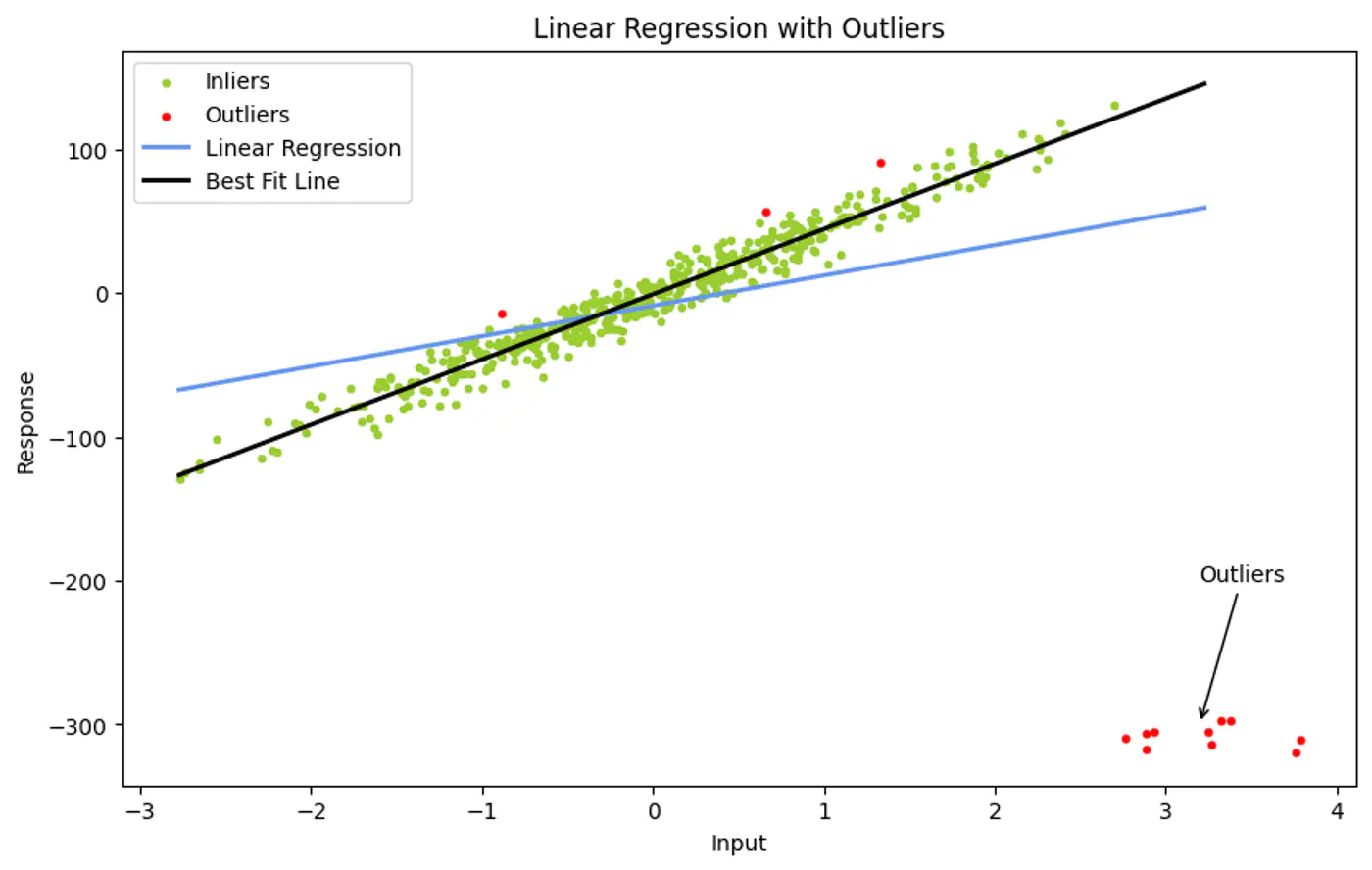
💡Instead of using all points, iteratively pick the smallest possible random subset to fit a model, then check (votes) how many other points in the dataset ‘agree’ with that model.
This gives the name to our algorithm:
- Random: Random subsets.
- Sampling: Small subsets.
- Consensus: Agreement with other points.
- Random Sampling:
- Randomly select a Minimal Sample Set (MSS) of ’n’ points from the input data ‘D’.
- e.g. n=2 for a line, or n=3 for a plane in 3D.
- Model Fitting:
- Compute the model parameters using only these ’n’ points.
- Test:
- For all other points in ‘D’, calculate the error relative to the model.
- 👉 Points with error < \(\tau\)(threshold) are added to the ‘Consensus Set’.
- Evaluate:
- If the consensus set is larger than the previous best, save this model and set.
- Repeat 🔁:
- Iterate ‘k’ times.
- Refine (Optional):
- Once the best model is found, re-estimate it using all points in the final consensus set (usually via Least Squares) for a more precise fit.
👉Example:
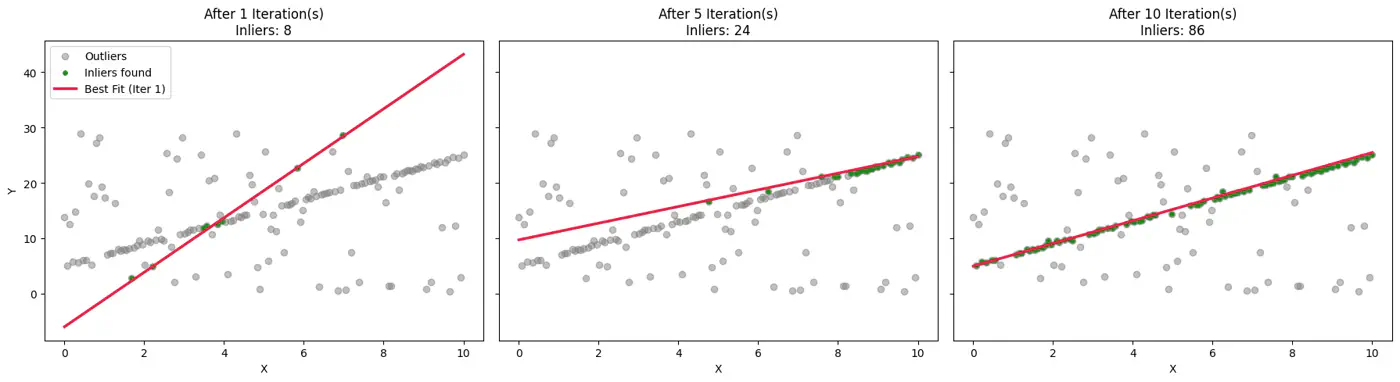
👉To ensure the algorithm finds a ‘clean’ sample set (no outliers) with a desired probability(often 99%), we use the following formula:
\[k=\frac{\log (1-P)}{\log (1-w^{n})}\]- k: Number of iterations required.
- P: Probability that at least one random sample contains only inliers.
- w: Ratio of inliers in the dataset (number of inliers / total points).
- n: Number of points in the Minimal Sample Set.
End of Section
1.3.6 - Dimensionality Reduction
End of Section
1.3.6.1 - PCA
- Data Compression
- Noise Reduction
- Feature Extraction: Create a smaller set of meaningful features from a larger one.
- Data Visualization: Project high-dimensional data down to 2 or 3 dimensions.
Assumption: Linear relationship between features.
💡‘Information = Variance’
☁️ Imagine a cloud of points in 2D space.
👀 Look for the direction (axis) along which the data is most ‘spread out’.
🚀 By projecting data onto this axis, we retain the maximum amount of information (variance).
👉Example 1: Var(Feature 1) < Var(Feature 1)
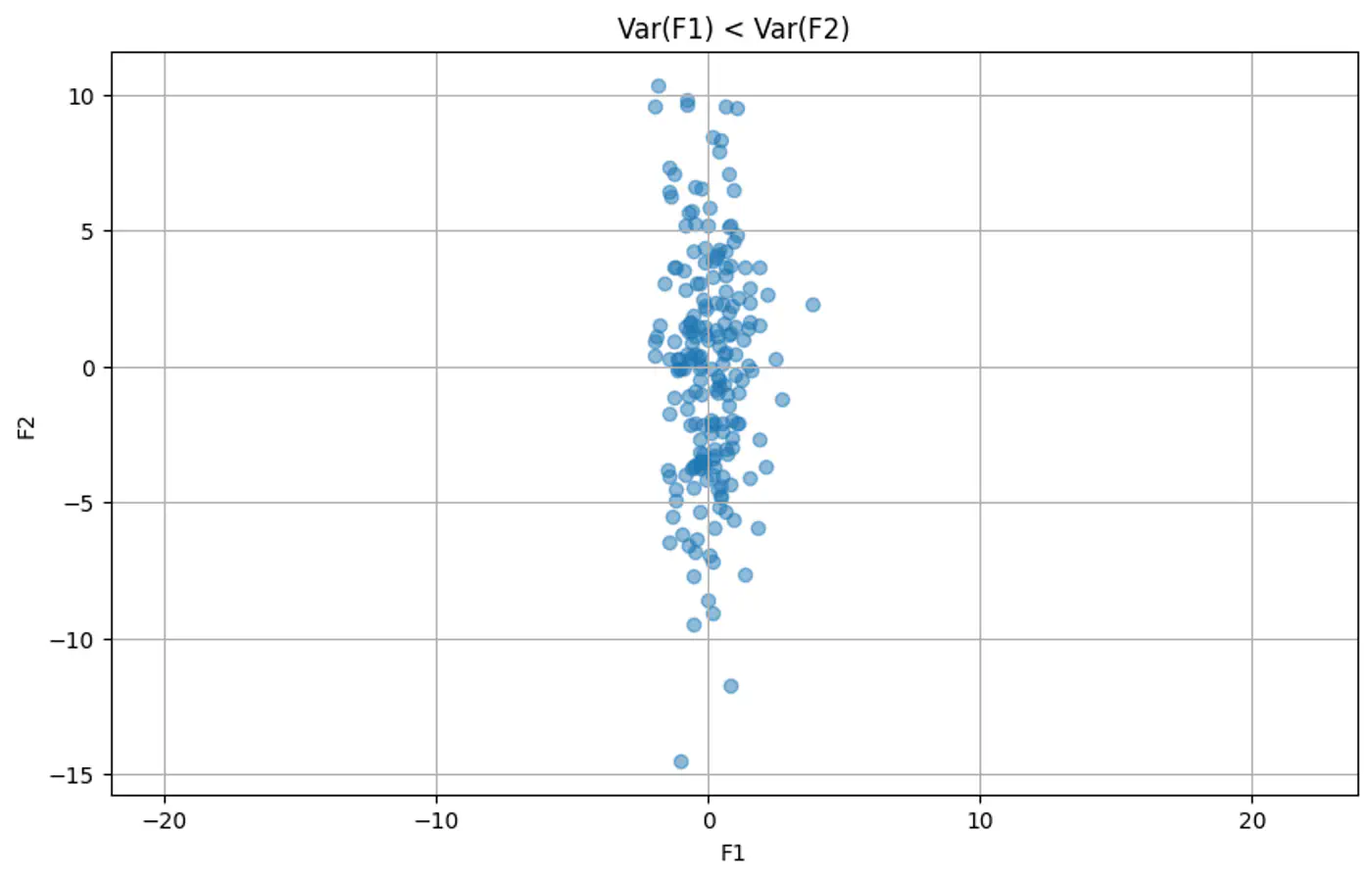
👉Example 2: Red line shows the direction of maximum variance
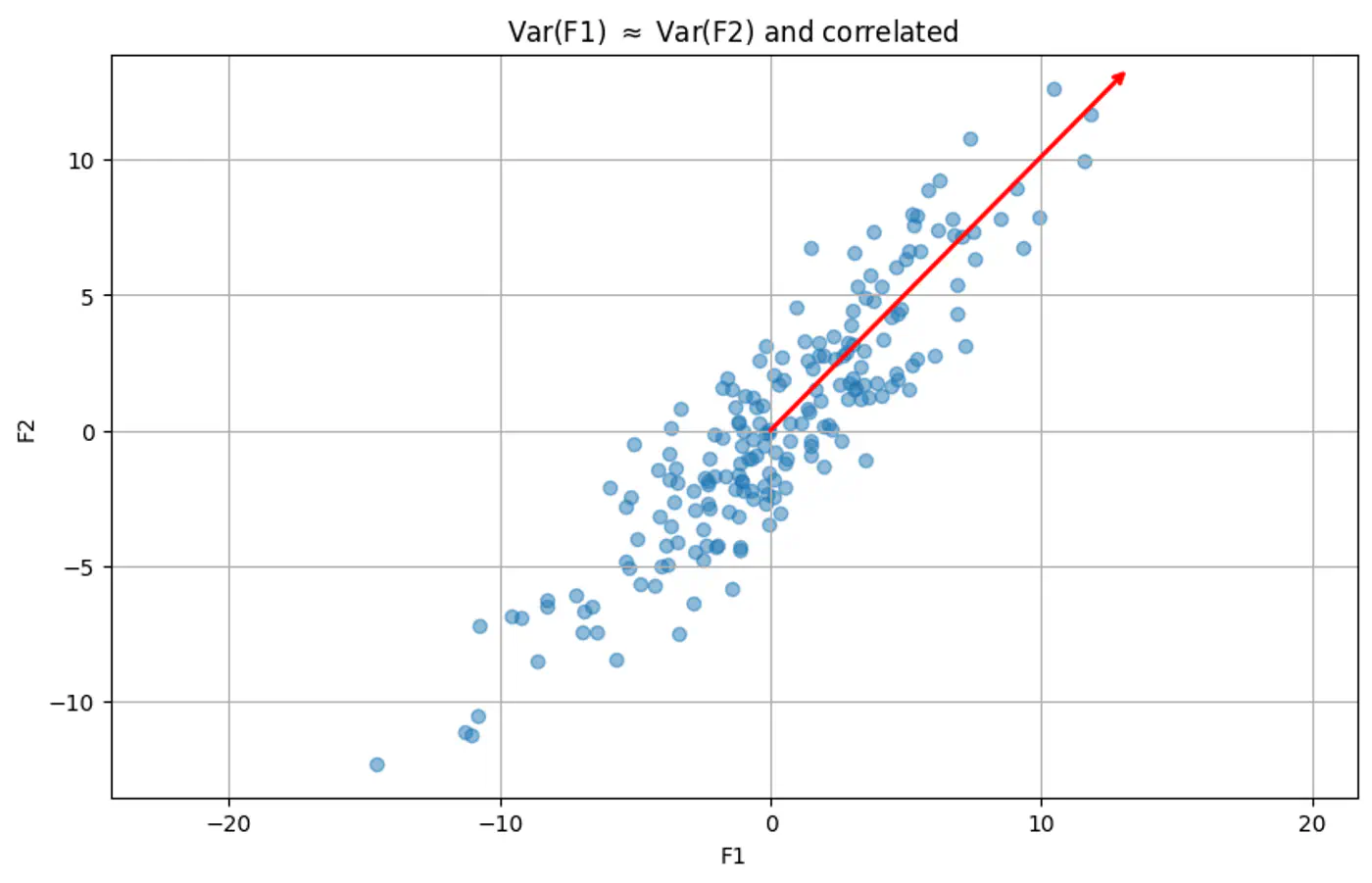
🧭 Finds the direction of maximum variance in the data.
Note: Some loss of information will always be there in dimensionality reduction, because there will be some variability in data along the direction that is dropped, and that will be lost.
- PCA seeks a direction 🧭, represented by a unit vector \(\hat{u}\) onto which data can be projected to maximize variance.
- The projection of a mean centered data point \(x_i\) onto \(u\) is \(u^Tx_i\).
- The variance of these projections can be expressed as \(u^{T}\Sigma u\), where \(\Sigma\) is the data’s covariance matrix.
- Data: Let \(X = \{x_{1},x_{2},\dots ,x_{n}\}\) is the mean centered (\(\bar{x} = 0\)) dataset.
- Projection of point \(x_i\) on \(u\) is \(z_i = u^Tx_i\)
- Variance of projected points (since \(\bar{x} = 0\)): \[\text{Var}(z)=\frac{1}{n}\sum _{i=1}^{n}z_{i}^{2}=\frac{1}{n}\sum _{i=1}^{n}(x_{i}^{T}u)^{2}\]
- 💡Since, \((x_{i}^{T}u)^{2} = (u^{T}x_{i})(x_{i}^{T}u)\) \( \implies\text{Var}(z)=u^{T}\left(\frac{1}{n}\sum _{i=1}^{n}x_{i}x_{i}^{T}\right)u\)
- 💡Since, Covariance Matrix, \(\Sigma = \left(\frac{1}{n}\sum _{i=1}^{n}x_{i}x_{i}^{T}\right)\)
- 👉 Therefore, \(\text{Var}(z)=u^{T}\Sigma u\)
👉 To prevent infinite variance, PCA constrains \(u\) to be a unit vector (\(\|u\|=1\)).
\[\text{maximize\ }u^{T}\Sigma u, \quad \text{subject\ to\ }u^{T}u=1\]Note: This constraint forces the optimization algorithm to focus purely on the direction that maximizes variance, rather than allowing it to artificially inflate the variance by increasing the length of the vector.
⏲️ Lagrangian function: \(L(u,\lambda )=u^{T}\Sigma u-\lambda (u^{T}u-1)\)
🔦 To find \(u\) that maximizes above constrained optimization:
💎 This is the standard Eigenvalue Equation.
🧭 So, the optimal directions \(u\) are the eigenvectors of the covariance matrix \(\Sigma\).
👉 To see which eigenvector maximizes variance, substitute the result back into the variance equation:
\[\text{Variance}=u^{T}\Sigma u=u^{T}(\lambda u)=\lambda (u^{T}u)=\lambda \]🧭 Since the variance equals the eigenvalue \(\lambda\), the direction \(u\) that maximizes variance is the eigenvector associated with the largest eigenvalue.
- Center the data: \(X = X - \mu\)
- Compute the Covariance Matrix: \(\Sigma = \frac{1}{n-1} X^T X\)
- Compute Eigenvectors and Eigenvalues of \(\Sigma\) .
- Sort eigenvalues in descending order and select the top ‘k’ eigenvectors.
- Project the original data onto the subspace: \(Z = X W_k\) where, \(W_{k}=[u_{1},u_{2},\dots ,u_{k}]\) , matrix formed by ‘k’ eigenvectors corresponding to ‘k’ largest eigenvalues.
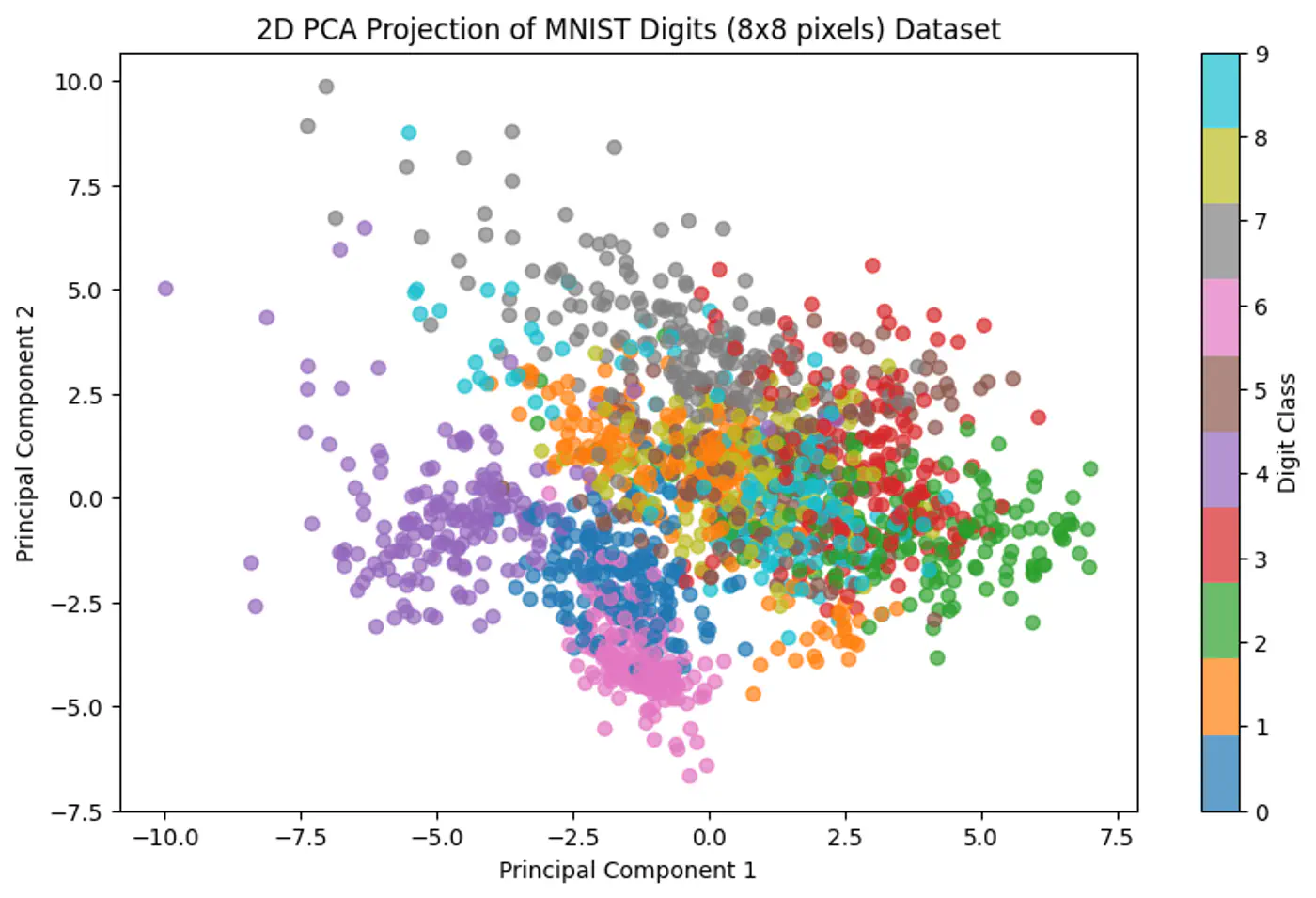
- Cannot model non-linear relationships.
- Sensitive to outliers.
End of Section
1.3.6.2 - t-SNE
👉 PCA preserves variance, not neighborhoods.
🔬 t-SNE focuses on the ‘neighborhood’ (local structure).
💡Tries to keep points that are close together in high-dimensional space close together in low-dimensional space.
👉 Map high-dimensional points to low-dimensional points , such that the pairwise similarities are preserved, while solving the ‘crowding problem’ (where points collapse into a single cluster).
👉 Crowding Problem
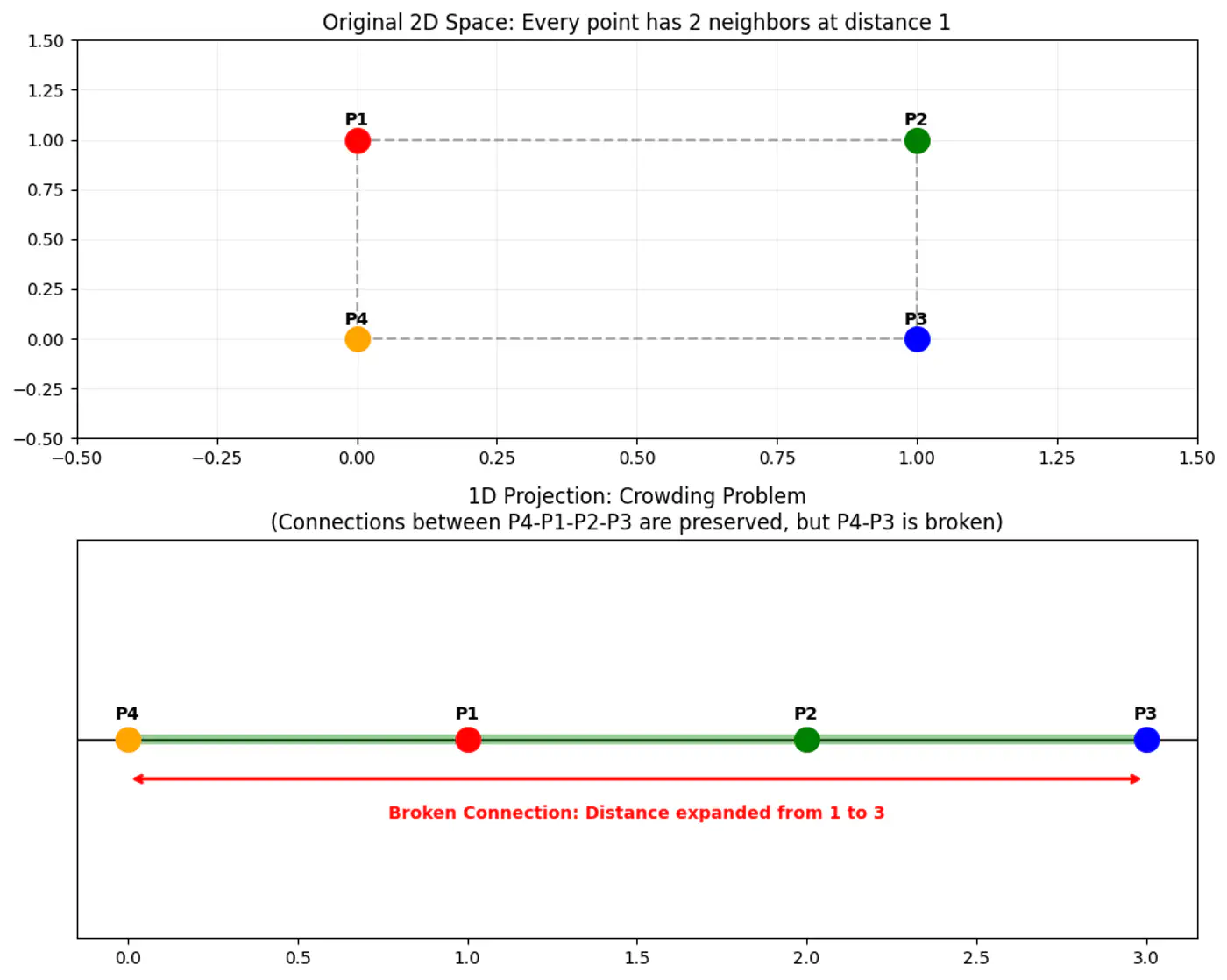
📌 Convert Euclidean distances into conditional probabilities representing similarities.
⚖️ Minimize the divergence between the probability distributions of the high-dimensional (Gaussian)
and low-dimensional (t-distribution) spaces.
Note: Probabilistic approach to defining neighbors is the core ‘stochastic’ element of the algorithm’s name.
💡The similarity of datapoint \(x_j\) to datapoint \(x_i\) is the conditional probability \(p_{j|i}\), that \(x_i\) would pick \(x_j\) as its neighbor.
Note: If neighbors are picked in proportion to their probability density under a Gaussian centered at \(x_i\).
\[p_{j|i} = \frac{\exp(-||x_i - x_j||^2 / 2\sigma_i^2)}{\sum_{k \neq i} \exp(-||x_i - x_k||^2 / 2\sigma_i^2)}\]- \(p_{j|i}\) high: nearby points.
- \(p_{j|i}\) low: widely separated points.
Note: Make probabilities symmetric: \(p_{ij} = \frac{p_{j|i} + p_{i|j}}{2n}\)
🧠 To solve the crowding problem, we use a heavy-tailed 🦨 distribution (Student’s-t distribution with degree of freedom \(\nu=1\)).
\[q_{ij} = \frac{(1 + ||y_i - y_j||^2)^{-1}}{\sum_{k \neq l} (1 + ||y_k - y_l||^2)^{-1}}\]Note: Heavier tail spreads out dissimilar points more effectively, allowing moderately distant points to be mapped further apart, preventing clusters from collapsing and ensuring visual separation and cluster distinctness.
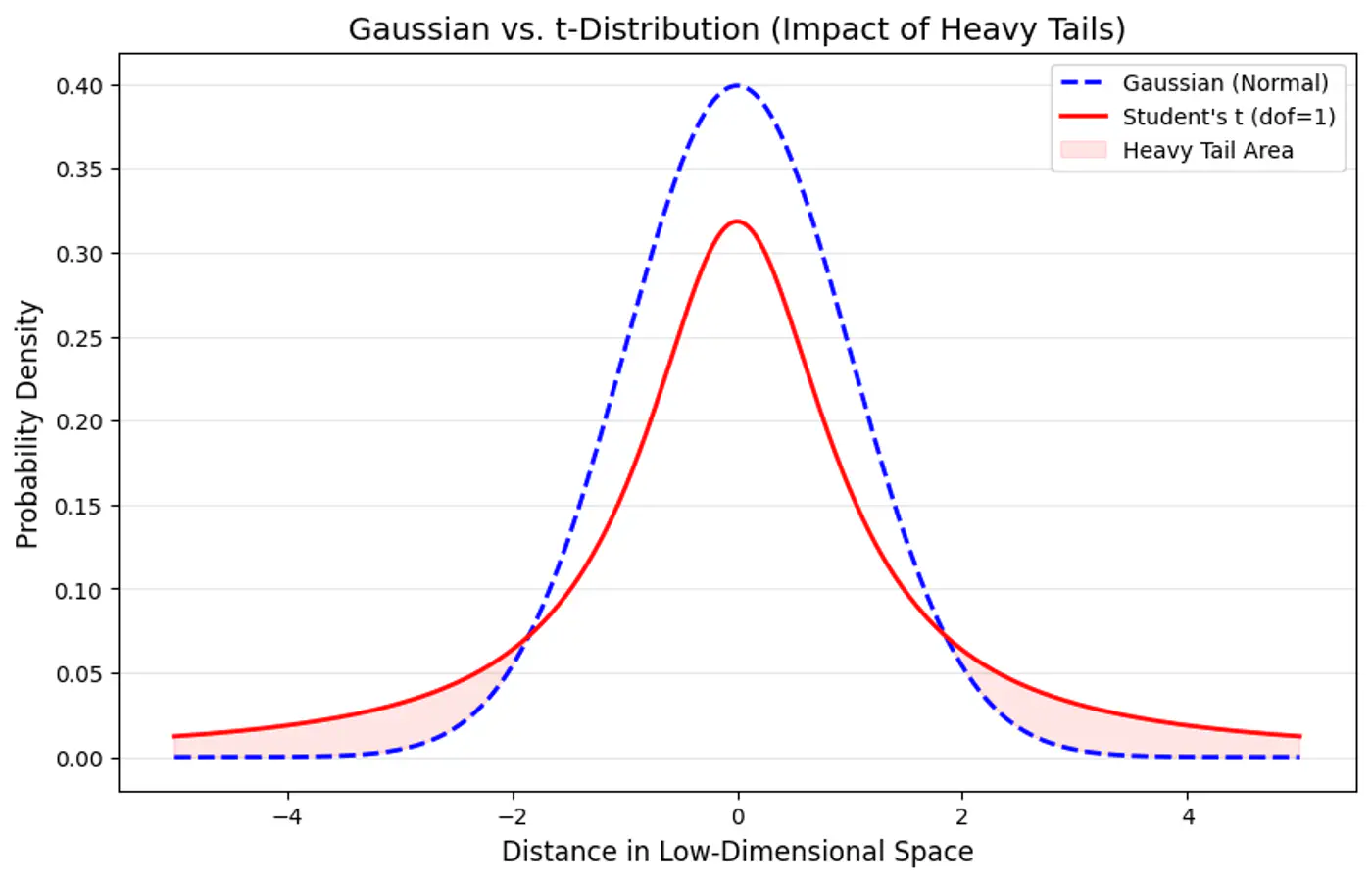
👉 Measure the difference between the distributions ‘p’ and ‘q’ using the Kullback-Leibler (KL) divergence, which we aim to minimize:
\[C = KL(P||Q) = \sum_i \sum_j p_{ij} \log \frac{p_{ij}}{q_{ij}}\]🏔️ Use gradient descent to iteratively adjust the positions of the low-dimensional points \(y_i\).
👉 The gradient of the KL divergence is:
\[\frac{\partial C}{\partial y_{i}}=4\sum _{j\ne i}(p_{ij}-q_{ij})(y_{i}-y_{j})(1+||y_{i}-y_{j}||^{2})^{-1}\]- C : t-SNE cost function (sum of all KL divergences).
- \(y_i, y_j\): coordinates of data points and in the low-dimensional space (typically 2D or 3D).
- \(p_{ij}\): high-dimensional similarity (joint probability) between points \(x_i\) and \(x_j\), calculated using a Gaussian distribution.
- \(q_{ij}\): low-dimensional similarity (joint probability) between points \(y_i\) and \(y_j\), calculated using a Student’s t-distribution with one degree of freedom.
- \((1+||y_{i}-y_{j}||^{2})^{-1}\): term comes from the heavy-tailed Student’s t-distribution, which helps mitigate the ‘crowding problem’ by allowing points that are moderately far apart to have a small attractive force.
💡 The gradient can be understood as a force acting on each point \(y_i\) in the low-dimensional map:
\[\frac{\partial C}{\partial y_{i}}=4\sum _{j\ne i}(p_{ij}-q_{ij})(y_{i}-y_{j})(1+||y_{i}-y_{j}||^{2})^{-1}\]- Attractive forces: If \(p_{ij}\) is large ⬆️ and \(q_{ij}\) is small ⬇️ (meaning two points were close in the high-dimensional space but are far in the low-dimensional space), the term \((p_{ij}-q_{ij})\) is positive, resulting in a strong attractive force pulling \(y_i\) and \(y_j\) closer.
- Repulsive forces: If \(p_{ij}\) is small ⬇️ and \(q_{ij}\) is large ⬆️ (meaning two points were far in the high-dimensional space but are close in the low-dimensional space), the term \((p_{ij}-q_{ij})\) is negative, resulting in a repulsive force pushing \(y_i\) and \(y_j\) apart.
👉 Update step of in low dimensions:
\[y_{i}^{(t+1)}=y_{i}^{(t)}-\eta \frac{\partial C}{\partial y_{i}}\]Attractive forces (\(p_{ij}>q_{ij}\)):
- The negative gradient moves \(y_i\) opposite to the (\(y_i - y_j\)) vector, pulling it toward \(y_j\).
Repulsive forces (\(p_{ij} < q_{ij}\)):
- The negative gradient moves \(y_i\) in the same direction as the (\(y_i - y_j\)) vector, pushing it away from \(y_j\).
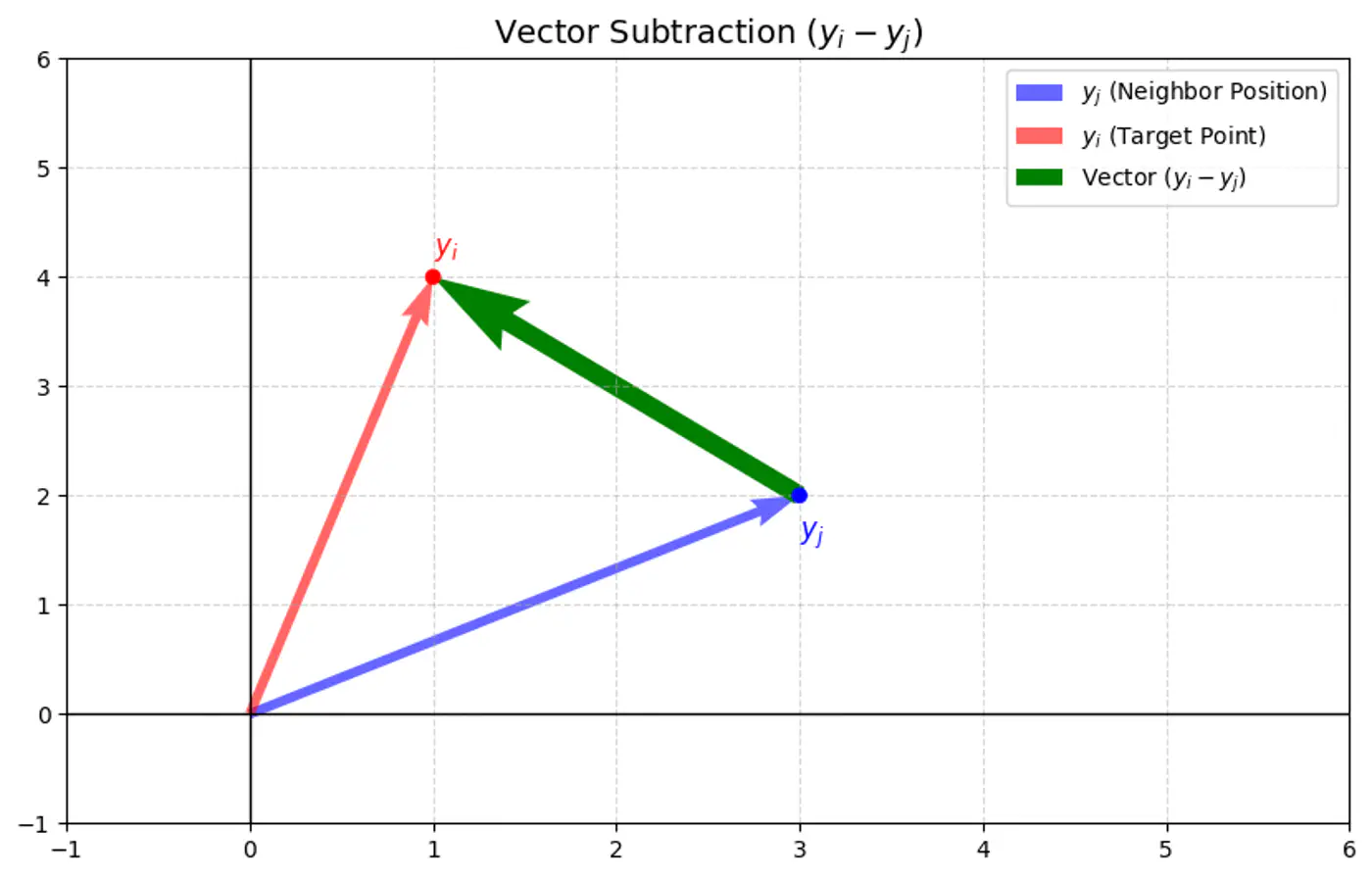
🏘️ User-defined parameter that loosely relates to the effective number of neighbors.
Note: Variance \(\sigma_i^2\) (Gaussian) is adjusted for each point to maintain a consistent perplexity.
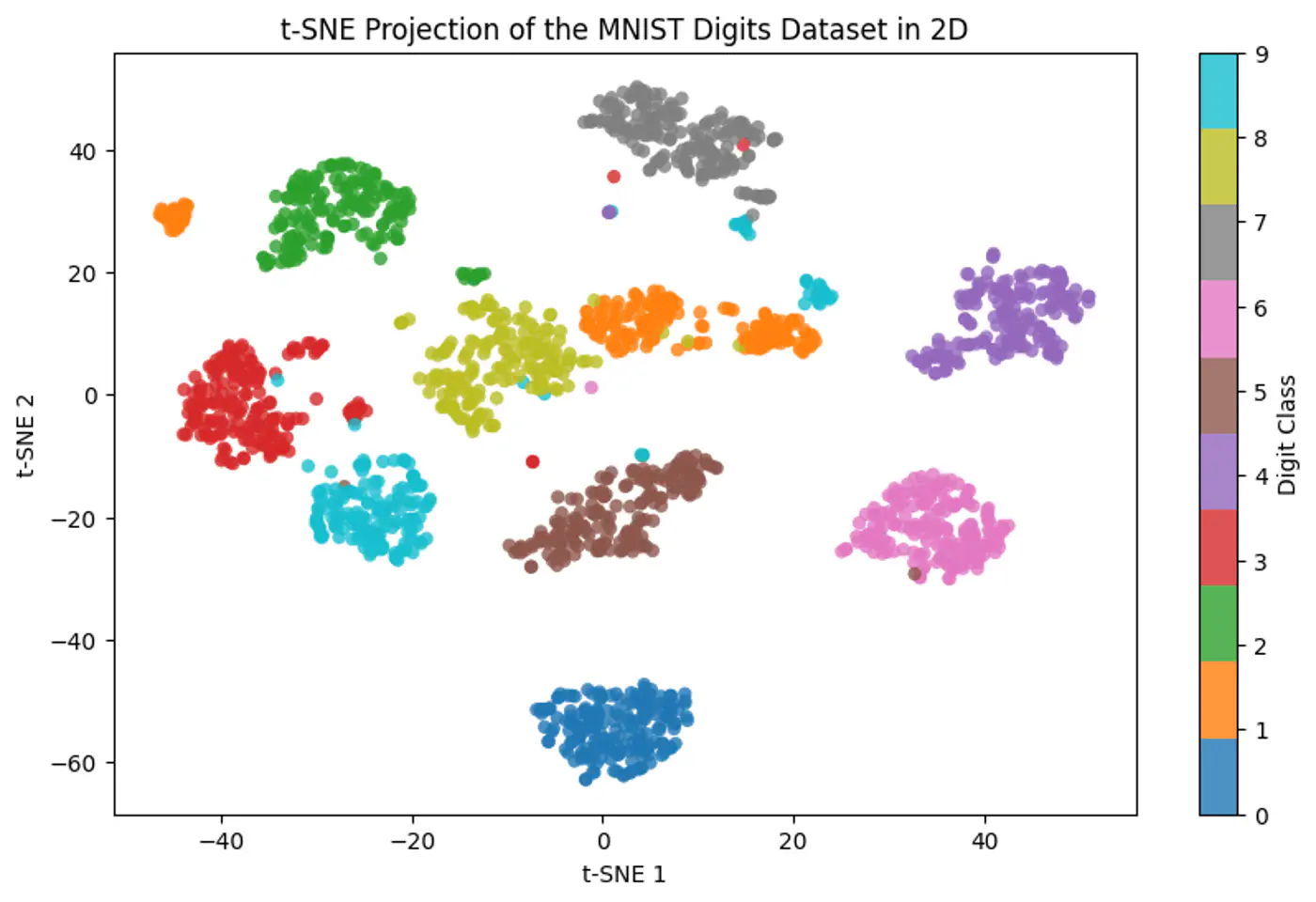
End of Section
1.3.6.3 - UMAP
🐢Visualizing massive datasets where t-SNE is too slow.
⭐️ Creating robust low-dimensional inputs for subsequent machine learning models.
⭐️ Using a world map 🗺️ (2D)instead of globe 🌍 for spherical(3D) earth 🌏.
👉It preserves the neighborhood relationships of countries (e.g., India is next to China), and to a good degree, the global structure.
⭐️ Non-linear dimensionality reduction technique to visualize high-dimensional data (like images, gene expressions)
in a lower-dimensional space (typically 2D or 3D), preserving its underlying structure and relationships.
👉 Constructs a high-dimensional graph of data points and then optimizes a lower-dimensional layout to closely match this graph, making complex datasets understandable by revealing patterns, clusters, and anomalies.
Note: Similar to t-SNE but often faster and better at maintaining global structure.
💡 Create a weighted graph (fuzzy simplicial set) representing the data’s topology and then find a low-dimensional graph that is as structurally similar as possible.
Note: Fuzzy means instead of using binary 0/1, we use weights in the range [0,1] for each edge.
- UMAP determines local connectivity based on a user-defined number of neighbors (n_neighbors).
- Normalizes distances locally using the distance to the nearest neighbor (\(\rho_i\)) and a scaling factor (\(\sigma_i\)) adjusted to enforce local connectivity constraints.
- The weight \(W_{ij}\) (fuzzy similarity) in high-dimensional space is: \[W_{ij}=\exp \left(-\frac{\max (0,\|x_{i}-x_{j}\|-\rho _{i})}{\sigma _{i}}\right)\]
Note: This ensures that the closest point always gets a weight of 1, preserving local structure.
👉In the low-dimensional space (e.g., 2D), UMAP uses a simple curve (similar to the t-distribution used in t-SNE) for edge weights:
\[Z_{ij}=(1+a\|y_{i}-y_{j}\|^{2b})^{-1}\]Note: The parameters ‘a’ and ‘b’ are typically fixed based on the ‘min_dist’ user parameter (e.g. min_dist = 0.1, then a≈1,577, b≈0.895).
⭐️ Unlike t-SNE’s KL divergence, UMAP minimizes the cross-entropy between the high-dimensional weights \(W_{ij}\) and the low-dimensional weights \(Z_{ij}\).
👉 Cost 💰 Function (C):
\[C=\sum _{i,j}\left(W_{ij}\log \frac{W_{ij}}{Z_{ij}}+(1-W_{ij})\log \frac{1-W_{ij}}{1-Z_{ij}}\right)\]Cost 💰 Function (C):
\[C=\sum _{i,j}\left(W_{ij}\log \frac{W_{ij}}{Z_{ij}}+(1-W_{ij})\log \frac{1-W_{ij}}{1-Z_{ij}}\right)\]🎯 Goal : Reduce overall Cross Entropy Loss.
- Attractive Force: \(W_{ij}\) high; make \(Z_{ij}\) high to make the log \(\frac{W_{ij}}{Z_{ij}} \)term small.
- Repulsive Force: \(W_{ij}\) low; make \(Z_{ij}\) low to make the \(log\frac{1-W_{ij}}{1-Z_{ij}}\) term small.
Note: This push and pull of 2 ‘forces’ will make the data in low dimensions settle into a position that is overall a good representation of the original data in higher dimensions.
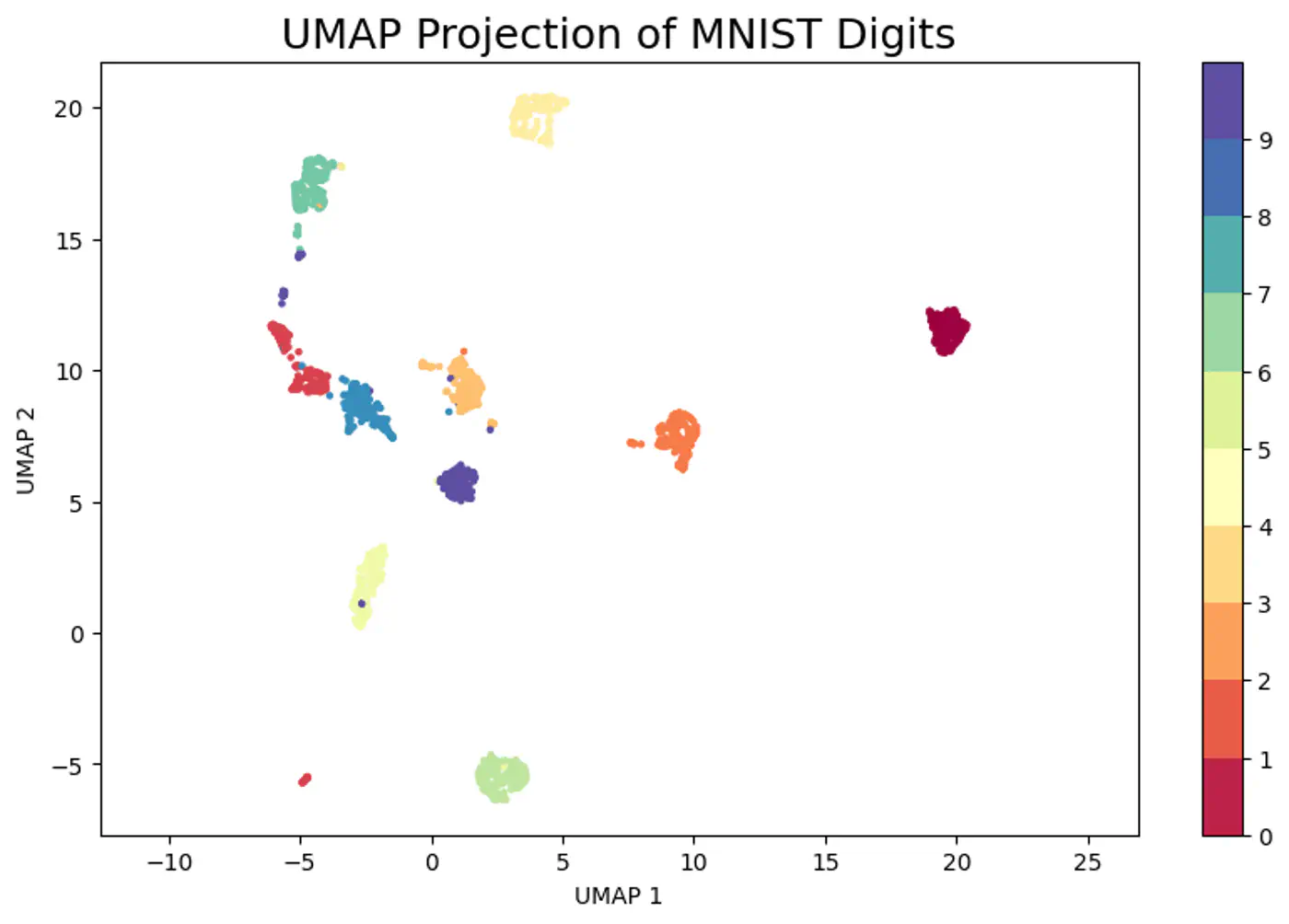
- Mathematically complex.
- Requires tuning (n_neighbors, min_dist).
End of Section
1.4.1 - Data Pre Processing
Messy and Incomplete.
We need to pre-process the data to make it:
- Clean
- Consistent
- Mathematically valid
- Computationally stable
👉 So that, the machine learning algorithm can safely consume the data.
- Missing Completely At Random (MCAR)
- The missingness occurs entirely by chance, such as due to a technical glitch during data collection or a random human error in data entry.
- Missing At Random (MAR)
- The probability of missingness depends on the observed data and not on the missing value itself.
- e.g. In some survey, the age of many females are missing, because they may not like to disclose the information.
- Missing Not At Random (MNAR)
- The probability of missingness is directly related to the unobserved missing value itself.
- e.g. Individuals with very high incomes 💰may intentionally refuse to report their salary due to privacy concerns, making the missing data directly dependent on the high income 💰value itself.
- Simple Imputation:
- Mean: Normally distributed numerical features.
- Median: Skewed numerical features.
- Mode: Categorical features, most frequent.
- KNN Imputation:
- Replace the missing value with mean/median/mode of ‘k’ nearest (similar) neighbors of the missing value.
- Predictive Imputation:
- Use another ML model to estimate missing values.
- Multivariate Imputation by Chained Equations (MICE):
- Iteratively models each variable with missing values as a function of other variables using flexible regression models (linear regression, logistic regression, etc.) in a ‘chained’ or sequential process.
- Creates multiple datasets, using slightly different random starting points.
🦄 Outliers are extreme or unusual data points, can mislead models, causing inaccurate predictions.
- Remove invalid or corrupted data.
- Replace (Impute): Median or capped value to reduce impact.
- Transform: Apply log or square root to reduce skew.
👉 For example: Log and Square Root Transformed Data
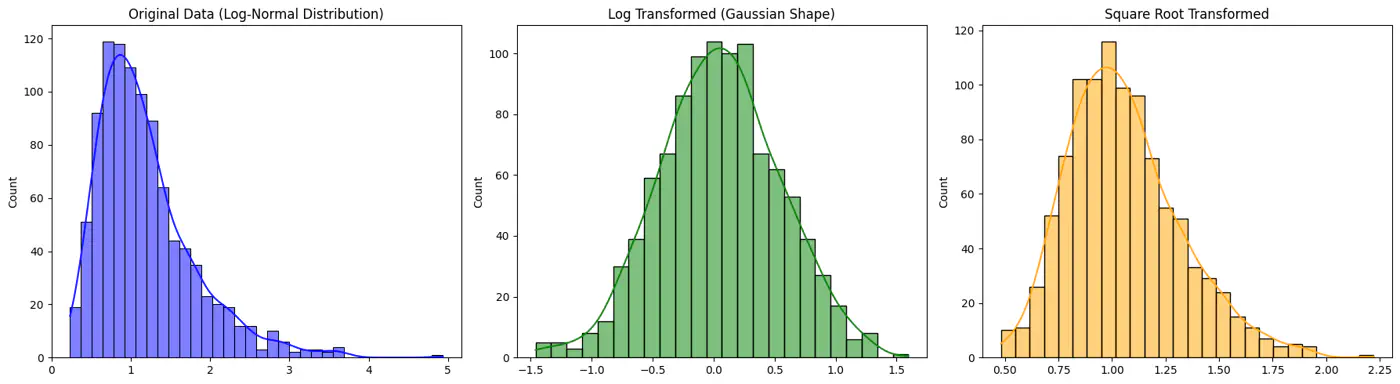
💡 If one feature ranges from 0-1 and another from 0-1000, larger feature will dominate the model.
- Standardization (Z-score) :
- μ=0, σ=1; less sensitive to outliers.
- \(x_{std} = (x − μ) / σ\)
- Min-Max Normalization:
- Maps data to specific range, typically [0,1]; sensitive to outliers.
- \(x_{minmax} = (x − min) / (max − min)\)
- Robust Scaling:
- Transforms features using median and IQR; resilient to outliers.
- \(x_{scaled}=(x-\text{median})/\text{IQR}\)
👉 Standardization Example
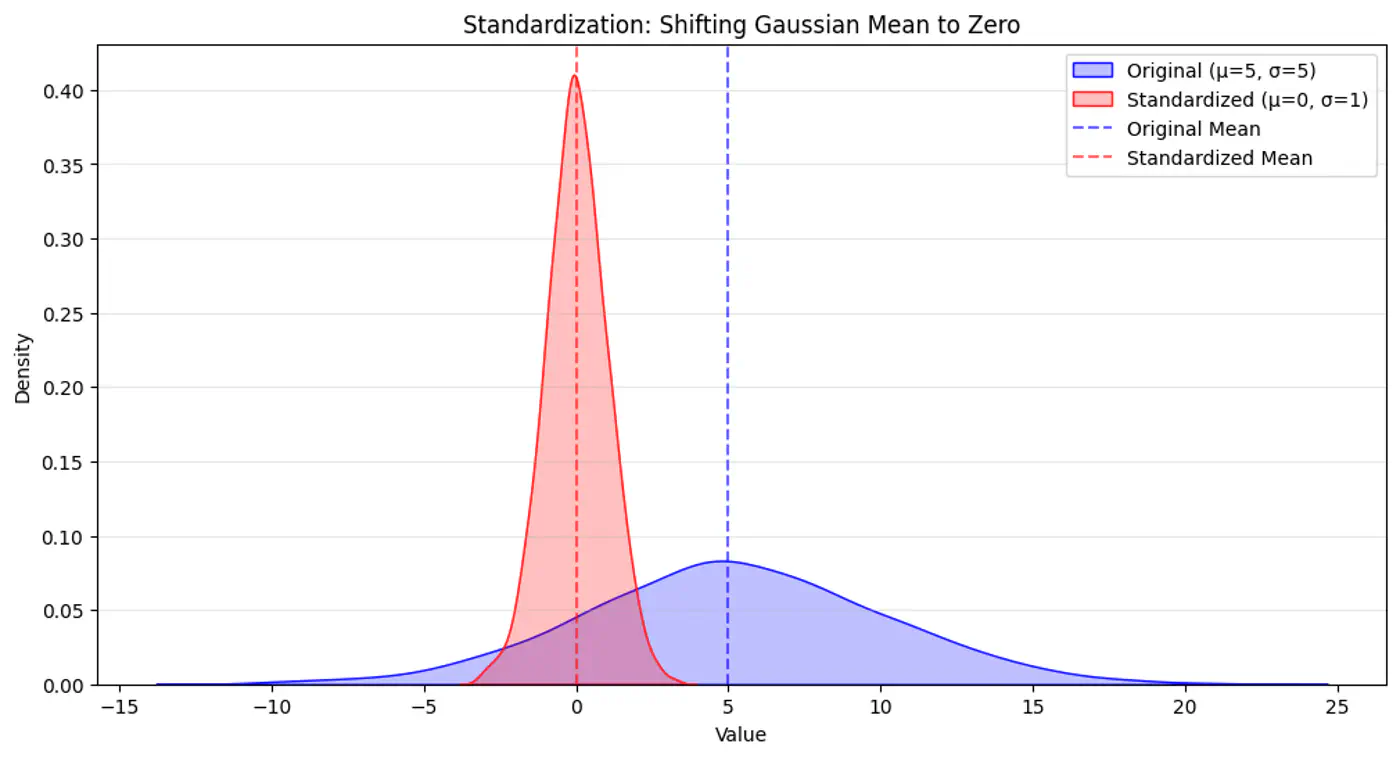
End of Section
1.4.2 - Categorical Variables
💡 ML models operate on numerical vectors.
👉 Categorical variables must be transformed (encoded) while preserving information and semantics.
- One Hot Encoding (OHE)
- Label Encoding
- Ordinal Encoding
- Frequency/Count Encoding
- Target Encoding
- Hash Encoding
⭐️ When the categorical data (nominal) is without any inherent ordering.
- Create binary columns per category.
- e.g.: Colors: Red, Blue, Green.
- Colors: [1,0,0], [0,1,0], [0,0,1]
Note: Use when low cardinality, or small number of unique values (<20).
⭐️ Assigns a unique integer (e.g., 0, 1, 2) to each category.
- When to use ?
- Target variable, i.e, unordered (nominal) data, in classification problems.
- e.g. encoding a city [“Paris”, “Tokyo”, “Amsterdam”] -> [1, 2, 0], (Alphabetical: Amsterdam=0, Paris=1, Tokyo=2).
- When to avoid ?
- For nominal data in linear models, because it can mislead the model to assume an order/hierarchy, when there is none.
⭐️ When categorical data has logical ordering.
Best for: Ordered (ordinal) input features.
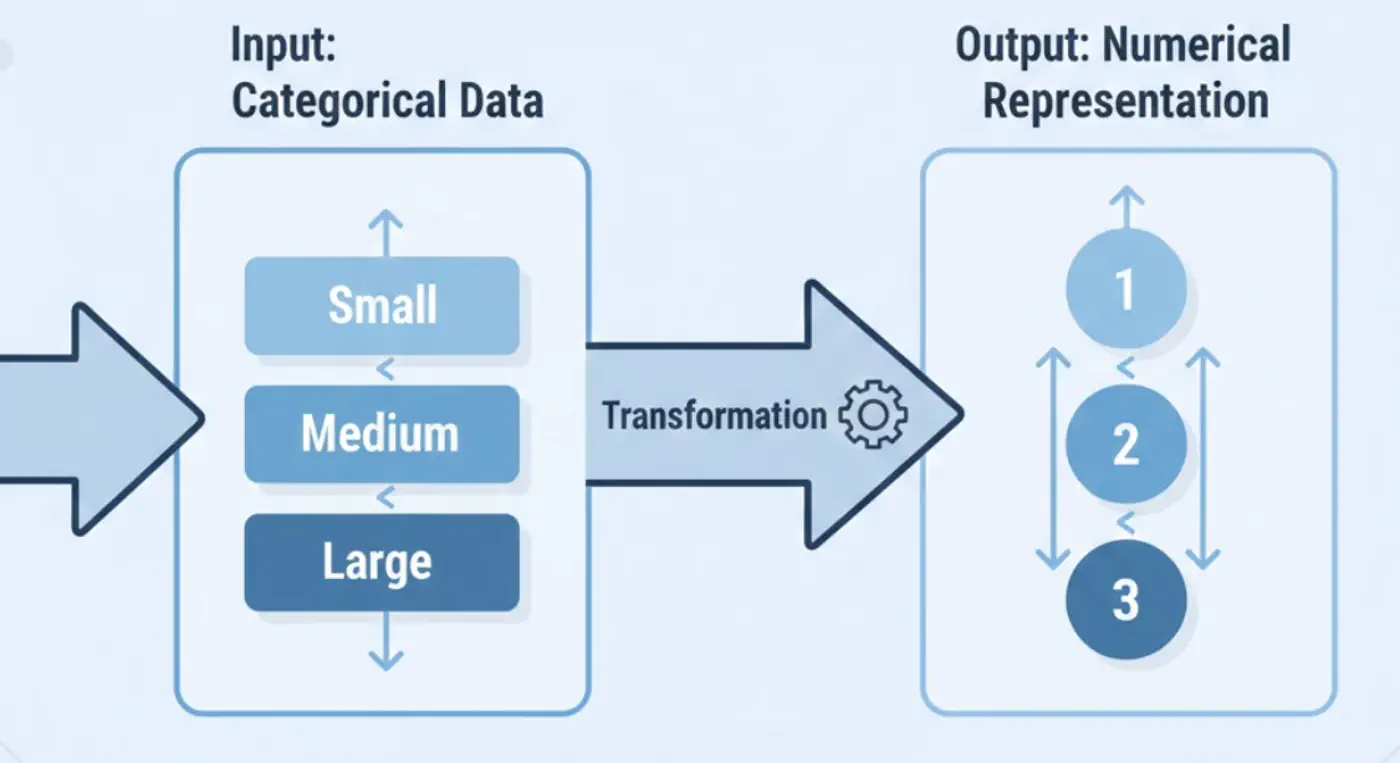
⭐️ Replace categories with their frequency or count in the dataset.
- Useful for high-cardinality features where many unique values exist.
👉 Example
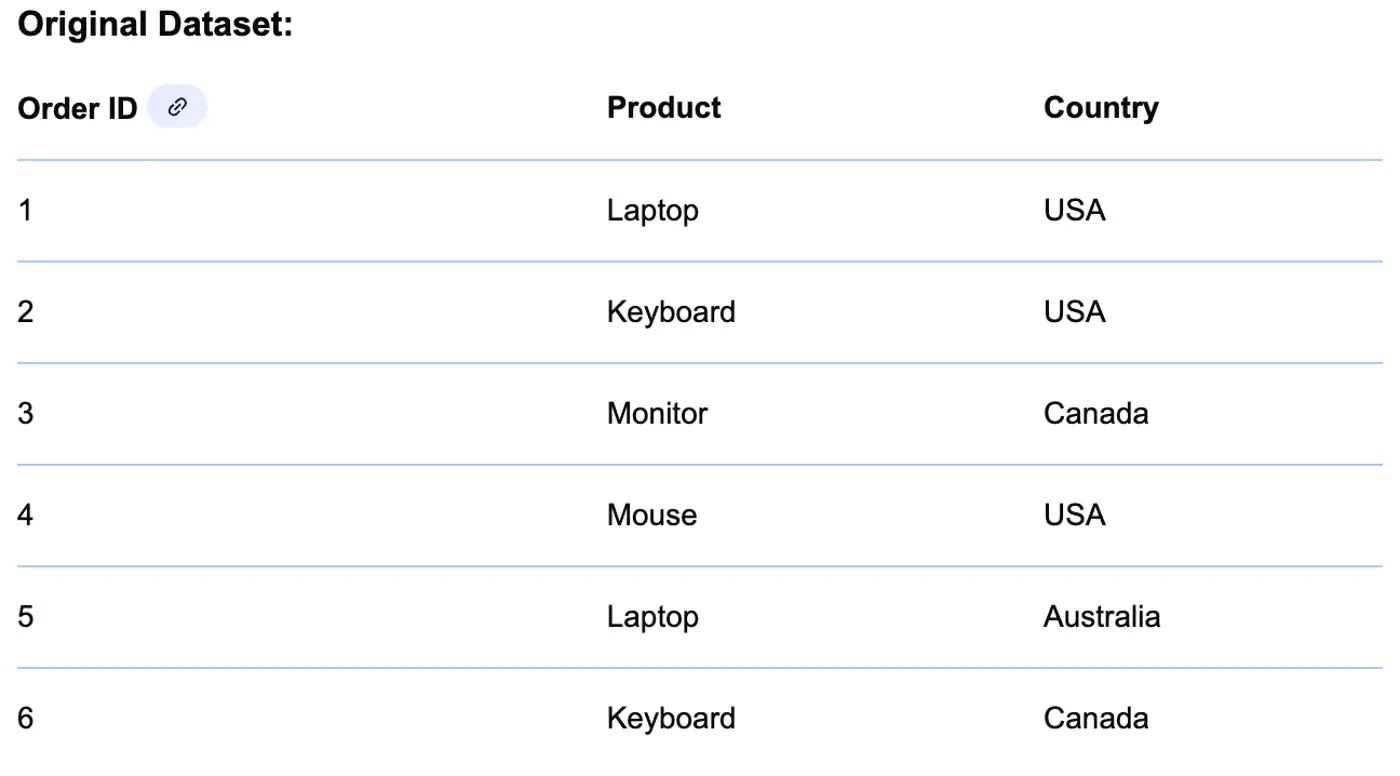
👉 Frequency of Country

👉 Country replaced with Frequency
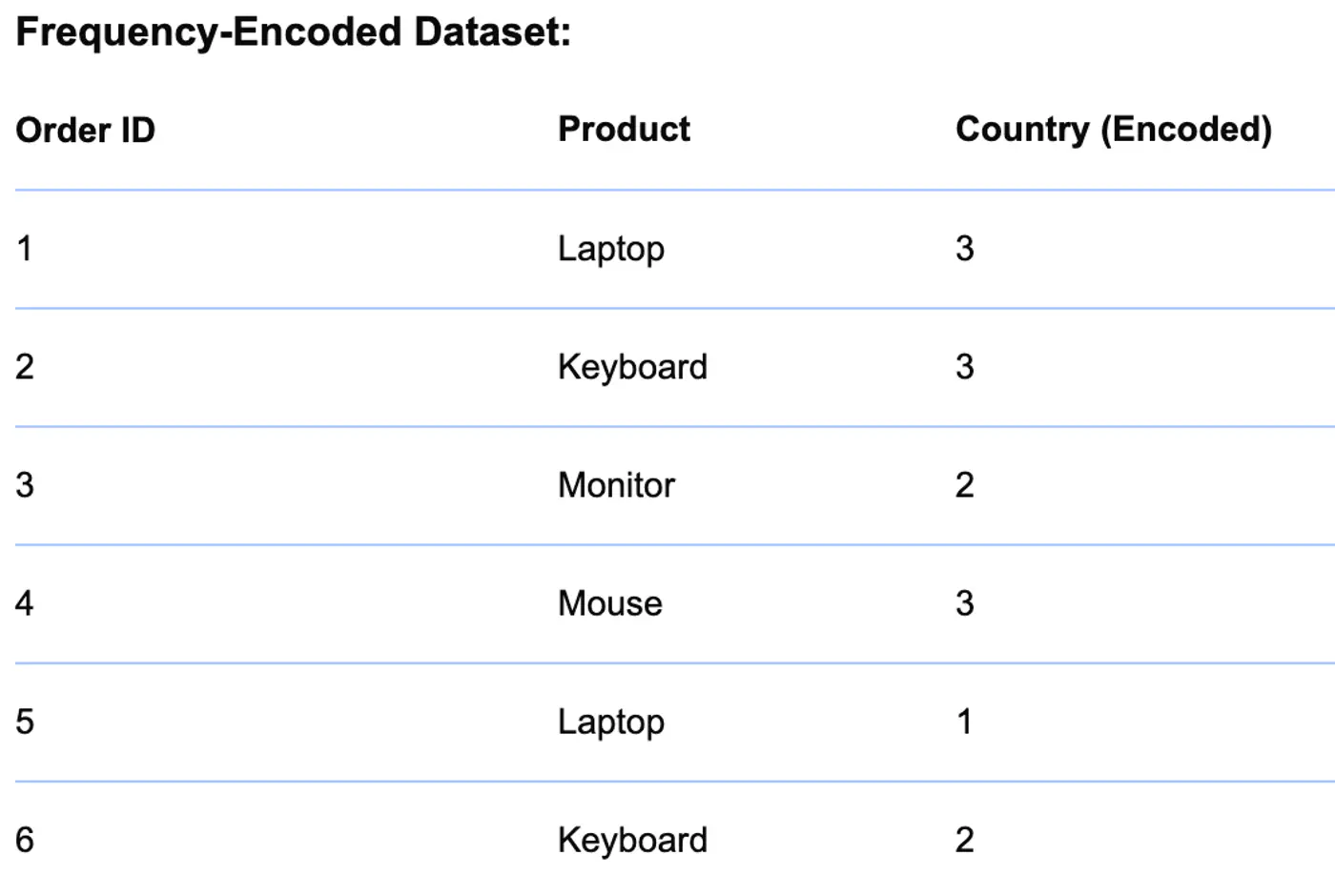
⭐️ Replace a category with the mean of the target variable for that specific category.
- When to use ?
- For high-cardinality nominal features, where one hot encoding is inefficient, e.g., zip code, product id, etc.
- Strong correlation between the category and the target variable.
- When to avoid ?
- With small datasets, because the category averages (encodings) are based on too few samples, making them unrepresentative.
- Also, it can lead to target leakage and overfitting unless proper smoothing or cross-validation techniques (like K-fold or Leave-One-Out) are used.
⭐️ Maps categories to a fixed number of features using a hash function.
Useful for high-cardinality features where we want to limit the dimensionality.
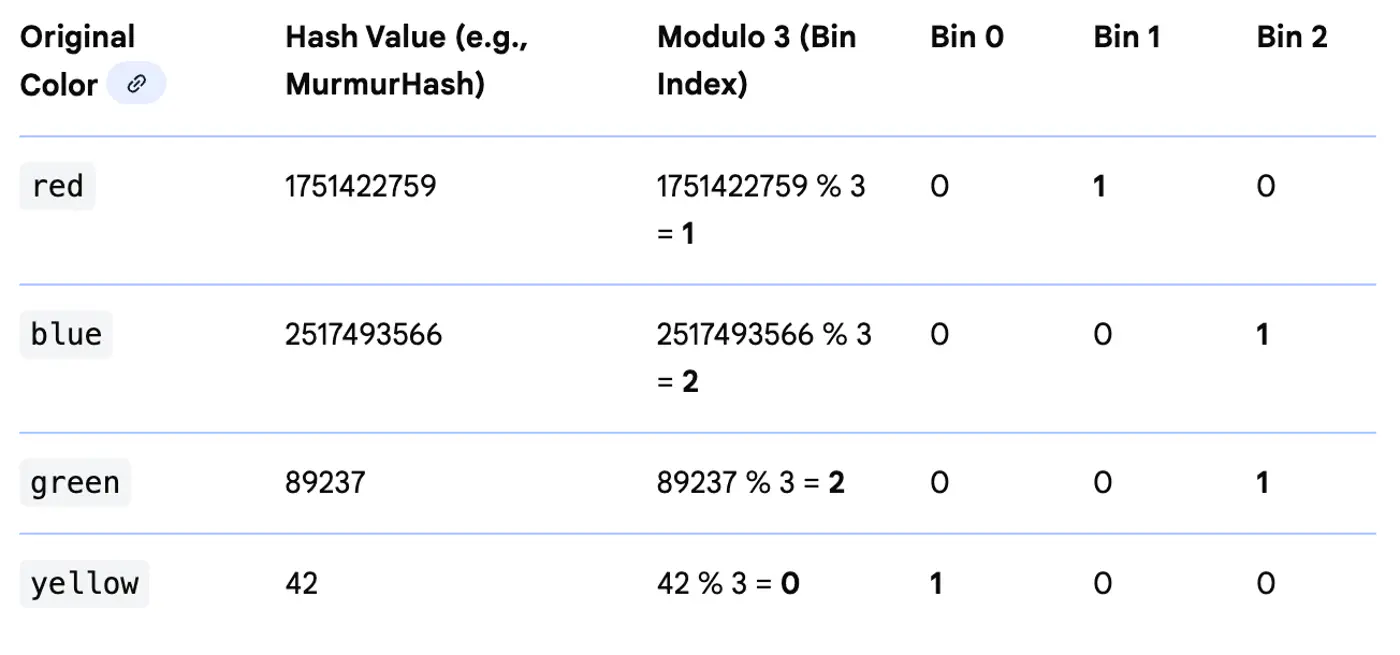
End of Section
1.4.3 - Feature Engineering
Create polynomial features, such as, x^2, x^3, etc., to learn non-linear relationship.
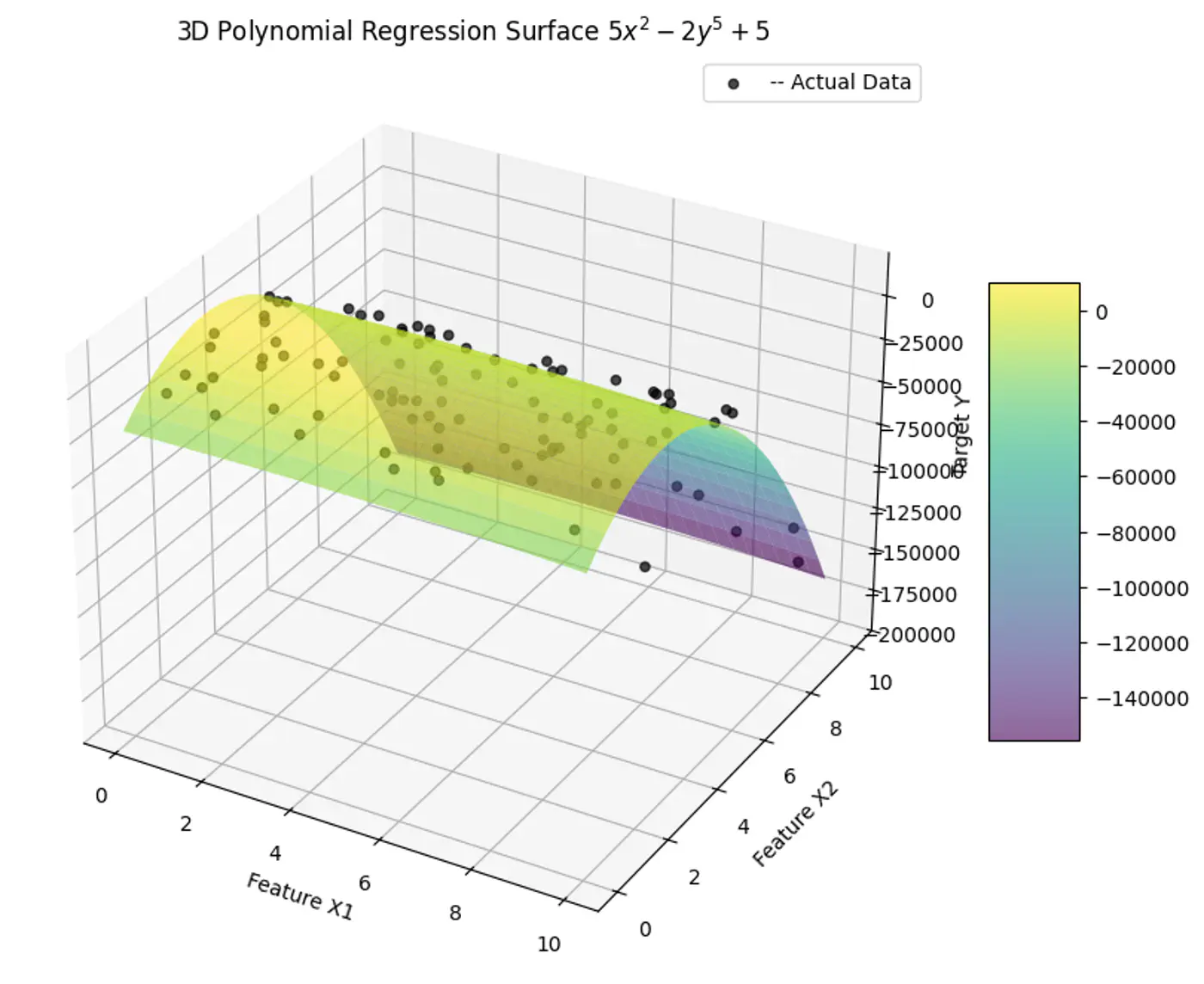
⭐️ Combine 2 or more features to capture non-linear relationship.
- e.g. combine latitude and longitude into one location feature ‘lat-long'.
⭐️ Memory-efficient 🧠 technique to convert categorical (string) data into a fixed-size numerical feature vector.
- Pros:
- Useful for high-cardinality features where we want to limit the dimensionality.
- Cons:
- Hash collisions.
- Reduced interpretability.
👉 Hash Encoding (Example)
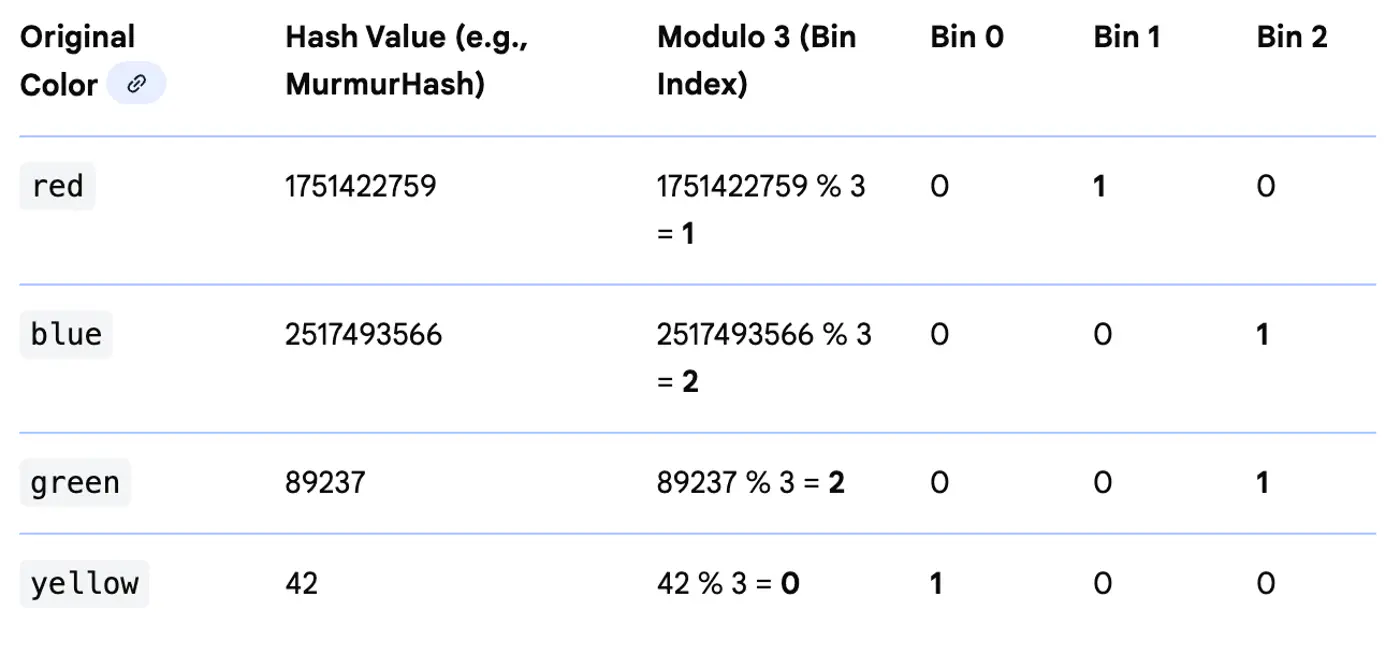
⭐️ Group continuous numerical values into discrete categories or ‘bin’.
- e.g. divide age into groups 18-24, 25-35, 35-45, 45-55, >55 years etc.
End of Section
1.4.4 - Data Leakage
⭐️ Occurs when a model is trained using data that would not be available during real-world predictions,
leading to good training performance, but poor real‑world 🌎 performance.
It is essentially the model ‘cheating’ by inadvertently accessing information about the target variable.
👉Any information from the validation/test set must NOT influence training, directly or indirectly.
❓So, how do we prevent this leakage of information or data leakage from training to validation or test set ?
- ❌ Wrong: Applying preprocessing (like global StandardScaler, Mean_Imputation, Target_Encoding etc.) on the entire dataset before splitting.
- ✅ Right: Compute mean, variance, etc. only on the training data and use the same for validation and test data.
Preventing Leakage in Cross-Validation:
- ❌ Wrong: Perform preprocessing (e.g., scaling, normalization, missing value imputation) on the entire dataset before passing it to cross_val_score.
- ✅ Right: Use sklearn.pipeline.Pipeline; Pipeline ensures that the ‘validation fold’ remains unseen until the transformation is applied using the training fold’s parameters.
This happens in Time Series ⏰ data.
- ❌ Wrong: Use standard random CV; it allows the model to ‘peek into the future’.
- ✅ Right: Use Time-Series Nested Cross-Validation (Forward Chaining) instead of random shuffling.
- ❌ Wrong: Include features that are only available after the event we are trying to predict and are proxy for the target.
- e.g. Including number_of_late_payments in a model to predict whether a person applying for a bank loan will default ?
- ✅ Right: Do not include such features during training.
Group Leakage:
- ❌ Wrong: If you have multiple rows that are correlated (same user).
- For the same patient or user, you put some rows in Train and others in Test.
- ✅ Right: Use GroupKFold to ensure all data from a specific group stays together in one fold.
End of Section
1.4.5 - Model Interpretability
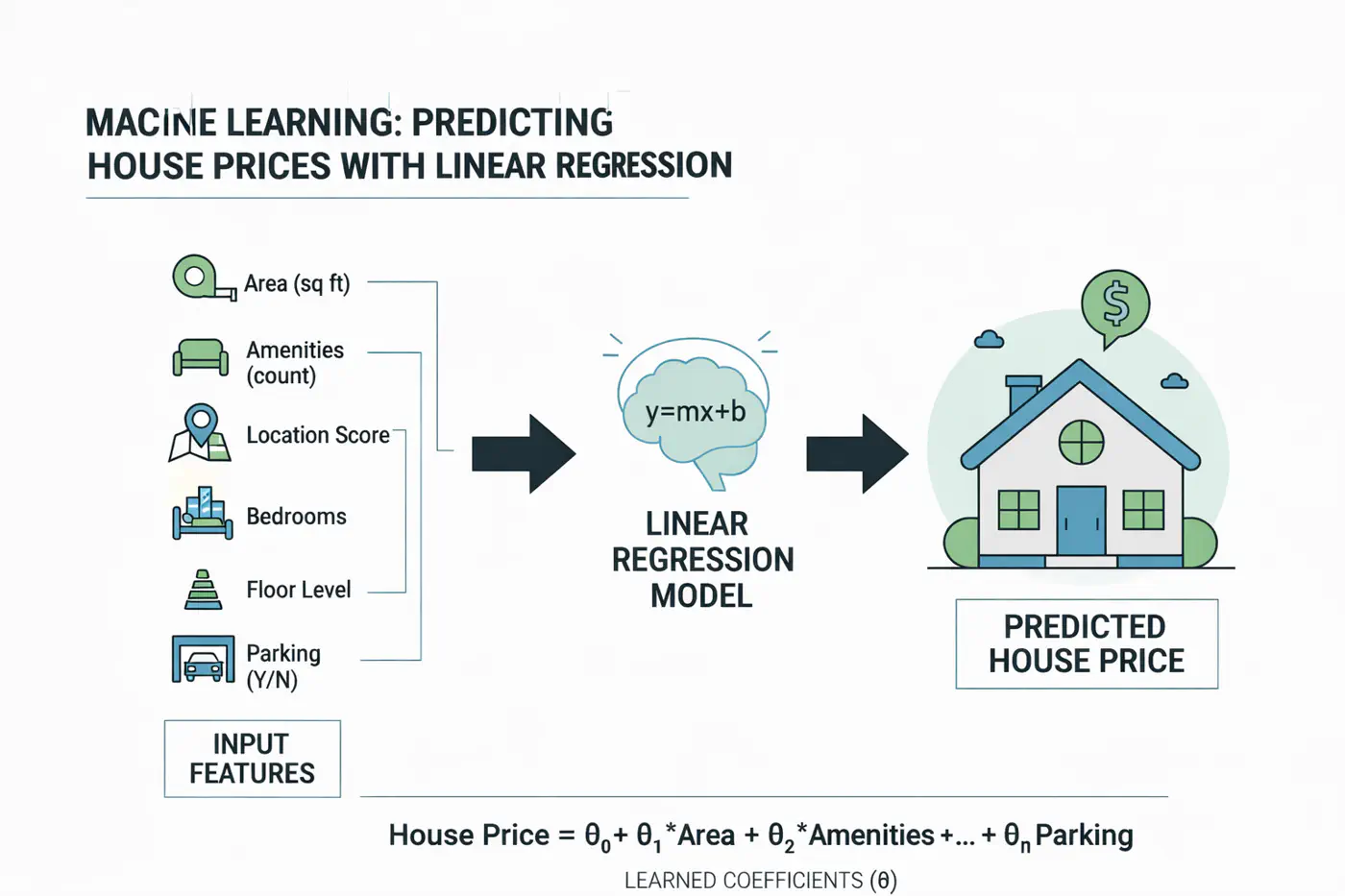
👉 Because without this capability the machine learning is like a black box to us.
👉 We should be able to answer which features had most influence on output.
⭐️ Let’s understand ‘Feature Importance’ and why the ML model output’s interpretability is important ?
Note: Weights 🏋️♀️ represent the importance of feature with standardized data.
💡 Overall model behavior + Why this prediction?
- Trust: Stakeholders must trust predictions.
- Model Debuggability: Detect leakage, spurious correlations.
- Feature engineering: Feedback loop.
- Regulatory compliance: Data privacy, GDPR.
⭐️ Stakeholders Must Trust Predictions.
- Users, executives, and clients are more likely to trust and adopt an AI system if they understand its reasoning.
- This transparency is fundamental, especially in high-stakes applications like healthcare, finance, or law, where decisions can have a significant impact.
⭐️ In many industries, regulations mandate the ability to explain decisions made by automated systems.
- For instance, the General Data Protection Regulation (GDPR) in Europe includes a “right to explanation” for individuals affected by algorithmic decisions.
- Interpretability ensures that organizations can meet these legal and ethical requirements.
End of Section
1.5.1 - Data Distribution Shift
- P(X | Y): Likelihood of X given Y (joint distribution)
- P(Y | X) : Model (Posterior)
- P(Y): Prior probability of the output Y.
- P(X): Evidence (marginal probability of the input X).
⭐️The input data distribution seen during training is different from the distribution seen during inference.
👉 P(X)(input) changes, but P(Y|X) (model) remains same.
- e.g. Self-driving car 🚗 trained on a bright, sunny day is used during foggy winter.
⭐️The output distribution changes, but for a given output, the input distribution remains the same.
👉 P(Y) (output) changes, but P(X|Y) remains the same.
- 😷 e.g. Flu-detection model is trained during summer, when only 1% of patients have flu.
- The same model is used during winter when 40% of patients have flu.
- 🍎 Prior probability of having flu P(Y) has changed from 1% to 40%, but the symptoms for a person to have flu P(X|Y) remains same.
⭐️ The relationship between inputs and outputs changes.
i.e the very definition of what you are trying to predict changes.
👉 Concept drifts are cyclic or seasonal.
- e.g. ‘Normal’ spending behavior in 2019 became ‘Abnormal’ during 2020 lockdowns 🔐.
End of Section
1.5.2 - Retraining Strategies
⭐️In a production ML environment, retraining is the ‘maintenance engine’ ⚙️ that keeps our models from becoming obsolete.
❌ Don’t ask: When do we retrain?
✅ Ask: “How do we automate the decision to retrain while balancing compute cost 💰, model risk, and data freshness?”
👉 The model is retrained on a regular schedule (e.g., daily, weekly, or monthly).
- Best for:
- Stable environments where data changes slowly.
(e.g. long-term demand forecast or a credit scoring model).
- Stable environments where data changes slowly.
- Pros:
- Highly predictable; easy to schedule compute resources; simple to implement via a cron job or Airflow DAG.
- Cons:
- Inefficient. You might retrain when not needed (wasting money 💵) or fail to retrain during a sudden market shift (losing accuracy).
👉 Retraining is initiated only when a specific performance or data metric crosses a pre-defined threshold.
- Metric Triggers:
- Performance Decay: A drop in Precision, Recall, or RMSE (requires ground-truth labels).
- Drift Detection: A high PSI (Population Stability Index) or K-S test score indicating covariate shift.
- Pros:
- Cost-effective; reacts to the ‘reality’ of the data rather than the calendar.
- Cons:
- Requires a robust monitoring stack 📺.
If the ‘trigger’ logic is buggy, the model may never update.
- Requires a robust monitoring stack 📺.
👉 Instead of retraining from scratch on a massive batch, the model is updated incrementally as new data ‘streams’ into the system.
- Mechanism: Using ‘Warm Starts’ where the model weights from the previous version are used as the starting point for the next few gradient descent steps.
- Best for:
- Recommendation engines (Netflix/TikTok) or High-Frequency Trading 💰where patterns change by the minute.
- Pros:
- Extreme ‘freshness’; low latency between data arrival and model update.
- Cons:
- High risk of ‘Catastrophic Forgetting’ (the model forgets old patterns) and high infrastructure complexity.
End of Section
1.5.3 - Deployment Patterns
⭐️In a production ML environment, retraining is only half the battle, we must also safely deploy the new version.
Types of deployment (most common):
- Shadow ❏ Deployment
- A/B Testing 🧪
- Canary 🦜 Deployment
👉 Safest way to deploy our model or any software update.
- Deploy the candidate model in parallel with the existing model.
- For each incoming request, route it to both models to make predictions, but only serve the existing model’s prediction to the user.
- Log the predictions from the new model for analysis purposes.
Note: When the new model’s predictions are satisfactory, we replace the existing model with the new model.
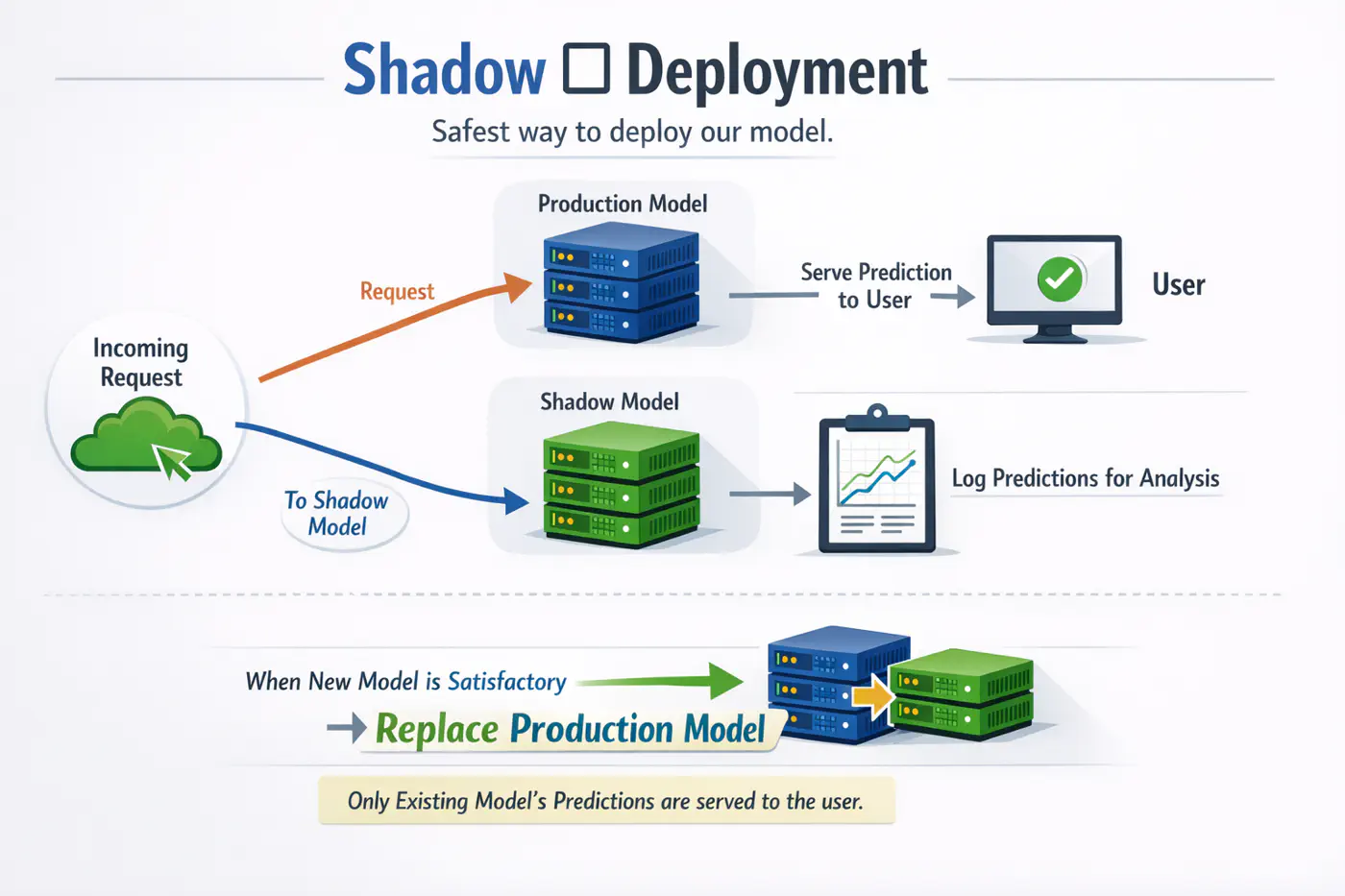
👉A/B testing is a way to compare two variants of a model.
- Deploy the candidate model in parallel with the existing model.
- A percentage of traffic🚦is routed to the candidate for predictions; the rest is routed to the existing model for predictions.
- Monitor 📺 and analyze the predictions, from both models to determine whether the difference in the two models’ performance is statistically significant.
Note: Say we run a two-sample test and get the result that model A is better than model B with the p-value of p = 0.05 or 5%.
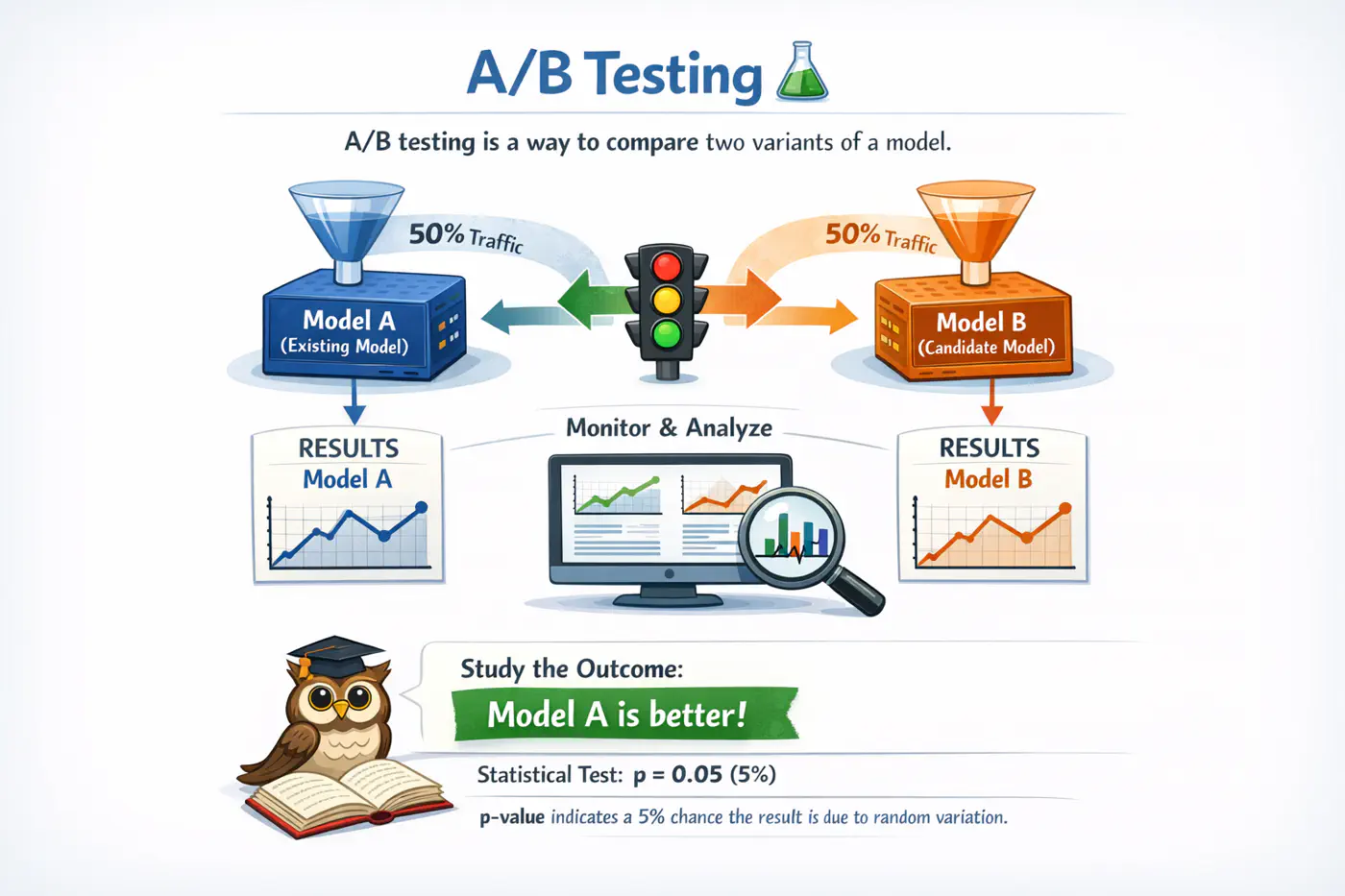
👉 Mitigates deployment risk by incrementally shifting traffic 🚦from a model version to a new version, allowing for real-world validation on a subset of users before a full-scale rollout.
- Deploy the candidate model in parallel with the existing model.
- A percentage of traffic🚦is routed to the candidate for predictions.
- If its performance is satisfactory, increase the traffic to the candidate model.If not, abort the canary and route all the traffic🚦 back to the existing model.
- Stop when either the canary serves all the traffic🚦 (the candidate model has replaced the existing model) or when the canary is aborted.
Note: Canary releases can be used to implement A/B testing due to the similarities in their setups. However, we can do canary analysis without A/B testing.
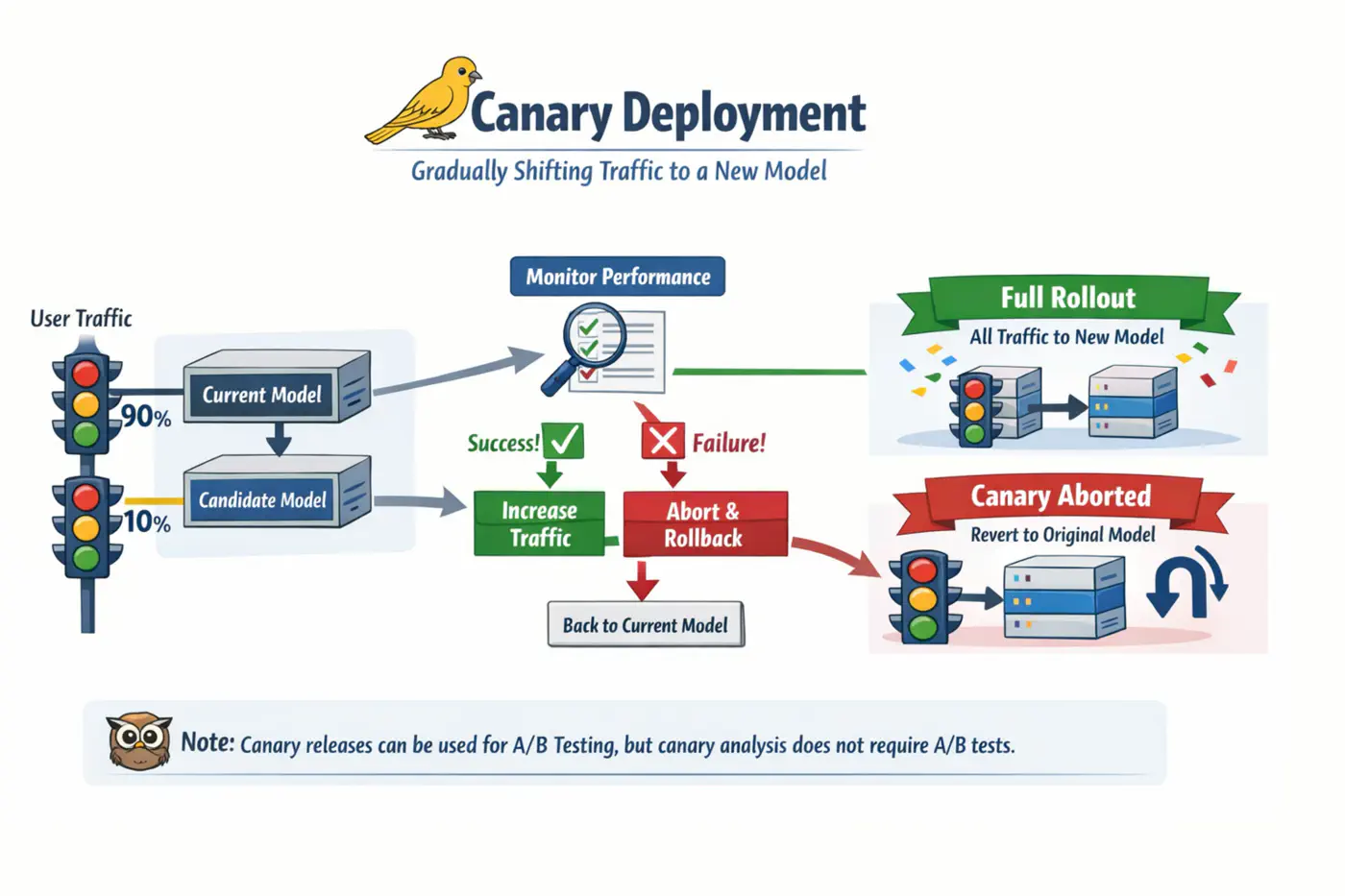
End of Section
1.6 - Interview Questions
Which metrics would you report and why?
We should evaluate an imbalanced classification model using metrics that that focus on performance for each class, especially the minority class.
Why ?
Say, we have a dataset with high imbalance, i.e, 99% of data belongs to positive class and only 1% of data belongs to the negative class.
In such a case, standard metrics, such as, accuracy is misleading, because a model can achieve 99% accuracy
by simply predicting positive class all the time.
So, what to do ?
First, of all, start with the confusion matrix. (focus on minority class)
It provides the raw counts of True Positives (TP), True Negatives (TN), False Positives (FP), and False Negatives (FN).
This is the foundation for all other metrics.
Confusion Matrix:
| Predicted Positive | Predicted Negative | |
|---|---|---|
| Actual Positive | True Positive (TP) | False Negative (FN) |
| Actual Negative | False Positive (FP) | True Negative (TN) |
- Precision: Of all instances the model predicted as positive, how many were actually positive?
- \( Precision = \frac{TP}{TP + FP} \)
- Use Case: The cost of a False Positive is high (e.g., marking a legitimate email as spam).
- Recall (Sensitivity): Of all actual positive instances, how many did the model find?
- \( Recall = \frac{TP}{TP + FN} \)
- Use Case: The cost of a False Negative is high (e.g., missing a cancer diagnosis or fraud transaction).
- F1-Score: Harmonic mean of precision and recall.
- \( F1 ~ Score = 2 * \frac{Precision \times Recall}{Precision + Recall}\)
- Why report F1-Score?: To balance precision and recall. A model with 1.0 precision and 0.0 recall will have an F1-score of 0.
- Precision-Recall (PR) AUC: Plots Precision against Recall for different classification thresholds.
Better than ROC curve because it uses Precision instead of False Positive Rate (FPR), which can be misleading for imbalanced data.- \(FPR = \frac{FP}{FP + TN}\)
ROC curve plots TPR vs FPR, \(FPR = \frac{FP}{FP + TN}\), so for an imbalanced data, FPR can be misleading.
So for imbalanced data, we better use Precision-Recall curve that uses Precision instead of FPR and hence is more reliable.
Let’s look at the fraud detection example below, N = 10,000 transactions, Fraud = 100, NOT fraud = 9900:
Confusion Matrix:
| Predicted Fraud | Predicted NOT Fraud | |
|---|---|---|
| Actual Fraud | 80 (TP) | 20 (FN) |
| Actual NOT Fraud | 220 (FP) | 9680 (TN) |
The FPR is very low due to the class imbalance, and hence Precision gives us a better view of the model’s performance.
👉Any information from the validation/test set must NOT influence training, directly or indirectly.
So, how do we prevent this leakage of information or data leakage from training to validation or test set ?
- Train-Test Contamination:
- ❌ Wrong: Applying preprocessing (like global StandardScaler, Mean_Imputation, Target_Encoding etc.) on the entire dataset before splitting.
- ✅ Right: Compute mean, variance, etc. only on the training data and use the same for validation and test data.
- Preventing Leakage in Cross-Validation:
- ❌ Wrong: Perform preprocessing (e.g., scaling, normalization, missing value imputation) on the entire dataset before passing it to cross_val_score.
- ✅ Right: Use sklearn.pipeline.Pipeline; Pipeline ensures that the ‘validation fold’ remains unseen until the transformation is applied using the training fold’s parameters.
- Time Series Data:
- ❌ Wrong: Use standard random CV; it allows the model to ‘peek into the future’.
- ✅ Right: Use Time-Series Nested Cross-Validation (Forward Chaining) instead of random shuffling.
- Target Leakage:
- ❌ Wrong: Include features that are only available after the event we are trying to predict and are proxy for the target.
- e.g. Including number_of_late_payments in a model to predict whether a person applying for a bank loan will default ?
- ✅ Right: Do not include such features during training.
- Group Leakage:
- ❌ Wrong: If you have multiple rows that are correlated (same user).
- For the same patient or user, you put some rows in Train and others in Test.
- ✅ Right: Use GroupKFold to ensure all data from a specific group stays together in one fold.
Bias-Variance Decomposition:
For Mean Squared Error (MSE) = \(\frac{1}{n} \sum_{i=1}^n (y_i - \hat{y_i})^2\)
Total Error = Bias^2 + Variance + Irreducible Error
- Bias = Systematic Error
- Bias measures how far the average prediction of a model is from the true value.
- Variance = Sensitivity to Data
- Variance measures how much the predictions of a model vary for different training datasets.
- Irreducible Error = Sensor noise, Human randomness
- Inherent uncertainty in the data generation process itself and cannot be reduced by any model.
Bias-Variance Trade-Off:
- High Bias (Underfitting): A model with high bias is too simple to capture the underlying patterns in the data
- e.g., fitting a straight line to curved data.
- High Variance (Overfitting): A model with high variance is too complex and learns the noise in the training data rather than the true relationship
- e.g., a high-degree polynomial curve that perfectly fits training data points but performs poorly on new data.
🎯 The goal is to find a balance; a ‘sweet spot’ that minimizes the total error.
🦉 A good model ‘generalizes’ well, i.e., neither too simple (low bias) nor too complex (low variance).
- L1 Regularization:
- \( \underset{w}{\mathrm{min}}\ J_{reg}(w) = \underset{w}{\mathrm{min}}\ J(w) + \lambda_1.\sum_{j=1}^n |w_j| \)
- L2 Regularization:
- \( \underset{w}{\mathrm{min}}\ J_{reg}(w) = \underset{w}{\mathrm{min}}\ J(w) + \lambda_2.\sum_{j=1}^n w_j^2 \)
L1 regularization produces sparse solutions when regularization coefficient \(\lambda_1\) is high.
- Because the gradient of L1 penalty (absolute function) is a constant value, i.e, \(\pm 1\), this means a constant reduction in weight at each step, making it gradually reach to 0 in finite steps.
- Whereas, the derivative of L2 penalty is proportional to the weight (\(2w_j\)) and as the weight reaches close to 0, the gradient also becomes very small, this means that the weight will become very close to 0, but not exactly equal to 0.
💡 Heteroscedasticity = Variance NOT Constant
Note: Linear regression assumes that the data has homoscedasticity (constant variance).
Ordinary Least Squares (OLS) is an unweighted estimator. It treats every data point as equally ‘informative’.
- Under Homoscedasticity: Every point has the same amount of noise, so giving them equal weight is logical.
- Under Heteroscedasticity: Some points have very low variance (high certainty) and some have very high variance (lots of noise).
👉 By treating all the points equally, OLS is ‘wasting’ the precision of the low-variance points and being ‘skewed’ by the high-variance points.
This is why OLS is no longer efficient; does not produce the smallest possible standard errors.
Which means:
- t-tests become unreliable.
- p-values become misleading.
- Confidence intervals are wrong.
👉 OLS is NO longer B.L.U.E. (Best Linear Unbiased Estimator).
- While the coefficients remain unbiased,
they are no longer the ‘best’ because there is another estimator (like Weighted Least Squares) that could provide a lower variance.
👉 How to Test for Heteroscedasticity ?
- Visual (Residual Plot):
- Heteroscedasticity: The points form a ‘fan’ or ‘funnel’ shape, widening or narrowing as values increase.
- Homoscedasticity: The points look like a random ‘cloud’ with consistent thickness.
- Breusch–Pagan Test
- White Test
- Goldfeld–Quandt Test
Probability vs. Likelihood:
Difference lies in which variable is fixed and which is varying.
- Probability(Forward View):
- Quantifies the chance of observing a specific outcome given a known, fixed parameters \(\theta\).
- Likelihood(Backward/Inverse View):
- Inverse concept used for inference (working backward from results to causes).
- It is a function of the parameters \(\theta\) and measures how ‘likely’ a specific set of parameters makes the observed (fixed) data appear.
MLE vs. MAP:
Both help us answer the question:
Which parameter \(\theta\) best explains the data we just saw?
- Maximum Likelihood Estimation (MLE):
- MLE believes the data should speak for itself.
- It asks: ‘Which value of \(\theta\) makes the observed data most probable?’
- It ignores any outside context or common sense.
- Maximum A Posteriori (MAP):
- MAP believes the data is important, but so is prior knowledge.
- It asks: ‘Given the data AND what we already know about the problem at hand, which value of \(\theta\) is most likely?’
The relationship between them is rooted in Bayes’ Theorem:
\[P(\theta \mid \text{data}) = \frac{P(\text{data} \mid \theta) P(\theta)}{P(\text{data})}\]- MLE maximizes only the Likelihood: \(P(\text{data} \mid \theta)\)
- MAP maximizes the Posterior: \(P(\text{data} \mid \theta) P(\theta)\)
Note:
- P(data) is a constant (a ’normalizing factor’), so we ignore it during maximization.
- For a prior with uniform distribution where every value is equally likely, MAP becomes MLE.
👉 Coin Toss Example:
- Data: Toss the coin 3 times, 2H + 1T.
- MLE: Probability of heads (\(\theta\))
- \(\theta_{MLE}\) = 2/3 = 0.67
- MAP (with prior belief that coin is fair)
- Assume prior: \(\theta \sim \beta(10,10)\)
- Posterior = \(\beta(12,11)\)
- \(\theta_{MAP}\) = 0.52 (Prior pulls estimate towards 0.5)
- MLE: Probability of heads (\(\theta\))
✅ Use MLE when:
- large dataset.
- no reliable prior knowledge.
✅ Use MAP when:
- small dataset.
- reliable prior/domain knowledge.
Entropy (H) is a measure of impurity or randomness in a dataset.
For binary classification, where the outcome is Yes/No, 0/1 etc., entropy will be:
\[H(S) = -p \log_2(p) - (1-p) \log_2(1-p)\]- Max Entropy: H(S) = 1, when the classes are split 50/50 (maximum uncertainty).
- Min Entropy: H(S) = 0 when the set is pure (all examples belong to one class).
Information Gain:
️Measures the reduction in entropy (uncertainty) achieved by splitting a dataset based on a specific attribute.
Note: The goal of a decision tree algorithm is to find the split that maximizes information gain, meaning it removes the most uncertainty from the data.
👉 To understand how a Decision Tree selects the ‘best’ root node, let’s use the example below:
The Dataset: “Will they buy the product?”
| ID | Age | Income | Credit Score | Buy? (Target) |
|---|---|---|---|---|
| 1 | Youth | High | Good | No |
| 2 | Youth | High | Excellent | No |
| 3 | Middle | High | Good | Yes |
| 4 | Senior | Medium | Good | Yes |
| 5 | Senior | Low | Good | Yes |
| 6 | Senior | Low | Excellent | No |
- Calculate parent node’s entropy:
- P(yes) = P(no) = 3/6 = 0.5
- \(H(Parent) = -(0.5 \log_2 0.5 + 0.5 \log_2 0.5) = \mathbf{1.0}\)
- Evaluate feature ‘Age’:
- Youth: 2 samples (0 Yes, 2 No). \(H(Youth) = \mathbf{0}\)
- Middle: 1 sample (1 Yes, 0 No). \(H(Middle) = \mathbf{0}\)
- Senior: 3 samples (2 Yes, 1 No). \(H(Senior) = -(\frac{2}{3} \log_2 \frac{2}{3} + \frac{1}{3} \log_2 \frac{1}{3}) \approx \mathbf{0.918}\)
- Weighted Entropy for Age:
- \((\frac{2}{6} \times 0) + (\frac{1}{6} \times 0) + (\frac{3}{6} \times 0.918) = \mathbf{0.459}\)
- Information Gain (Age): \(1.0 - 0.459 = \mathbf{0.541}\)
- Evaluate feature ‘Income’:
- High: 3 samples (1 Yes, 2 No). \(H(High) \approx \mathbf{0.918}\)
- Medium: 1 sample (1 Yes, 0 No). \(H(Medium) = \mathbf{0}\)
- Low: 2 samples (1 Yes, 1 No). \(H(Low) = \mathbf{1.0}\)
- Weighted Entropy for Income:
- \((\frac{3}{6} \times 0.918) + (\frac{1}{6} \times 0) + (\frac{2}{6} \times 1.0) = \mathbf{0.792}\)
- Information Gain (Income): \(1.0 - 0.792 = \mathbf{0.208}\)
- Evaluate feature ‘Credit Score’
- Good: 4 samples (3 Yes, 1 No). \(H(Good) \approx \mathbf{0.811}\)
- Excellent: 2 samples (0 Yes, 2 No). \(H(Excellent) = \mathbf{0}\)
- Weighted Entropy for Credit Score:
- \((\frac{4}{6} \times 0.811) + (\frac{2}{6} \times 0) = \mathbf{0.541}\)
- Information Gain (Credit Score): \(1.0 - 0.541 = \mathbf{0.459}\)
- The Decision Tree algorithm compares the information gain for all the features, and splits on the feature with maximum information gain.
- Here in our case it is “Age”, IG = 0.541.
- The algorithm chooses ‘Age’ as the root node.
- Splits the data into three branches (Youth, Middle, Senior).
End of Section
2 - Maths
Maths for AI & ML
This sheet contains all the topics that will be covered for Maths for AI & ML.
2.1.1 - Introduction to Probability
Because the world around us is very uncertain, and Probability acts as -
the fundamental language
to understand, express and deal with this uncertainty.
- Toss a fair coin, \(P(H) = P(T) = 1/2\)
- Roll a die, \(P(1) = P(2) = P(3) = P(4) = P(5) = P(6) = 1/6\)
- Email classifier, \(P(spam) = 0.95 ,~ P(not ~ spam) = 0.05\)
Range: \([0,1]\)
\(P=0\): Highly unlikely
\(P=1\): Almost certain
Symbol: \(\Omega\)
\(P(\Omega) = 1\)
- Toss a fair coin, sample space: \(\Omega = \{H,T\}\)
- Roll a die, sample space: \(\Omega = \{1,2,3,4,5,6\}\)
- Choose a real number \(x\) from the interval \([2,3]\), sample space: \(\Omega = [2,3]\); sample size = \(\infin\)
Note: There can be infinitely many points between 2 and 3, e.g: 2.21, 2.211, 2.2111, 2.21111, … - Randomly put a point in a rectangular region; sample size = \(\infin\)
Note: There can be infinitely many points in any rectangular region.
A,B,…⊆Ω
- Toss a fair coin, set of possible outcomes: \(\{H,T\}\)
- Roll a die, set of possible outcomes: \(\{1,2,3,4,5,6\}\)
- Roll a die, event \(A = \{1,2\} => P(A) = 2/6 = 1/3\)
- Email classifier, set of possible outcomes: \(\{spam,not ~spam\}\).
- Toss a fair coin, possible outcomes: \(\Omega = \{H,T\}\)
- Roll a die, possible outcomes: \(\Omega = \{1,2,3,4,5,6\}\)
- Choose a real number \(x\) from the interval \([2,3]\) with decimal precision, sample space: \(\Omega = [2,3]\).
Note: There are 99 real numbers between 2 and 3 with 2 decimal precision i.e from 2.01 to 2.99. - Number of cars passing a specific traffic signal in 1 hour.
- A line segment between 2 and 3 - forms a continuum.
- Randomly put a point in a rectangular region.
graph TD
A[Sample Space] --> |Discrete| B(Finite)
A --> C(Infinite)
C --> |Discrete| D(Countable)
C --> |Continuous| E(Uncountable)No overlapping or common outcomes.
If one event occurs, then the other event does NOT occur.
- Roll a die, sample space: \(\Omega = \{1,2,3,4,5,6\}\)
Odd outcome = \(A = \{1,3,5\}\)
Even outcome = \(B = \{2,4,6\}\) are mutually exclusive.
\(P(A \cap B) = 0\)
Since, \(P(A \cup B) = P(A) + P(B) - (P(A \cap B)\)
Therefore, \(P(A \cup B) = P(A) + P(B)\)
Note: If we know that event \(A\) has occurred, then we can say for sure that the event \(B\) did NOT occur.
- Roll a die twice , sample space: \(\Omega = \{1,2,3,4,5,6\}\)
Odd number in 1st throw = \(A = \{1,3,5\}\)
Odd number in 2nd throw = \(B = \{1,3,5\}\)
Note: A and B are independent because whether we get an odd number in 1st roll has NO impact of getting an odd number in second roll.
\(P(A \cap B) = P(A)*P(B)\)
Note: If we know that event \(A\) has occurred, then that gives us NO new information about the event \(B\).
Let’s understand this answer with an example.
Probability of choosing exactly one point on the number line or a real number, say 2.5,
from the interval \([2,3]\) is almost = 0, because there are infinitely many points between 2 and 3.
Also, we can NOT say that choosing exactly 2.5 is impossible, because it exists there on the number line.
But, for all practical purposes, \(P(2.5) = 0\).
Therefore, we say that \(P=0\) means “Highly Unlikely” and NOT “Impossible”.
Extending this line of reasoning, we can say that probability of NOT choosing 2.5, \(P(!2.5) = 1\).
Theoretically yes, because there are infinitely many points between 2 and 3.
But, we cannot say for sure that we cannot choose 2.5 exactly.
There is some probability of choosing 2.5, but it is very small.
Therefore, we say that \(P=1\) means “Almost Sure” and NOT “Certain”.
Here, in this case we can say that \(P(7)=0\) and that means Impossible.
Similarly, we can say that \(P(get ~any ~number ~between ~1 ~and ~6)=1\) and \(P=1 => \) Certain.
End of Introduction
2.1.2 - Conditional Probability
It is the probability of an event occurring, given that another event has already occurred.
Allows us to update probability when additional information is revealed.
\(P(A \cap B) = P(A)*P(B \mid A)\)
- Roll a die, sample space: \(\Omega = \{1,2,3,4,5,6\}\)
Event A = Get a 5 = \(\{5\} => P(A) = 1/6\)
Event B = Get an odd number = \(\{1, 3, 5\} => P(B) = 3/6 = 1/2\)
It is a formula that uses conditional probability.
It allows us to update our belief about an event’s probability based on new evidence.
We know from conditional probability and chain rule that:
Combining all the above equations gives us the Bayes’ Theorem:
$$ \begin{aligned} P(A \mid B) = \frac{P(A)*P(B \mid A)}{P(B)} \end{aligned} $$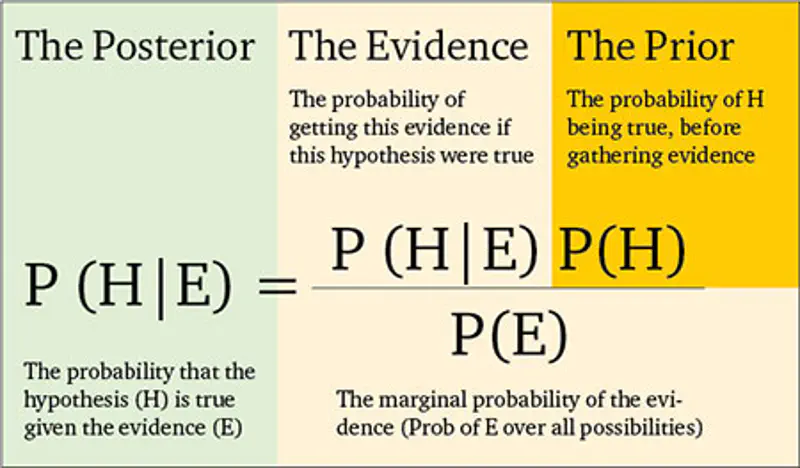
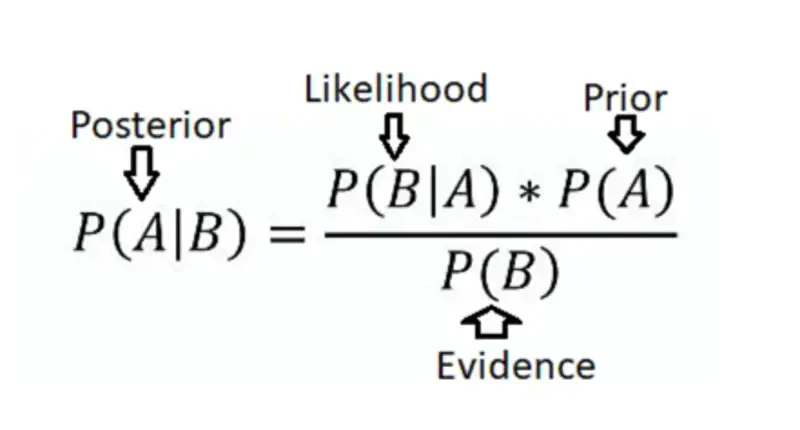
- Roll a die, sample space: \(\Omega = \{1,2,3,4,5,6\}\)
Event A = Get a 5 = \(\{5\} => P(A) = 1/6\)
Event B = Get an odd number = \(\{1, 3, 5\} => P(B) = 3/6 = 1/2\)
Task: Find the probability of getting a 5 given that you rolled an odd number.
\(P(B \mid A) = 1\) = Probability of getting an odd number given that we have rolled a 5.
Now, let’s understand another concept called Law of Total Probability.
Here, we can say that the sample space \(\Omega\) is divided into 2 parts - \(A\) and \(A ^ \complement \)
So, the probability of an event \(B\) is given by:
\[ B = B \cap A + B \cap A ^ \complement \\ P(B) = P(B \cap A) + P(B \cap A ^ \complement ) \\ By ~Chain ~Rule: P(B) = P(A)*P(B \mid A) + P(A ^ \complement )*P(B \mid A ^ \complement ) \]Overall probability of an event B, considering all the different, mutually exclusive ways it can occur.
If A₁, A₂, …, Aₙ are a set of events that partition the sample space, such that they are -
- Mututally exclusive : \(A_i \cap A_j = \emptyset\) for all \(i, j\)
- Exhaustive: \(A₁ \cup A₂ ... \cup Aₙ = \Omega\) for all \(i \neq j\) \[ P(B) = \sum_{i=1}^{n} P(A_i)*P(B \mid A_i) \] where \(n\) is the number of mutually exclusive partitions of the sample space \(\Omega\) .
Now, we can also generalize the Bayes’ Theorem using the Law of Total Probability.
End of Section
2.1.3 - Independence of Events
Two events are independent if the occurrence of one event does not affect the probability of the other event.
There are 3 types of independence of events:
- Mutual Independence
- Pair-Wise Independence
- Conditional Independence
Joint probability of two events is equal to the product of the individual probabilities of the two events.
\(P(A \cap B) = P(A)*P(B)\)
Joint probability: The probability of two or more events occurring simultaneously.
\(P(A \cap B)\) or \(P(A, B)\)
- Toss a coin and roll a die -
\(A\) = Get a heads; \(P(A)=1/2\)
\(B\) = Get an odd number; \(P(B)=1/2\)
=> A and B are mutually independent.
Pair-wise independence != Mutual independence.
- Toss 3 coins;
For 2 tosses, sample space: \(\Omega = \{HH,HT, TH, TT\}\)
\(A\) = First and Second toss outcomes are same i.e \(\{HH, TT\}\); \(P(A)= 2/4 = 1/2\)
\(B\) = Second and Third toss outcomes are same i.e \(\{HH, TT\}\); \(P(B)= 2/4 = 1/2\)
\(C\) = Third and First toss outcomes are same i.e \(\{HH, TT\}\); \(P(C)= 2/4 = 1/2\)
Now, pair-wise independence of the above events A & B is - \(P(A \cap B)\)
\(P(A \cap B)\) => Outcomes of first and second toss are same &
outcomes of second and third toss are same.
=> Outcomes of all the three tosses are same.
Total number of outcomes = 8
Desired outcomes = \(\{HHH, TTT\}\) = 2
=> \(P(A \cap B) = 2/8 = 1/4 = P(A) * P(B) = 1/2 * 1/2 = 1/4\)
Therefore, \(A\) and \(B\) are pair-wise independent.
Similarly, we can also prove that \(A\) and \(C\) and \(B\) and \(C\) are also pair-wise independent.
Now, let’s check for mutual independence of the above events A, B & C.
\(P(A \cap B \cap C) = P(A)*P(B)*P(C)\)
\(P(A \cap B \cap C)\) = Outcomes of all the three tosses are same i.e \(\{HHH, TTT\}\)
Total number of outcomes = 8
Desired outcomes = \(\{HHH, TTT\}\) = 2
So, \(P(A \cap B \cap C)\) = 2/8 = 1/4
But, \(P(A)*P(B)*P(C) = 1/2*1/2*1/2 = 1/8\)
Therefore \(P(A \cap B \cap C)\) ≠ \(P(A)*P(B)*P(C)\)
=> \(A, B, C\) are NOT mutually independent but only pair wise independent.
if they are independent given that C has occurred.
Occurrence of C changes the context, causing the events A & B to become independent of each other.
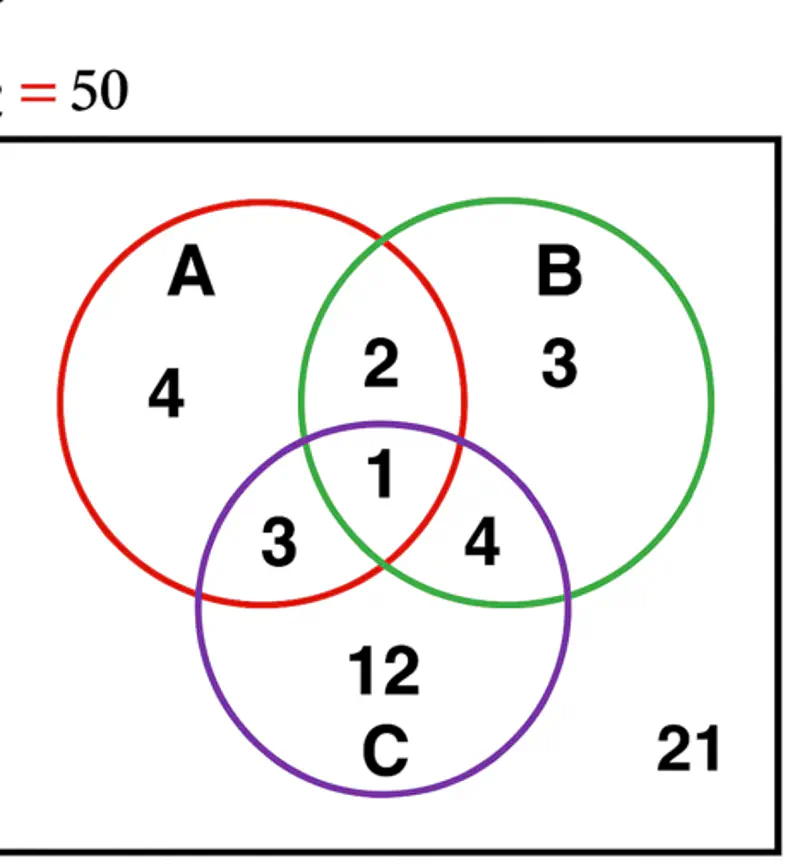
=> A & B are NOT independent.
Now, let’s check for conditional independence of A & B given C.
Therefore, A & B are conditionally independent given C.
End of Section
2.1.4 - Cumulative Distribution Function
Random Variable X is represented as, \(X: \Omega \to \mathbb{R} \)
👉 It maps abstract outcomes of a random experiment to concrete numerical values required for mathematical analysis.
- Toss a coin 2 times, sample space: \(\Omega = \{HH,HT, TH, TT\}\)
The above random experiment of coin tosses can be mapped to a random variable \(X: \Omega \to \mathbb{R} \)
\(X: \Omega = \{HH,HT, TH, TT\} \to \mathbb{R}) \)
Say, if we count the number of heads, then
\[ \begin{aligned} X(0) = \{TT\} = 1 \\ X(1) = \{HT, TH\} = 2 \\ X(2) = \{HH\} = 1 \\ \end{aligned} \] Similar, output will be observed for number of tails.
Depending upon the nature of output, random variables are of 2 types - Discrete and Continuous.
Typically obtained by counting.
Discrete random variable cannot take any value between 2 consecutive values.
Possible outcomes are infinite.
- A person’s height in a given range of say 150cm-200cm.
Height can take any value, not just round values, e.g: 150.1cm, 167.95cm, 180.123cm etc.
Now, that we have understood that how random variable can be used to map outcomes of abstract random experiment to real values for mathematical analysis, we will understand its applications.
CDF = \(F(X) = P(X \leq x)\) i.e Probability a random variable \(X\) will take for a value \(<=x\).
- Discrete random variable - Toss a coin 2 times, sample space: \(\Omega = \{HH,HT, TH, TT\}\)
Count the number of heads.
\[ \begin{aligned} X(0) = \{TT\} = 1 => P(X = 0) = 1/4 \\ X(1) = \{HT, TH\} = 2 => P(X = 1) = 1/2\\ X(2) = \{HH\} = 1 => P(X = 2) = 1/4 \\ \\ CDF = F(X) = P(X \leq x) \\ F(0) = P(X \leq 0) = P(X < 0) + P(X = 0) = 1/4 \\ F(1) = P(X \leq 1) = P(X < 1) + P(X = 1) = 1/4 + 1/2 = 3/4 \\ F(2) = P(X \leq 2) = P(X < 2) + P(X = 2) = 3/4 + 1/4 = 1 \\ \end{aligned} \]
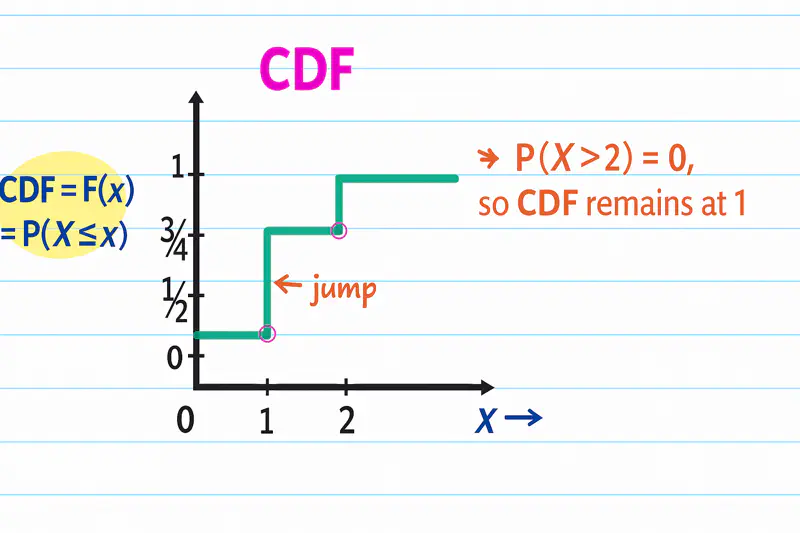
- Continuous random variable - Consider a line segment/interval from \(\Omega = [0,2] \)
Random variable \(X(\omega) = \omega\)
i.e \(X(1) = 1 ~and~ X(1.1) = 1.1 \)
\[ \begin{aligned} P[(0,1/2)] = (1/2)/2 = 1/4 \\ P[(3/4, 2)] = (2-3/4)/2 = 5/8 \\ P(X<=1.1) = P(-\infty, 1.1)/2 = 1.1/2 \end{aligned} \] \[ F_X(x) = P(X \leq x) = \begin{cases} \frac{x}{2} & \text{if } x \in [0,2] \\ 1 & \text{if } x > 2 \\ 0 & \text{if } x < 0 \end{cases} \]
- Non-Decreasing:
For any 2 values \(x_1\) and \(x_2\) such that \(x_1 \leq x_2\), corresponding CDF must satisfy -
\(F(x_1) \leq F(x_2)\)
Note: Cumulative function can never decrease as x increases. - Bounded:
Range of CDF is always between 0 and 1, because CDF represents total probability, which cannot be negative or greater than 1.
\(\lim_{x \to -\infty} F(X) = 0\); as \(x \to -\infty\), event \((X \le x)\) becomes an impossible event
i.e \(P(X \le x) =0\)
\(\lim_{x \to \infty} F(X) = 1\); as \(x \to \infty\), event \((X \le x)\) includes all possible outcomes of the event, making sure that \(P(X \le x) =1\) - Right Continuous:
Function’s value at a point is same as the limit of the function, as we approach that point from right hand side (RHS).
Note: Reason for only right continuity is because how the cumulative distribution function is defined, as we may have a jump on the left side of the point.
\(\lim_{h \to o^+} F(X+h) = F(X)\)
For example:
In the above coin toss CDF example, if we approach X=1 from right, say 1.001, 1.01 etc, the value of \(F(X) = P(X \leq x) = F(1) = 3/4\).
But, if we approach \(X=1\) from left, say 0.99, 0.999 etc, the value of \(F(X) = 1/4\), as these values do NOT yet include the value of the probability at \(X=1\).
Value of the probability of a random variable X, at any given value x, is calculated by summing up all the probabilities for values \(\le x\).
\( CDF = F_X(x) = \sum_{x_i \le x} P(X=x_i) \), where \(P(X=x_i)\) is the Probability Mass Function (PMF) at \(x_i\)
In the above coin toss example -
Height of the jump at ‘x’ = Probability at that value ‘x’.
e.g: Jump at (x=1) = 1/2 = Probability at (x=1).
CDF for continuous random variable is calculated by integrating the probability density function (PDF) from \(-\infty\) to the given value \(x\).
\( CDF = F_X(x) = \int_{-\infty}^{x} f(x) \,dx \), where \(f(x)\) is the Probability Density Function (PDF) of random variable.
Note: We can also say that PDF is the derivative of CDF for continuous random variable.
\(PDF = f(x) = F'(X) = \frac{dF_X(x)}{dx} \)
Read more about Integration
End of Section
2.1.5 - Probability Mass Function
It assigns a “non-zero” mass or probability to a specific countable outcome.
Note: Called ‘mass’ because probability is concentrated at a single discrete point.
\(PMF = P(X=x)\)
e.g: Bernoulli, Binomial, Multinomial, Poisson etc.
Commonly visualised as a bar chart.
Note: PMF = Jump at a given point in CDF.
\(PMF = P_x(X=x_i) = F_X(X=x_i) - F_X(X=x_{i-1})\)

- Non-Negative: Probability of any value ‘x’ must be non-negative i.e \(P(X=x) \ge 0 ~\forall x~\).
- Sum = 1: Sum of probabilities of all possible outcomes must be 1.
\( \sum_{x} P(X=x) = 1 \) - For any value that the discrete random variable can NOT take, the probability must be zero.
p = Probability of success
1-p = Probability of failure
Mean = p
Variance = p(1-p)
Note: A single trial that adheres to these conditions is called a Bernoulli trial.
\(PMF, P(x) = p^x(1-p)^{1-x}\), where \(x \in \{0,1\}\)
- Toss a coin, we get heads or tails.
- Result of a test, pass or fail.
- Machine learning, binary classification model.
It extends the Bernoulli distribution by modeling the number of successes that occur over a fixed number of
independent trials.
n = Number of trials
k = Number of successes
p = Probability of success
Mean = np
Variance = np(1-p)
\(PMF, P(x=k) = \binom{n}{k}p^k(1-p)^{n-k}\), where \(k \in \{0,1,2,3,...,n\}\)
\(\binom{n}{k} = \frac{n!}{k!(n-k)!}\) i.e number of ways to achieve ‘k’ successes in ‘n’ independent trials.
Note: Bernoulli is a special case of Binomial distribution where n = 1.
Also read about Multinomial distribution i.e where number of possible outcomes is > 2.
- Counting number of heads(success) in ’n’ coin tosses.
- Trials are independent.
- Probability of success remains constant for every trial.
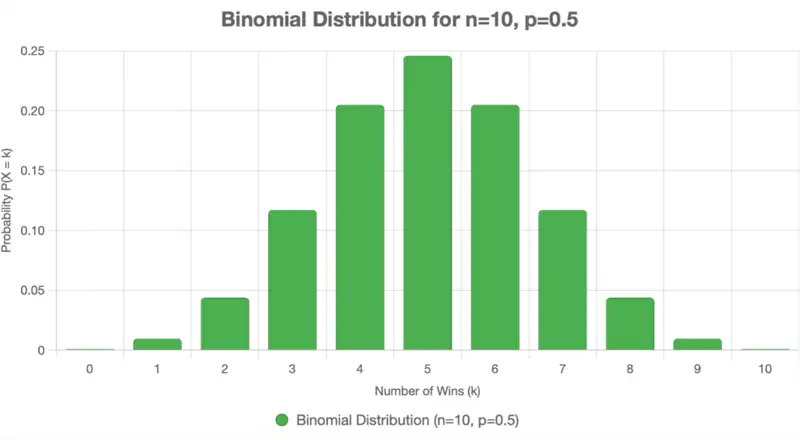
Total number of outcomes in 3 coin tosses = 2^3 = 8
Desired outcomes i.e 2 heads in 3 coin tosses = \(\{HHT, HTH, THH\}\) = 3
Probability of getting exactly 2 heads in 3 coin tosses = \(\frac{3}{8}\) = 0.375
Now lets solve the question using the binomial distribution formula.
Number of successes, k = 1
Number of trials, n = 10
Probability of success, p = 1/3
Probability of winning lottery, P(k=1) =
It expresses the probability of an event happening a certain number of times ‘k’ within a fixed interval of time.
Given that:
- Events occur with a known constant average rate.
- Occurrence of an event is independent of the time since the last event.
Parameters:
\(\lambda\): Expected number of events per interval
\(k\) = Number of events in the same interval
Mean = \(\lambda\)
Variance = \(\lambda\)
PMF = Probability of occurrence of ‘k’ events in the same interval
Note: Useful to count data where total population size is large but the probability of an individual event is small.
- Model the number of customers arrival at a service center per hour.
- Number of website clicks in a given time period.
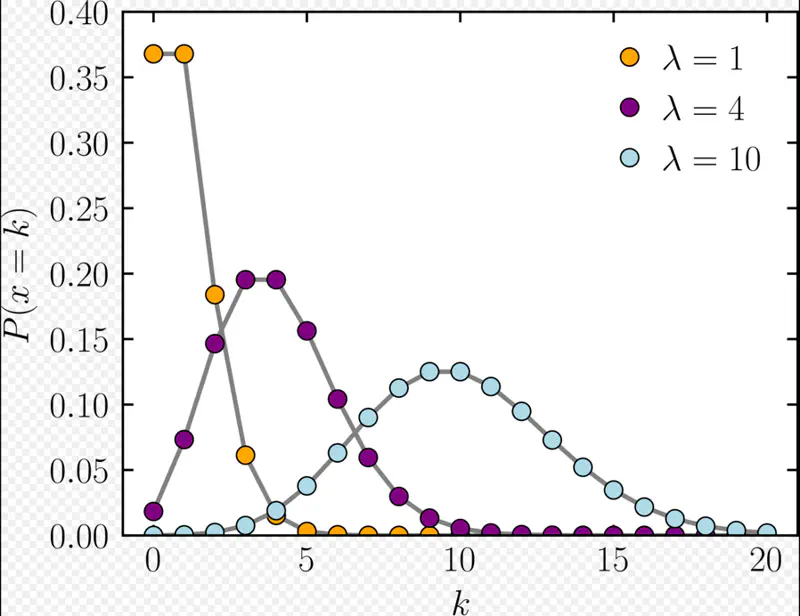
Expected (average) number of emails per hour, \(\lambda\) = 5
Probability of receiving exactly k=3 emails in the next hour =
End of Section
2.1.6 - Probability Density Function
This is a function used for continuous random variables to describe the likelihood of the variable taking on a value
within a specific range or interval.
Since, at any given point the probability of a continuous random variable is zero,
we find the probability within a given range.
Note: Called ‘density’ because probability is spread continuously over a range of values
rather than being concentrated at a single point as in PMF.
e.g: Uniform, Gaussian, Exponential, etc.
Note: PDF is a continuous function \(f(x)\).
It is also the derivative of Cumulative Distribution Function (CDF) \(F_X(x)\)
\(PDF = f(x) = F'(X) = \frac{dF_X(x)}{dx} \)
- Non-Negative: Function must be non-negative everywhere i.e \(f(x) \ge 0 \forall x\).
- Sum = 1: Total area under curve must be equal to 1.
\( \int_{-\infty}^{\infty} f(x) \,dx = 1\) - Probability of a continuous random variable in the range [a,b] is given by -
\( P(a \le x \le b) = \int_{a}^{b} f(x) \,dx\)
Consider a line segment/interval from \(\Omega = [0,2] \)
Random variable \(X(\omega) = \omega\)
i.e \(X(1) = 1 ~and~ X(1.1) = 1.1 \)
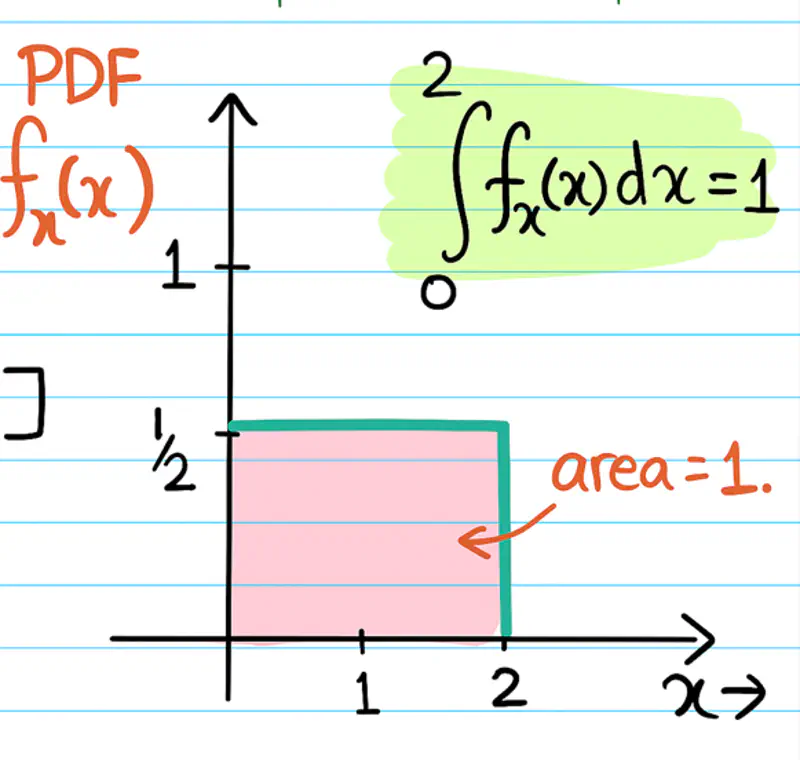
Note: If we know the PDF of a continuous random variable, then we can find the probability of any given region/interval by calculating the area under the curve.
All the outcomes within the given range are equally likely to occur.
Also known as ‘fair’ distribution.
Note: This is a natural starting point to understand randomness in general.
Mean = Median = \( \frac{a+b}{2} ~if~ x \in [a,b] \)
Variance = \( \frac{(b-a)^2}{12} \)
Standard uniform distribution: \( X \sim U(0,1) \)
PDF in terms of mean(\(\mu\)) and standard deviation(\(\sigma\)) -
Random number generator that generates a random number between 0 and 1.
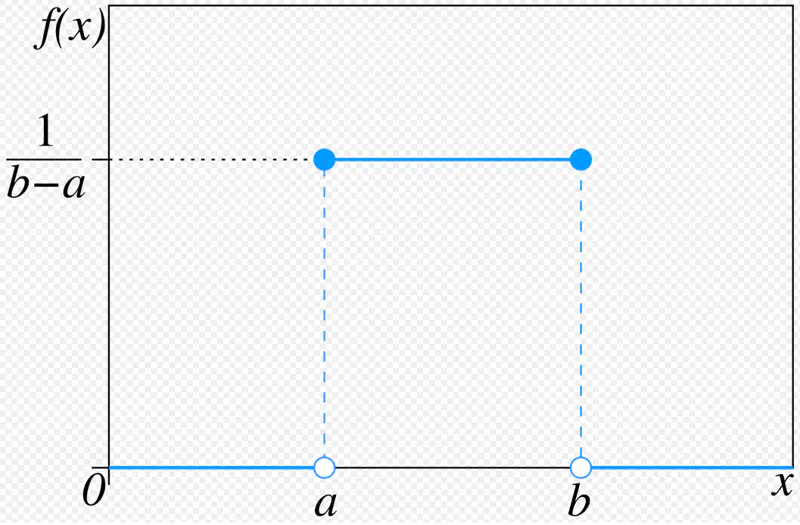
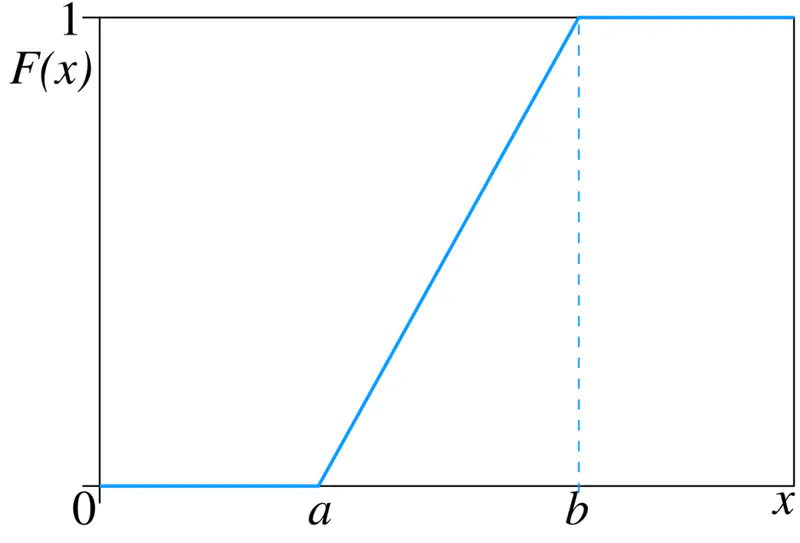
It is a continuous probability distribution characterized by a symmetrical, bell-shaped curve with
most data clustered around the central average, with frequency of values decreasing as they move away from the center.
- Most outcomes are average; extremely low or extremely high values are rare.
- Characterised by mean and standard deviation/variance.
- Peak at mean = median, symmetric around mean.
Note: Most important and widely used distribution.
\[ X \sim N(\mu, \sigma^2) \] $$ \begin{aligned} PDF = f(x) = \dfrac{1}{\sqrt{2\pi}\sigma}e^{-\dfrac{(x-\mu)^2}{2\sigma^2}} \\ \end{aligned} $$
Mean = \(\mu\)
Variance = \(\sigma^2\)
Standard normal distribution:
\[ Z \sim N(0,1) ~i.e~ \mu = 0, \sigma^2 = 1 \]
Any normal distribution can be standardized using Z-score transformation:
- Human height, IQ scores, blood-pressure etc.
- Measurement of errors in scientific experiments.
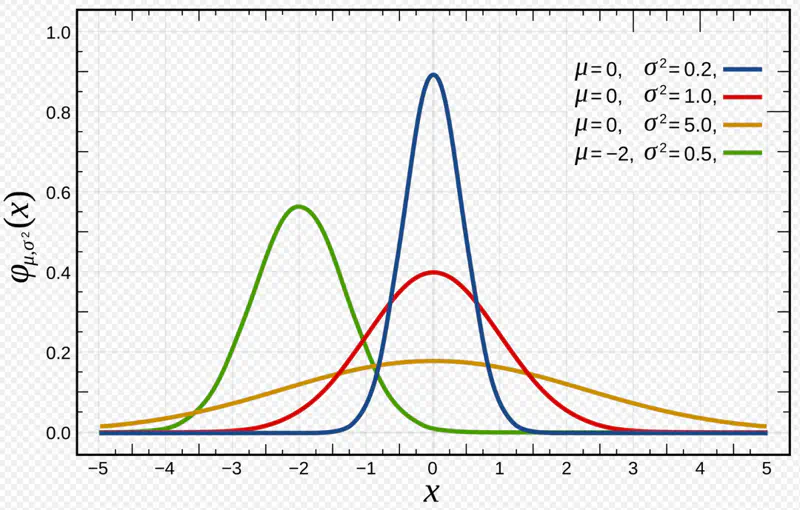
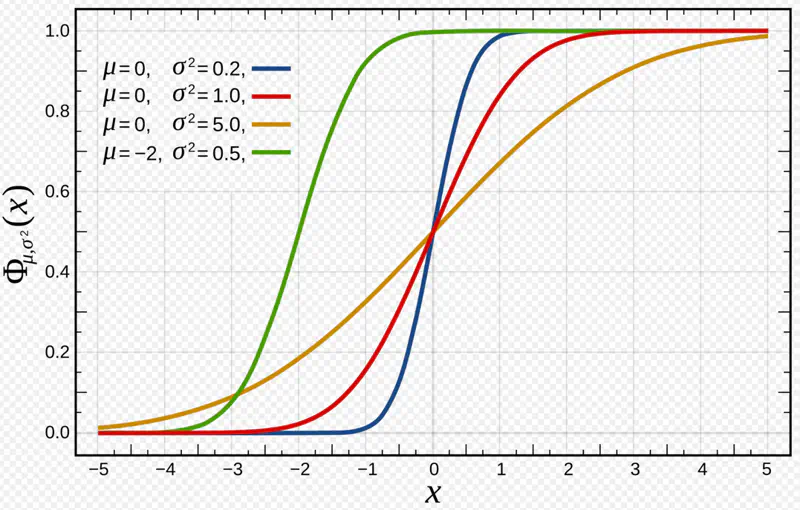
- 68.27% of the data lie within 1 standard deviation of the mean i.e \(\mu \pm \sigma\)
- 95.45% of the data lie within 2 standard deviations of the mean i.e \(\mu \pm 2\sigma\)
- 99.73% of the data lie within 3 standard deviations of the mean i.e \(\mu \pm 3\sigma\)
It is used to model the amount of time until a specific event occurs.
Given that:
- Events occur with a known constant average rate.
- Occurrence of an event is independent of the time since the last event.
Parameters:
Rate parameter: \(\lambda\): Average number of events per unit time
Scale parameter: \(\mu ~or~ \beta \): Mean time between events
Mean = \(\frac{1}{\lambda}\)
Variance = \(\frac{1}{\lambda^2}\)
\[ \lambda = \dfrac{1}{\beta} = \dfrac{1}{\mu}\]
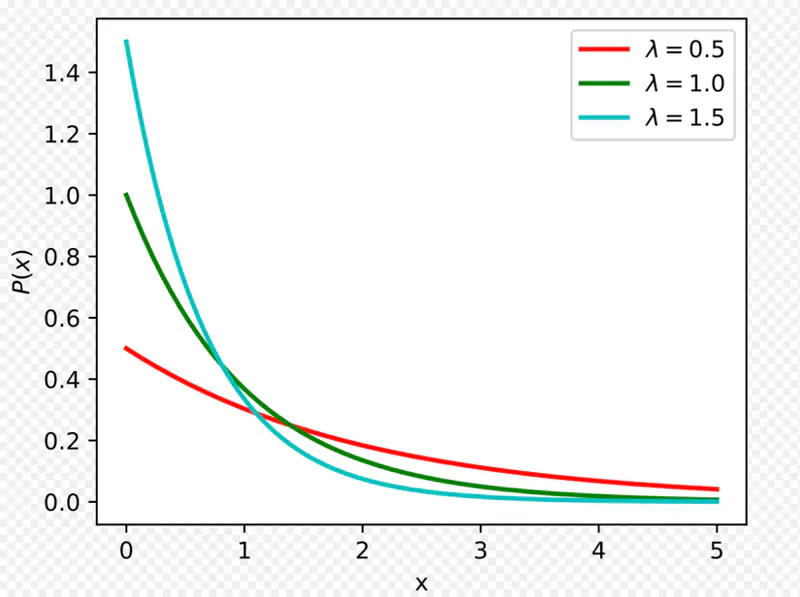
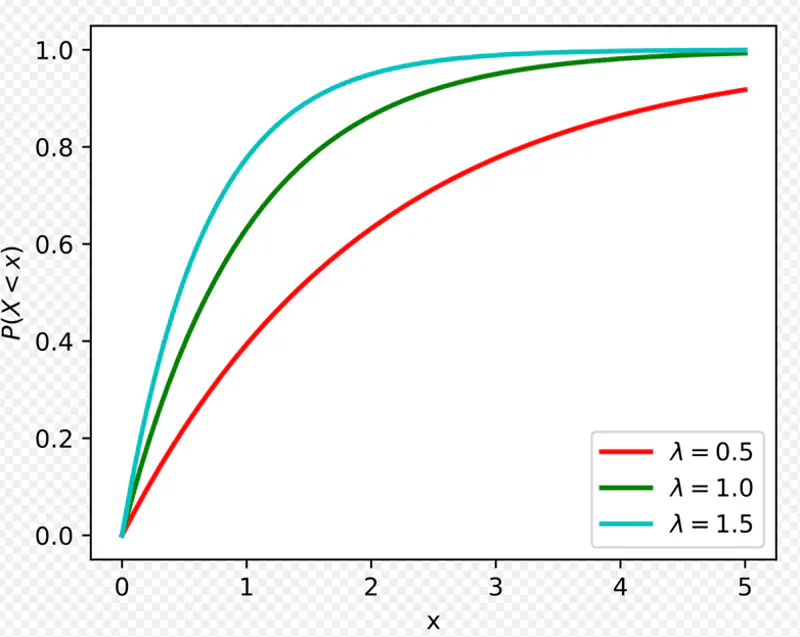
Mean time to serve 1 customer = \(\mu\) = 4 minutes
So, \(\lambda\) = average number of customers served per unit time = \(1/\mu\) = 1/4 = 0.25 minutes
Probability to serve a customer in less than 3 minutes can be found using CDF -
So, probability of a customer being served in less than 3 minutes is 53%(approx).
In this case x = 2 minutes, and \(\lambda\) = 0.25 so,
So, probability of a customer being served in greater than 2 minutes is 60%(approx).
Probability of waiting for an additional period of time for an event to occur is independent of
how long you have already waited.
e.g: Lifetime of electronic items follow exponential distribution.
- Probability of a computer part failing in the next 1 hour is the same regardless of whether
it has been working for 1 day or 1 year or 5 years.
Note: Memoryless property makes exponential distribution particularly useful for -
- Modeling systems that do not experience ‘wear and tear’; where failure is due to a constant random rate rather than degradation over time.
- Also, useful for ‘reliability analysis’ of electronic systems where a ‘random failure’ model is more appropriate than
a ‘wear out’ model.
Task: We want to find the probability of the electronic item lasting for \(x > t+s \) days,
given that it has already lasted for \(x>t\) days.
This is a ‘conditional probability’.
Since,
We want to find: \( P(X > t+s \mid X > t) = ~? \)
Hence, probability that the electronic item will survive for an additional time ’s’ days
is independent of the time ’t’ days it has already survived.
Poisson distribution models the number of events occurring in a fixed interval of time,
given a constant average rate \(\lambda \).
Exponential distribution models the time interval between those successive events.
- 2 faces of the same coin.
- \(\lambda_{poisson}\) is identical to rate parameter \(\lambda_{exponential}\).
Note: If the number of events in a given interval follow a Poisson distribution, then the waiting time between those events will necessarily follow an Exponential distribution.
Lets see the proof for the above statement.
Poisson Case:
The probability of observing exactly ‘k’ events in a time interval of length ’t’
with an average rate of \( \lambda \) events per unit time is given by -
The event that waiting time until next event > t, is same as observing ‘0’ events in that interval.
=> We can use the PMF of Poisson distribution with k=0.
Exponential Case:
Now, lets consider exponential distribution that models the waiting time ‘T’ until next event.
The event that waiting time ‘T’ > t, is same as observing ‘0’ events in that interval.
Observation:
Exponential distribution is a direct consequence of Poisson distribution probability of ‘0’ events.
Using Poisson:
Average failure rate, \(\lambda\) = 1/20 = 0.05
Time interval, t = 10 hours
Number of events, k = 0
Average number of events in interval, (\(\lambda t\)) = (1/20) * 10 = 0.5
Probability of having NO failures in the next 10 hours = ?
Using Exponential:
What is the probability that wait time until next failure > 10 hours?
Waiting time, T > t = 10 hours.
Average number of failures per hour, \(\lambda\) = 1/20 per hour
Therefore, we have seen that this problem can be solved using both Poisson and Exponential distribution.
End of Section
2.1.7 - Expectation
Long run average of the outcomes of a random experiment.
When we talk about ‘Expectation’ we mean - on an average.
- Value we expect to get when we repeated an experiment multiple times and took an average of the results.
- Measures the central tendency of a data distribution.
- Similar to ‘Center of Mass’ in Physics.
Note: If the probability of each outcome is different, then we take a weighted average.
Discrete Case:
\(E[X] = \sum_{i=1}^{n} x_i.P(X = x_i) \)
Continuous Case:
\(E[X] = \int_{-\infty}^{\infty} x.f(x) dx \)
if its a tail, then you lose Rs. 50. What is the expected value of the amount that you will win per toss?
Possible outcomes are: \(x_1 = 100 ,~ x_2 = -50 \)
Probability of each outcome is \(P(X = 100) = 0.5,~ P(X = -50) = 0.5 \)
Therefore, the expected value of the amount that you will win per toss is Rs. 25 in the long run.
It is the average of the squared differences from the mean.
- Measures the spread or variability in the data distribution.
- If the variance is low, then the data is clustered around the mean i.e less variability;
and if the variance is high, then the data is widely spread out i.e high variability.
Variance in terms of expected value, where \(E[X] = \mu\), is given by:
\[ \begin{aligned} Var[X] &= E[(X - E[X])^2] \\ &= E[X^2 + E[X]^2 - 2XE[X]] \\ &= E[X^2] + E[X]^2 - 2E[X]E[X] \\ &= E[X^2] + E[X]^2 - 2E[X]^2 \\ Var[X] &= E[X^2] - E[X]^2 \\ \end{aligned} \]Note: This is the computational formula for variance, as it is easier to calculate than the
average of square distances from mean.
PDF of continuous uniform random variable =
$$ \begin{aligned} f_X(x) &= \begin{cases} \dfrac{1}{b-a}, & x \in [a,b] \\ 0, & \text{otherwise.} \end{cases} \\ \end{aligned} $$\[ \begin{aligned} \text{Expected value = mean } = \\[0.5em] E[X] = \dfrac{b+a}{2} \\[0.5em] Var[X] = E[X^2] - E[X]^2 \end{aligned} \]We know \(E[X]\) already, now we will calculate \(E[X^2]\):
\[ \begin{aligned} E[X] &= \int_{-\infty}^{\infty} x.f(x) dx \\ E[X^2] &= \int_{-\infty}^{\infty} x^2.f(x) dx \\ &= \int_{-\infty}^{a} x^2f(x) dx + \int_{a}^{b} x^2f(x) dx + \int_{b}^{\infty} x^2f(x) dx\\ &= 0 + \int_{a}^{b} x^2f(x) dx + 0 \\ &= \int_{a}^{b} x^2f(x) dx \\ &= \dfrac{1}{b-a} \int_{a}^{b} x^2 dx \\ &= \dfrac{1}{b-a} * \{\frac{x^3}{3}\}_{a}^{b} \\ &= \dfrac{1}{b-a} * \{\frac{b^3 - a^3}{3}\} \\ &= \dfrac{1}{b-a} * \{\frac{(b-a)(b^2+ab+a^2)}{3}\} \\ E[X^2] &= \dfrac{b^2+ab+a^2}{3} \end{aligned} \]Now, we know both \(E[X]\) and \(E[X^2]\), so we can calculate \(Var[X]\):
\[ \begin{aligned} Var[X] &= E[X^2] - E[X]^2 \\ &= \dfrac{b^2+ab+a^2}{3} - \dfrac{(b+a)^2}{4} \\ &= \dfrac{b^2+ab+a^2}{3} - \dfrac{b^2+2ab+a^2}{4} \\ &= \dfrac{4b^2+4ab+4a^2-3b^2-6ab-3a^2}{12} \\ &= \dfrac{b^2-2ab+a^2}{12} \\ Var[X]&= \dfrac{(b-a)^2}{12} \end{aligned} \]It is the measure of how 2 variables X & Y vary together.
It gives the direction of the relationship between the variables.
Note: For both direction as well as magnitude, we use Correlation.
Let’s use expectation to compute the co-variance of two random variables X & Y:
Note:
- In a multivariate setting, relationship between all the pairs of random variables are summarized in
a square symmetrix matrix called ‘Co-Variance Matrix’ \(\Sigma\).
- Covariance of a random variable with self gives the variance, hence the diagonals of covariance matrix are variances.
End of Section
2.1.8 - Moment Generating Function
They are statistical measures that describe the characteristics of a probability distribution,
such as its central tendency, spread/variability, and asymmetry.
Note: We will discuss ‘raw’ and ‘central’ moments.
Important moments:
- Mean : Central tendency
- Variance : Spread/variability
- Skewness : Asymmetry
- Kurtosis : Tailedness
It is a function that simplifies computation of moments, such as, the mean, variance, skewness, and kurtosis,
by providing a compact way to derive any moment of a random variable through differentiation.
- Provides an alternative to PDFs and CDFs of random variables.
- A powerful property of the MGF is that its \(n^{th}\) derivative, when evaluated at \((t=0)\) = the \(n^{th}\) moment of the random variable.
Not all probability distributions have MGFs; sometimes, integrals may not converge and moment may not exist.
Since, \(MGF = M_X(t) = E[e^{tX}] \), we can write the MGF as:
Say, \( E[X^n] = m_n \), where \(m_n\) is the \(n^{th}\) moment of the random variable, then:
\[ M_X(t) = 1 + \frac{t}{1!}m_1 + \frac{t^2}{2!}m_2 + \frac{t^3}{3!}m_3 + \cdots + \frac{t^n}{n!}m_n + \cdots \\ \]Important: Differentiating \(M_X(t)\) with respect to \(t\), \(i\) times, and setting \((t =0)\) gives us the \(i^{th}\) moment of the random variable.
\[ \frac{dM_X(t) }{dt} = M'_X(t) = 0 + m_1 + \frac{2t}{2!}m_2 + \frac{3t^2}{3!}m_3 + \cdots + \frac{nt^n-1}{n!}m_n + \cdots \\ \]Set \(t=0\), to get the first moment:
\[ \frac{dM_X(t) }{dt} = M'_X(t) = m_1 \]Similarly, second moment = \(M''_X(t) = m_2 = E[X^2]\)
And, \(n^{th}\) derivative = \(M^{(n)}_X(t) = m_n = E[X^n]\)
e.g: \( Var[X] = M''_X(0) - (M'_X(0))^2 \)
Note: If we know the MGF of a random variable, then we do NOT need to do integration or summation to find the moments.
Continuous:
Discrete:
\[ M_X(t) = E[e^{tX}] = \sum_{i=1}^{\infty} e^{tx} P(X=x) \]End of Section
2.1.9 - Joint & Marginal
It describes the probability of 2 or more random variables occurring simultaneously.
- The random variables can be from different distributions, such as, discrete and continuous.
Joint CDF:
Discrete Case:
Continuous Case:
Joint PMF:
Key Properties:
- \(P(X = x, Y = y) \ge 0 ~ \forall (x,y) \)
- \( \sum_{i} \sum_{j} P(X = x_i, Y = y_j) = 1 \)
Joint PDF:
Key Properties:
- \(f_{X,Y}(x,y) \ge 0 ~ \forall (x,y) \)
- \( \int_{-\infty}^{\infty} \int_{-\infty}^{\infty} f_{X,Y}(x,y) dy dx = 1 \)
- If we consider 2 random variables, say, height(X) and weight(Y), then the joint distribution will tell us the probability of finding a person having a particular height and weight.
A ball is picked at random from each bag, such that both draws are independent of each other.
Let’s use this example to understand joint probability.
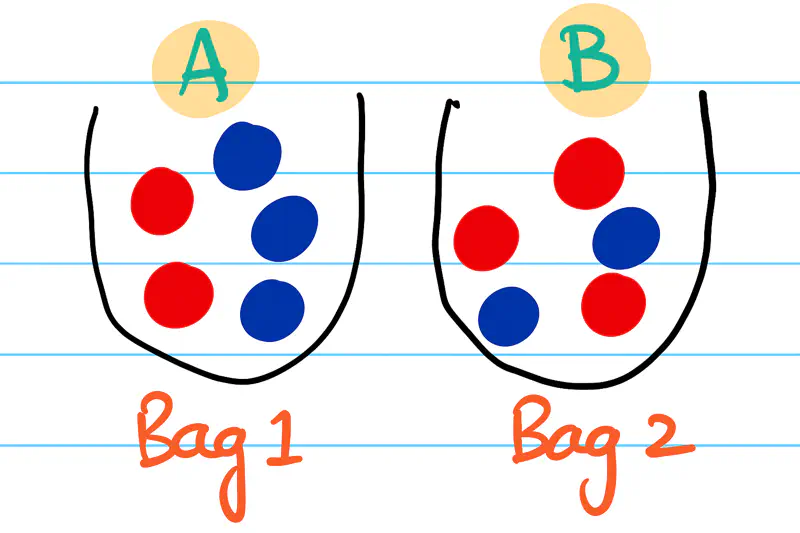
Let A & B be discrete random variables associated with the outcome of the ball drawn from first and second bags
respectively.
| A = Red | A = Blue | |
|---|---|---|
| B = Red | 2/5*3/5 = 6/25 | 3/5*3/5 = 9/25 |
| B = Blue | 2/5*2/5 = 4/25 | 3/5*2/5 = 6/25 |
Since, the draws are independent, joint probability = P(A) * P(B)
Each of the 4 cells in above table shows the probability of combination of results from 2 draws or joint probability.
It describes the probability distribution of an individual random variable in a joint distribution,
without considering the outcomes of other random variables.
- If we have the joint distribution, then we can get the marginal distribution of each random variable from it.
- Marginal probability equals summing the joint probability across other random variables.
Marginal CDF:
We know that Joint CDF =
Marginal CDF =
\[ F_X(a, \infty) = P(X \le a, Y < \infty) = P(X \le a) \]Discrete Case:
Continuous Case:
Law of Total Probability
We know that Joint Probability Distribution =
The events \((Y=y)\) partition the sample space, such that:
- \( (Y=y_1) \cap (Y=y_2) \cap ... \cap (Y=y_n) = \Phi \)
- \( (Y=y_1) \cup (Y=y_2) \cup ... \cup (Y=y_n) = \Omega \)
From Law of Total Probability, we get:
Marginal PMF:
Marginal PDF:
X: Roll a die ; \( \Omega = \{1,2,3,4,5,6\} \)
Y: Toss a coin ; \( \Omega = \{H,T\} \)
Joint PMF = \( P_{X,Y}(x,y) = P(X=x, Y=y) = 1/6*1/2 = 1/12\)
Marginal PMF of X = \( P_X(x) =\sum_{y \in \mathbb{\{H,T\}}} P_{X,Y}(x,y) = = 1/12+1/12 = 1/6\)
=> Marginally, X is uniform over 1-6 i.e a fair die.
Marginal PMF of Y = \( P_Y(y) = \sum_{1}^6 P_{X,Y}(x,y) = 6*(1/12) = 1/2 \)
=> Marginally, Y is uniform over H,T i.e a fair coin.
\( X \sim U(0,1) \)
\( Y \sim U(0,1) \)
Marginal PDF = \(f_X(x) = \int_{-\infty}^{\infty} f_{X,Y}(x,y) dy \)
Joint PDF =
Marginal PDF =
\[ \begin{aligned} f_X(x) &= \int_{0}^{1} f_{X,Y}(x,y) dy \\ &= \int_{0}^{1} 1 dy \\ &= 1 \\ f_X(x) &= \begin{cases} 1 & \text{if } x \in [0,1] \\ 0 & \text{otherwise } \end{cases} \end{aligned} \]There are 2 bags; bag_1 has 2 red balls & 3 blue balls, bag_2 has 3 red balls & 2 blue balls.
A ball is picked at random from each bag, such that both draws are independent of each other.
Let’s use this example to understand marginal probability.

Let A & B be discrete random variables associated with the outcome of the ball drawn from first and second bags
respectively.
| A = Red | A = Blue | P(B) (Marginal) | |
|---|---|---|---|
| B = Red | 2/5*3/5 = 6/25 | 3/5*3/5 = 9/25 | 6/25 + 9/25 = 15/25 = 3/5 |
| B = Blue | 2/5*2/5 = 4/25 | 3/5*2/5 = 6/25 | 4/25 + 6/25 = 10/25 = 2/5 |
| P(A) (Marginal) | 6/25 + 4/25 = 10/25 = 2/5 | 9/25 + 6/25 = 15/25 = 3/5 |
We can see from the table above - P(A=Red) is the sum of joint distribution over all possible values of B i.e Red & Blue.
It measures the probability of an event occurring given that another event has already happened.
- It provides a way to update our belief about the likelihood based on new information.
P(A, B) = Joint Probability of A and B
P(B) = Marginal Probability of B
=> Conditional Probability = Joint Probability / Marginal Probability
Conditional CDF:
Discrete Case:
Continuous Case:
Conditional PMF:
Conditional PDF:
- Generative machine learning models, such as, GANs, learn the conditional distribution of pixels, given the style of input image.
Note: We only have information about the joint and marginal probabilities.
What is the conditional probability of drawing a red ball in the first draw, given that a blue ball is drawn in second draw?

Let A & B be discrete random variables associated with the outcome of the ball drawn from first and second bags
respectively.
A = Red ball in first draw
B = Blue ball in second draw.
| A = Red | A = Blue | P(B) (Marginal) | |
|---|---|---|---|
| B = Red | 6/25 | 9/25 | 3/5 |
| B = Blue | 4/25 | 6/25 | 2/5 |
| P(A) (Marginal) | 2/5 | 3/5 |
Therefore, probability of drawing a red ball in the first draw, given that a blue ball is drawn in second draw = 2/5.
This gives us the conditional expectation of a random variable X, given another random variable Y=y.
Discrete Case:
Continuous Case:
- Conditional expectation of of a person’s weight, given his/her height = 165 cm, will give us the average weight of all people with height = 165 cm.
Applications:
- Linear regression algorithm is conditional expectation of target variable ‘Y’, given input feature variable ‘X’.
- Expectation Maximisation(EM) algorithm is built on conditional expectation.
This gives us the variance of a random variable calculated after taking into account the value(s) of another related variable.
For example:
- Variance of car’s mileage for city driving might be small, but the variance will be large for mix of city and highway driving.
Note: Models that take into account the change in variance or heteroscedasticity tend to be more accurate.
End of Section
2.1.10 - Independent & Identically Distributed
Let’s revisit and understand the independence of random variables first.
It means that the knowing the outcome of one random variable does not impact the probability of the other random variable.
Two random variables X & Y are independent if:
- We know that if 2 random variables X,Y are independent, then their covariance is zero,
since there is no linear relationship between them.
- But the converse may NOT be true, i.e, if the covariance is zero, then we can say for sure that the random variables
are independent.
Let’s go through few examples to understand the independence of random variables.
For example:
- Toss a coin + Roll a die.
\[ X = \begin{cases} 1 & \text{if Heads} \\ \\ 0 & \text{if Tails} \end{cases} \\ \text{} \\ Y = \{1,2,3,4,5,6\} \\ \text{} \\ \text{ Joint probability is all possible combinations of X and Y i.e 2x6 = 12 } \\[10pt] \text{ Sample space: } \Omega = \{ (H,1), (H,2), (H,3), (H,4), (H,5), (H,6), \\ (T,1), (T,2), (T,3), (T,4), (T,5), (T,6) \} \]
Here, clearly, X and Y are independent.
- Toss a coin twice.
\(X\) = Number of heads = \(\{0,1,2\}\)
\(Y\) = Number of tails = \(\{0,1,2\}\)
X, Y are NOT independent, but mutually exclusive, because if we know about one, then we automatically know about the other.
- \(X\) is a continuous uniform random variable \( X \sim U(-1,1) \).
\(Y = 2X\).
Let’s check for independence of \(X\) and \(Y\) i.e \(E[XY] = E[X]E[Y]\) or not?
Now, lets calculate the value of \(E[XY]\):
From (1) and (2), we can say that:
\[ E[XY] ⍯ E[X]E[Y] \\ \]Hence, \(X\) and \(Y\) are NOT independent.
Read more about Integration
Similarly, random variables X & Y are said to be identically distributed if they belong to the same probability distribution.
I.I.D assumption for samples(data points) in a dataset means that the samples are:
- Independent, i.e, each sample is independent of the other.
- Identically distributed, i.e, each sample is drawn from the same probability distribution.
- We take the heights of a random sample of people to estimate the average height of the population of a city.
- Here ‘independent’ assumption means that the height of each person in the sample is independent of the other person.
Usually, heights of members of the same family may be highly correlated.
However, for practical purposes, we can assume that all the heights are independent of one another. - And, for ‘identically distributed’ - we can assume that all the heights are from the same Gaussian distribution with some mean and variance.
- Here ‘independent’ assumption means that the height of each person in the sample is independent of the other person.
End of Section
2.1.11 - Convergence
A sequence of random variables \(X_1, X_1, \dots, X_n\) is said to converge in probability
to a known random variable \(X\),
if for every number \(\epsilon >0 \), the following is true:
where,
\(X_n\): is the estimator or sample based random variable.
\(X\): is the known or limiting or target random variable.
\(\epsilon\): is the tolerance level or margin of error.
- Toss a fair coin:
Estimator:
\[ X_n = \begin{cases} \frac{n}{n+1} & \text{, if Head } \\ \\ \frac{1}{n} & \text{, if Tail} \\ \end{cases} \]
Known random variable (Bernoulli):
\[ X = \begin{cases} 1 & \text{, if Head } \\ \\ 0 & \text{, if Tail} \\ \end{cases} \]
Say, tolerance level \(\epsilon = 0.1\).
Then,
If n=5;
So, if n=5, then \(|X_n - X| > (\epsilon = 0.1)\).
\( \implies P(|X_n - X| \ge (\epsilon=0.1)) = 1\).
if n=20;
So, if n=20, then \(|X_n - X| < (\epsilon=0.1)\).
\(\implies P(|X_n - X| \ge (\epsilon=0.1)) = 0\).
\( \implies P(|X_n - X| \ge (\epsilon=0.1)) = 0 ~\forall ~n \ge 10\)
Therefore,
Similarly, we can prove that if \(\epsilon = 0.01\), then the probability will be equal to 0 for \( n\ge 100 \).
Note: Task is to check whether the sequence of randome variables \(X_1, X_2, \dots, X_n\) converges in probability to a known random variable \(X\) as \(n\rightarrow\infty\).
So, we can conclude that, if \(n > \frac{1}{\epsilon}\), then:
A sequence of random variables \(X_1, X_2, \dots, X_n\) is said to almost surely converge to a known random variable \(X\),
for \(n \ge 1\), if the following is true:
where,
\(X_n\): is the estimator or sample based random variable.
\(X\): is the known or limiting or target random variable.
If, \(X_n \xrightarrow{Almost ~ Sure} X \), \( \implies X_n \xrightarrow{Probability} X \)
But, converse is NOT true.
*Note: Almost Sure convergence is hardest to satisfy amongst all convergence, such as, convergence in probability,
convergence in distribution, etc.
Read more about Limits
- \(X\) is random variable such that \(X = \frac{1}{2} \), a constant, i.e \(X_1 = X_2 = \dots = X_n = \frac{1}{2}\).
\(Y_1, Y_2,\dots ,Y_n \) are another sequence of random variables, such that :
\[ Y_1 = X_1 \\[10pt] Y_2 = \frac{X_1 + X_2}{2} \\[10pt] Y_3 = \frac{X_1 + X_2 + X_3}{3} \\[10pt] \dots \\ Y_n = \frac{1}{n} \sum_{i=1}^{n} X_i \xrightarrow{Almost ~ Sure} \frac{1}{2} \]
End of Section
2.1.12 - Law of Large Numbers
This law states that given a sequence of independent and identically distributed (IID) samples \(X_1, X_1, \dots, X_n\) from a random variable with finite mean, the sample mean (\(\bar{X_n}\)) converges in probability to the expected value \(E[X]\) or population mean (\( \mu \)).
\[ \lim_{n\rightarrow\infty} P(|\bar{X_n} - E[X]| \ge \epsilon) = 0, \forall ~ \epsilon >0 \\[10pt] \\[10pt] \frac{1}{n} \sum_{i=1}^{n} X_i \xrightarrow{Probability} E[X], \text{ as } n \rightarrow \infty \]Note: Does NOT guarantee that sample mean will be close to population mean,
but instead says that - the probability of sample mean being far away from the population mean is low.
Read more about Limits
- Toss a coin large number of times \('n'\), as \(n \rightarrow \infty\), the proportion of heads will probably be very
close to \(0.5\).
However, it does NOT rule out the possibility of a rare sequence, e.g., getting 10 consecutive heads.
But, the probability of such a rare event is extremely low.
This law states that given a sequence of independent and identically distributed (IID) samples \(X_1, X_1, \dots, X_n\) from a random variable with finite mean, the sample mean (\(\bar{X_n}\)) converges almost surely to the expected value \(E[X]\) or population mean (\( \mu \)).
\[ P(\lim_{n\rightarrow\infty} \bar{X_n} = E[X]) = 1, \text{ as } n \rightarrow \infty \\[10pt] \frac{1}{n} \sum_{i=1}^{n} X_i \xrightarrow{Almost ~ Sure} E[X], \text{ as } n \rightarrow \infty \]Note:
- It guarantees that the sequence of sample averages itself converges to population mean, with exception of set of outcomes that has probability = 0.
- Almost certain guarantee; Much stronger statement than Weak Law of Large Numbers.
- Toss a coin large number of times \('n'\), as \(n \rightarrow \infty\), the proportion of heads will converge
to \(0.5\), with probability = 1.
This means that a sequence where the proportion of heads never settles down to 0.5, is a probability = 0 event.
- Almost sure convergence ensures ML model’s reliability by guaranteeing that the average error on a large dataset will
converge to the true error.
Thus, providing confidence that model will perform consistently and accurately on unseen data.
End of Section
2.1.13 - Markov's Inequality
It gives an upper bound for the probability that a non-negative random variable will NOT exceed, based on its expected value.
Note: It gives a very loose upper bound.
Read more about Expectation
What is the probability that the restaurant will serve more than 200 customers in the next hour?
Hence, a 25% chance of serving more than 200 customers.
What is the probability that a randomly selected student gets a score of 90 marks or more?
Hence, there is a 78% chance that a randomly selected student gets a score of 90 marks or more.
It states that the probability of a random variable deviating from its mean is small if its variance is small.
- It is a more powerful version of Markov’s Inequality.
- It uses variance of the distribution in addition to the expected value or mean.
- Also, it does NOT assume the random variable to be non-negative.
- It uses more information about the data i.e mean and variance.
Note: It gives a tighter upper bound than Markov’s Inequality.
What is the probability that a randomly selected student gets a score of 90 marks or more?
Given the standard deviation of the test score is 10 marks.
Given, the standard deviation of the test score \(\sigma\) = 10 marks.
=> Variance = \(\sigma^2\) = 100
Hence, Chebyshev’s Inequality gives a far tighter upper bound of 25% than Markov’s Inequality of 78%(approx).
It is an upper bound on the probability that a random variable deviates from its expected value.
It’s an exponentially decreasing bound on the “tail” of a random variable’s distribution,
which can be calculated using its moment generating function.
- It is used for sum or average of independent random variables (not necessarily identically distributed).
- It provides exponentially tighter bounds, better than Chebyshev’s Inequality’s quadratic decay.
- It uses all moments to capture the full shape of the distribution, using the moment generating function(MGF).
Proof:
For the sum of ’n’ independent random variables,
Note: Used to compute how far the sum of independent random variables deviate from their expected value.
Read more about Moment Generating Function
End of Section
2.1.14 - Cross Entropy & KL Divergence
It is a measure of the amount of information gained when a specific, individual event occurs, and is defined based on
the probability of that event.
It is defined as the negative logarithm of the probability of the event.
- It is also called ‘Surprisal’.
Note: Logarithm(for base > 1) is a monotonically increasing function, so as x increases, log(x) also increases.
- So, if probability P(x) increases, then surprise factor S(x) decreases.
- Common events have a high probability of occurrence, hence a low surprise factor.
- Rare events have a low probability of occurrence, hence a high surprise factor.
Units:
- The unit of surprise factor with log base 2 is bits.
- with base ’e’ or natural log its nats (natural units of information).
It conveys how much ‘information’ we expect to gain from a random event.
- Entropy is the average or expected value of surprise factor of a random variable.
- More uniform the distribution ⇒ greater average surprise factor, since all possible outcomes are equally likely.
Case 1:
Toss a fair coin; \(P(H) = P(T) = 0.5 = 2^{-1}\)
Surprise factor (Heads or Tails) = \(-log(P(x)) = -log_2(0.5) = -log_2(2^{-1}) = 1~bit\)
Entropy = \( \sum_{x \in X} P(x)log(-P(x)) = 0.5*1 + 0.5*1 = 1 ~bit \)Case 2:
Toss a biased coin; \(P(H) = 0.9 P(T) = 0.1 \)
Surprise factor (Heads) = \(-log(P(x)) = -log_2(0.9) \approx 0.15 ~bits \)
Surprise factor (Tails) = \(-log(P(x)) = -log_2(0.1) \approx 3.32 ~bits\)
Entropy = \( \sum_{x \in X} P(x)log(-P(x)) = 0.9*0.15 + 0.1*3.32 \approx 0.47 ~bits\)
Therefore, a biased coin is less surprising on an average than a fair coin, hence lower entropy.
It is a measure of the average ‘information gain’ or ‘surprise’ when using an imperfect model \(Q\) to encode events
from a true model \(P\).
- It measures how surprised we are on an average, if the true distribution is \(P\), but we predict using another distribution
\(Q\).
A model is trained to classify images as ‘cat’ or ‘dog’. Say, for an input image the true label is ‘cat’,
so the true distribution:
\(P = [1.0 ~(cat), 0.0 ~(dog)]\).
Let’s calculate the cross-entropy for the outputs of 2 models A & B.
Model A:
\(Q_A = [0.8 ~(cat), 0.2 ~(dog)]\)
Cross Entropy = \(H(P, Q_A) = -\sum_{i=1}^n P(x_i)log(Q_A(x_i)) \)
\(= -[1*log_2(0.8) + 0*log_2(0.2)] \approx 0.32 ~bits\)
Note: This is very low cross-entropy, since the predicted value is very close to actual, i.e low surprise.Model B:
\(Q_B = [0.2 ~(cat), 0.8 ~(dog)]\)
Cross Entropy = \(H(P, Q_B) = -\sum_{i=1}^n P(x_i)log(Q_B(x_i)) \)
\(= -[1*log_2(0.2) + 0*log_2(0.8)] \approx 2.32 ~bits\)
Note: Here the cross-entropy is very high, since the predicted value is quite far from the actual truth, i.e high surprise.
It measures the information lost when one probability distribution \(Q\) is used to approximate another distribution \(P\).
It quantifies the ‘extra cost’ in bits needed to encode data using the approximate distribution \(Q\) instead of
the true distribution \(P\).
KL Divergence = Cross Entropy(P,Q) - Entropy(P)
\[ \begin{aligned} D_{KL}(P \parallel Q) &= H(P, Q) - H(P) \\ &= -\sum_{i=1}^n P(x_i)log(Q(x_i)) - [-\sum_{i=1}^n P(x_i)log(P(x_i))] \\[10pt] &= \sum_{i=1}^n P(x_i)[log(P(x_i)) - log(Q(x_i)) ] \\[10pt] D_{KL}(P \parallel Q) &= \sum_{i=1}^n P(x_i)log(\frac{P(x_i)}{Q(x_i)}) \\[10pt] \text{ For continuous case: } \\ D_{KL}(P \parallel Q) &= \int_{-\infty}^{\infty} p(x)log(\frac{p(x)}{q(x)})dx \end{aligned} \]Note:
- If \(P = Q\) ,i.e, P and Q are the same distributions, then KL Divergence = 0.
- KL divergence is NOT symmetrical ,i.e, \(D_{KL}(P \parallel Q) ⍯ (D_{KL}(Q \parallel P)\).
Using the same cat, dog classification problem example as mentioned above.
A model is trained to classify images as ‘cat’ or ‘dog’. Say, for an input image the true label is ‘cat’,
so the true distribution:
\(P = [1.0 ~(cat), 0.0 ~(dog)]\).
Let’s calculate the KL divergence for the outputs of the 2 models A & B.
Let’s calculate entropy first, and we can reuse the cross-entropy values calculated above already.
Entropy: \(H(P) = -\sum_{x \in X} P(x)log(P(x)) = -[1*log_2(1) + 0*log_2(0)] = 0 ~bits\)
Note: \(0*log_2(0) = 0\), is an approximation hack, since \(log(0)\) is undefined.
Model A:
\( D_{KL}(P \parallel Q) = H(P, Q) - H(P) = 0.32 - 0 = 0.32 ~bits \)
Model A incurs an additional 0.32 bits of surprise due to its imperfect prediction.Model B:
\( D_{KL}(P \parallel Q) = H(P, Q) - H(P) = 2.32 - 0 = 2.32 ~bits \)
Model B has much more ‘information loss’ or incurs higher ‘penalty’ of 2.32 bits as its prediction was from from the truth.
It is a smoothed and symmetric version of the Kullback-Leibler (KL) divergence and is calculated by averaging the
KL divergences between each of the two distributions and their combined average distribution.
- Symmetrical and smoothed version of KL divergence.
- Always finite; KL divergence can be infinite if \( P ⍯ 0 ~and~ Q = 0 \).
- Makes JS divergence more stable for ML models where some predicted probabilities may be exactly 0.
Let’s continue the cat and dog image classification example discussed above.
A model is trained to classify images as ‘cat’ or ‘dog’. Say, for an input image the true label is ‘cat’,
so the true distribution:
\(P = [1.0 ~(cat), 0.0 ~(dog)]\).
Step 1: Calculate the average distribution M.
Step 2: Calculate \(D_{KL}(P \parallel M)\).
Step 3: Calculate \(D_{KL}(Q \parallel M)\).
Step 4: Finally, lets put all together to calculate \(D_{JS}(P \parallel Q)\).
Therefore, lower JS divergence value => P and Q are more similar.
End of Section
2.1.15 - Parametric Model Estimation
It is the process of finding the best-fitting finite set of parameters \(\Theta\) for a model that assumes a
specific probability distribution for the data.
It involves using the dataset to estimate the parameters (like the mean and standard deviation of a normal distribution)
that define the model.
\(P(X \mid \Theta) \) : Probability of seeing data \(X: (X_1, X_2, \dots, X_n) \), given the parameters \(\Theta\) of the
underlying probability distribution from which the data is assumed to be generated.
Goal of estimation:
We observed data, \( D = \{X_1, X_2, \dots, X_n\} \), and we want to infer the the unknow parameters \(\Theta\)
of the underlying probability distribution, assuming that the data is generated I.I.D.
Read more for I.I.D
Note: Most of the time, from experience, we know the underlying probability distribution of data, such as, Bernoulli, Gaussian, etc.
There are 2 philosophical approaches to estimate the parameters of a parametric model:
- Frequentist:
- Parameters \(\Theta\) is fixed, but unknown, only data is random.
- It views probability as the long-run frequency of events in repeated trials; e.g toss a coin ’n’ times.
- It is favoured when the sample size is large.
- For example, Maximum Likelihood Estimation(MLE), Method of Moments, etc.
- Bayesian:
- Parameters \(\Theta\) itself is unknown, so we model it as a random variable with a probability distribution.
- It views probability as a degree of belief that can be updated with new evidence, i.e. data.
Thus, integrating prior knowledge with data to express the uncertainty about the parameters. - It is favoured when the sample size is small, as it uses prior belief about the data distribution too.
- For example, Maximum A Posteriori Estimation(MAP), Minimum Mean Square Error Estimation(MMSE), etc..
It is the most popular frequentist approach to estimate the parameters of a model.
This method helps us find the parameters \(\Theta\) that make the data most probable.
Likelihood Function:
Say, we have data, \(D = X_1, X_2, \dots, X_n\) are I.I.D discrete random variable with PMF = \(P_{\Theta}(.)\)
Then, the likelihood function is the probability of observing the data, \(D = \{X_1, X_2, \dots, X_n\}\),
given the parameters \(\Theta\).
Task:
Find the value of parameter \(\Theta_{ML}\) that maximizes the likelihood function.
In order to find the parameter \(\Theta_{ML}\) that maximises the likelihood function,
we need to take the first derivative of the likelihood function with respect to \(\Theta\) and equate it to zero.
But, taking derivative of a product is challenging, so we will take the logarithm on both sides.
Note: Log is a monotonically increasing function, i.e, as x increases, log(x) increases too.
Let us denote the log-likelihood function as \(\bar{L}\).
\[ \begin{aligned} \mathcal{\bar{L}_{X_1, X_2, \dots ,X_n}}(\Theta) &= log [\prod_{i=1}^{n} P_{\Theta}(X_i)] \\ &= \sum_{i=1}^{n} log P_{\Theta}(X_i) \\ \end{aligned} \]Therefore, Maximum Likelihood Estimation is the parameter \(\Theta_{ML}\) that maximises the log-likelihood function.
\[ \Theta_{ML}(X_1, X_2, \dots, X_n) = \underset{\Theta}{\mathrm{argmax}}\ \bar{L}_{X_1, X_2, \dots ,X_n}(\Theta) \]Given \(X_1, X_2, \dots, X_n\) are I.I.D. Bernoulli random variable with PMF as below:
Estimate the parameter \(\theta\) using Maximum Likelihood Estimation.
Let, \(n_1 = \) number of 1’s in the dataset.
Likelihood Function:
Log-Likelihood Function:
Maximum Likelihood Estimation:
In order to find the parameter \(\theta_{ML}\), we need to take the first derivative of the log-likelihood function
with respect to \(\theta\) and equate it to zero.
Read more about Derivative
Say, e.g., we have 10 observations for the Bernoulli random variable as: 1,0,1,1,0,1,1,0,1,0.
Then, the parameter \(\Theta_{ML} = \frac{6}{10} = 0.6\) i.e proportion of 1’s.
\(x_1, x_2, \dots, x_n\) are the realisations/observations of the random variable.
Estimate the parameters \(\mu\) and \(\sigma\) of the Gaussian distribution using Maximum Likelihood Estimation.
Likelihood Function:
Log-Likelihood Function:
Note: Here, the first term \( -\frac{n}{2} log (2\pi) \) is a constant wrt both \(\mu\) and \(\sigma\), so we can ignore the term.
Maximum Likelihood Estimation:
Instead of finding \(\mu\) and \(\sigma\) that maximises the log-likelihood function,
we can find \(\mu\) and \(\sigma\) that minimises the negative of the log-likelihood function.
Now, lets differentiate the log likelihood function wrt \(\mu\) and \(\sigma\) separately to get \(\mu_{ML}, \sigma_{ML}\).
Lets, calculate \(\mu_{ML}\) first by taking the derivative of the log-likelihood function wrt \(\mu\) and equating it to 0.
Read more about Derivative
Similarly, we can calculate \(\sigma_{ML}\) by taking the derivative of the log-likelihood function wrt \(\sigma\)
and equating it to 0.
Note: In general MLE is biased, i.e does NOT give an unbiased estimate => divides by \(n\) instead of \((n-1)\).
Bayesian statistics model parameters by updating initial beliefs (prior probabilities) with observed data to form
a final belief (posterior probability) using Bayes’ Theorem.
Instead of a single point estimate, it provides a probability distribution over possible parameter values,
which allows to quantify uncertainty and yields more robust models, especially with limited data.
Bayes’ Theorem:
Read more about Bayes’ Theorem
\(P(\Theta)\): Prior: Initial distribution of \(\Theta\) before seeing the data.
\(P(X \mid \Theta)\): Likelihood: Conditional distribution of data \(X\), given the parameter \(\Theta\).
\(P(\Theta \mid X)\): Posterior: Conditional distribution of parameter \(\Theta\), given the data \(X\).
\(P(X)\): Evidence: Probability of seeing the data \(X\).
\(x_1, x_2, \dots, x_n\) are the realisations, \(\Theta \in [0, 1] \).
Estimate the parameter \(\Theta\) using Bayesian statistics.
Let, \(n_1 = \) number of 1’s in the dataset.
Prior: \(P(\Theta)\) : \(\Theta \sim U(0, 1) \), i.e, parameter \(\Theta\) comes from a continuos uniform
distribution in the range [0,1].
\(f_{\Theta}(\theta) = 1, \theta \in [0, 1] \)
Likelihood: \(P_{X \mid \Theta}(x \mid \theta) = \theta^{n_1} (1 - \theta)^{n - n_1} \).
Posterior:
\[ f_{\Theta \mid X} (\theta \mid x) = \frac{f_{\Theta}(\theta) P_{X \mid \Theta}(x \mid \theta)}{f_{X}(x)} \\ \]Note: The most difficult part is to calculate the denominator \(f_{X}(x)\).
So, either we try NOT to compute it all together, or we try to map it to some known functions to make calculations easier.
We know that we can get the marginal probability by integrating the joint probability over another variable.
Also from conditional probability, we know:
\[ \tag{2} f_{X \mid Y}(x \mid y) = \frac{f_{X,Y}(x,y)}{f_{Y}(y)} \\ => f_{X,Y}(x,y) = f_{Y}(y) f_{X \mid Y}(x \mid y) \]From equations 1 and 2, we have:
\[ \tag{3} f_{X}(x) = \int_{Y}f_{Y}(y) f_{X \mid Y}(x \mid y)dy \]Now let’s replace the value of \(f_{X}(x) \) in the posterior from equation 3:
Posterior:
Beta Function:
It is a special mathematical function denoted by B(a, b) or β(a, b) that is defined by the integral formula:
Note: We can see that the denominator of the posterior is of the form of Beta function.
Posterior:
Suppose, in the above example, we are told that the parameter \(\Theta\) is closer to 1 than 0.
How will we incorporate this useful information (apriori knowledge) into our parameter estimation?
Since, we know apriori that \(\Theta\) is closer to 1 than 0, we should take this initial belief into account
to do our parameter estimation.
Prior:
Posterior:
\[ \begin{aligned} f_{\Theta \mid X} (\theta \mid x) &= \frac{f_{\Theta}(\theta) P_{X \mid \Theta}(x \mid \theta)}{f_{X}(x)} \\[10pt] &= \frac{f_{\Theta}(\theta) P_{X \mid \Theta}(x \mid \theta)}{\int_{\Theta}f_{\Theta}(\theta) P_{X \mid \Theta}(x \mid \theta)d\theta} \\[10pt] &= \frac{f_{\Theta}(\theta) P_{X \mid \Theta}(x \mid \theta)}{\int_{0}^1f_{\Theta}(\theta) P_{X \mid \Theta}(x \mid \theta)d\theta} \\[10pt] \text{ We know that: } f_{\Theta}(\theta) = 2\Theta, \theta \in [0, 1] \\ &= \frac{2\Theta * P_{X \mid \Theta}(x \mid \theta)}{\int_{0}^1 2\Theta* P_{X \mid \Theta}(x \mid \theta)d\theta} \\[10pt] & = \frac{2\Theta * \theta^{n_1} (1 - \theta)^{n - n_1}}{\int_{0}^1 2\Theta * \theta^{n_1} (1 - \theta)^{n - n_1}} \\[10pt] => f_{\Theta \mid X} (\theta \mid x) &= \frac{\theta^{n_1+1} (1 - \theta)^{n - n_1}}{β(n_1+2, n-n_1+1)} \end{aligned} \]Note: If we do NOT have enough data, then we should NOT ignore our initial belief.
However, if we have enough data, then the data will override our initial belief and the posterior will be dominated by data.
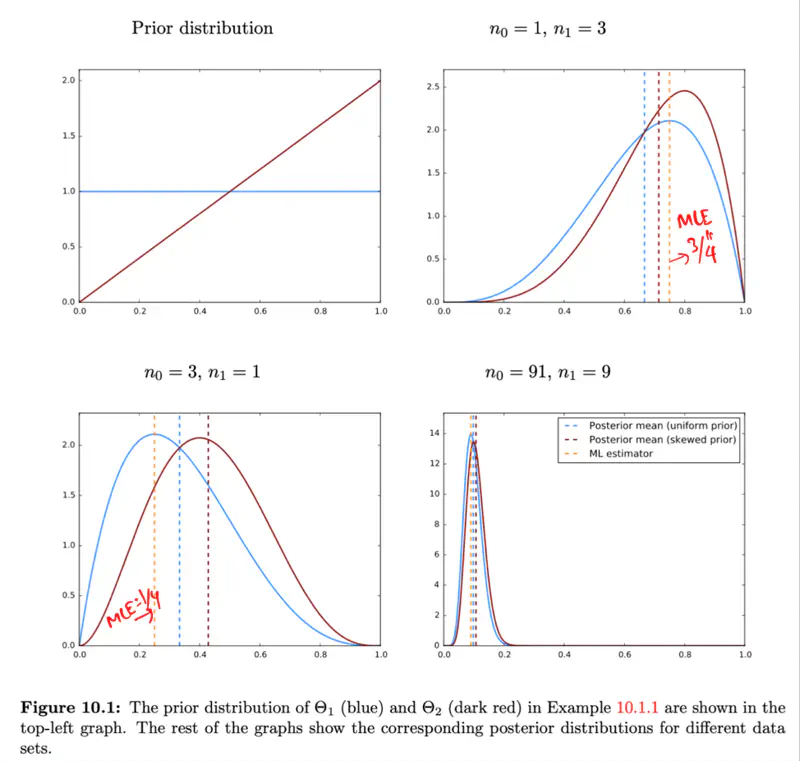
Note: Bayesian approach gives us a probability distribution of parameter \(\Theta\).
We can use Bayesian Point Estimators, after getting the posterior distribution, to summarize it with a single value for
practical use, such as,
- Maximum A Posteriori (MAP) Estimator
- Minimum Mean Square Error (MMSE) Estimator
It finds the mode(peak) of the posterior distribution.
- MAP has the minimum probability of error, since it picks single most probable value.
Estimate the unknown parameter \(\mu\) using MAP.
Given that :
\(\Theta\) is discrete, with probability 0.5 for both 0 and 1.
=> The Gaussian distribution is equally likely to be centered at 0 or 1.
Variance: \(\sigma^2 = 1\)
Prior:
Likelihood:
Posterior:
We need to find:
\[ \underset{\Theta}{\mathrm{argmax}}\ P_{\Theta \mid X} (\theta \mid x) \]Taking log on both sides:
Let’s calculate the log-likelihood function first -
\[ \begin{aligned} \log f_{X \mid \Theta}(x \mid \theta) &= \log \prod_{i=1}^n \frac{1}{\sqrt{2\pi}} e^{-\frac{(x_i-\theta)^2}{2}} \\ &= log(\frac{1}{\sqrt{2\pi}})^n + \sum_{i=1}^n \log (e^{-\frac{(x_i-\theta)^2}{2}}), \text{ since \(\sigma = 1\)} \\ &= n\log(\frac{1}{\sqrt{2\pi}}) + \sum_{i=1}^n -\frac{(x_i-\theta)^2}{2} \\ => \tag{2} \log f_{X \mid \Theta}(x \mid \theta) &= n\log(\frac{1}{\sqrt{2\pi}}) - \sum_{i=1}^n \frac{(x_i-\theta)^2}{2} \end{aligned} \]Here computing \(f_{X}(x)\) is difficult.
So, instead of differentiating above equation wrt \(\Theta\), and equating to 0,
we will calculate the above expression for \(\theta=0\) and \(\theta=1\) and compare the values.
This way we can get rid of the common value \(f_{X}(x)\) in both expressions that is not dependent on \(\Theta\).
When \(\theta=1\):
Similarly, when \(\theta=0\):
So, we can say that \(\theta = 1\) only if :
the value of the above expression for \(\theta = 1\) > the value of the above expression for \(\theta = 0\)
From equation 3 and 4:
Note: If the prior is uniform then \(\Theta_{MAP} = \Theta_{MLE}\), because uniform prior does NOT give any information about the initial bias, and all possibilities are equally likely.
What if, in the above example, we know that the initial belief is not uniform but biased towards 0?
Prior:
Now, let’s compare the log-posterior for both the cases i.e \(\theta=0\) and \(\theta=1\) as we did earlier.
But, note that this time the probabilities for \(\theta=0\) and \(\theta=1\) are different.
So, we can say that \(\theta = 1\) only if :
the value of the above expression for \(\theta = 1\) > the value of the above expression for \(\theta = 0\)
From equation 3 and 4 above:
Therefore, we can see that \(\Theta_{MAP}\) is extra biased towards 0.
Note: For a non-uniform prior \(\Theta_{MAP}\) estimate will be pulled towards the prior’s mode.
e.g: Patient has a certain disease or not.
But, what if we want to minimize average magnitude of errors, over time, say predicting a stock’s price?
Just getting to know whether the prediction was right or wrong is not sufficient here.
We also want to know that the prediction was wrong by how much, so that we can minimize the loss over time.
Minimizes the expected value of squared error.
Mean of the posterior distribution is the conditional expectation of parameter \(\Theta\), given the data.
- Posterior mean minimizes the the mean squared error.
\(\hat\Theta(X)\): Predicted value.
\(\Theta\): Actual value.
Let’s revisit the above examples that we used for MLE.
Case 1: Uniform Continuous Prior for parameter \(\Theta\) of Bernoulli distribution
Prior:
Posterior:
Let’s calculate the \(\Theta_{MMSE}\):
Similarly, for the second case where the prior is biased towards 1.
Case 2: Prior is biased towards 1 for parameter \(\Theta\) of Bernoulli distribution
Prior:
Posterior:
So, here in this case our \(\Theta_{MMSE}\) is:
- MMSE is the average of the posterior distribution, whereas MAP is the mode/peak.
- If posterior distribution is symmetric and unimodal (only 1 peak), then MAP and MMSE are very close.
- If posterior distribution is skewed and multimodal (many peaks), then MAP and MMSE can differ a lot.
- MMSE considers all the values of the posterior distribution; hence, it is more accurate than MAP, especially for skewed or multimodal distributions.
End of Section
2.2.1 - Data Distribution
A single number that describes the central, typical, or representative value of a dataset, e.g, mean, median, and mode.
The mean is the average, the median is the middle value in a sorted list, and the mode is the most frequently occurring value.
- A single representative value can be used to compare different groups or distributions.
The artihmetic average of a set of numbers i.e sum all values and divide by the number of values.
\(mean = \frac{1}{n}\sum_{i=1}^{n}x_i\)
- Most common measure of central tendency.
- Represents the ‘balancing point’ of data.
- Sample mean is denoted by \(\bar{x}\), and population mean by \(\mu\).
Pros:
- Uses all datapoints in its calculation, providing a comprehensive measure.
Cons:
- Highly sensitive to outliers i.e exteme values.
- mean\((1,2,3,4,5) = \frac{1+2+3+4+5}{5} = 3 \)
- With outlier: mean\((1,2,3,4,100) = \frac{1+2+3+4+100}{5} = \frac{110}{5} = 22\)
Note: Just a single extreme value of 100 has pushed the mean from 3 to 22.
The middle value of a sorted list of numbers. It divides the dataset into 2 equal halves.
Calculation:
- Arrange the data points in ascending order.
- If the number of data points is even, the median is the average of the two middle values.
- If the number of data points is odd, the median is the middle value i.e \((\frac{n+1}{2})^{th}\) element.
Pros:
- Not impacted by outliers, making it a more robust/reliable measure, especially for skewed distributions.
Cons:
- Does NOT use all the datapoints in its calculation.
- median\((1,2,3,4,5) = 3\)
- median\((1,2,3,4,5,6) = \frac{3+4}{2} = 3.5\)
- With outlier: median\((1,2,3,4,100) = 3\)
Note: No impact of outlier.
The most frequently occurring value in a dataset.
- Dataset can have 1 mode i.e unimodal, 2 modes i.e bimodal, and more than 2 modes i.e multimodal.
- If NO value repeats, then NO mode.
Pros:
- Only measure of central tendency that can be used for categorical/nominal data, such as, gender, blood group, level of education, etc.
- It can reveal important peaks in data distribution.
Cons:
- A dataset can have multiple modes, or no mode at all, which can make mode less informative.
Quantifies how spread out or scattered the data points are.
E.g: Range, Variance, Standard Deviation, Median Absoulute Deviation(MAD), Skewness, Kurtosis, etc.
The difference between the largest and smallest values in a dataset. Simplest measure of dispersion
\(range = max - min\)
Pros:
- Easy to calculate and understand.
Cons:
- Only considers the the 2 extreme values of dataset and ignores the distribution of data in between.
- Highly sensitive to outliers.
- range\((1,2,3,4,5) = 5 - 1 = 4\)
The average of the squared distance of each value from the mean.
Measures the spread of data points.
\(sample ~ variance = s^2 = \frac{1}{n}\sum_{i=1}^{n}(x_i - \bar{x})^2\)
\(population ~ variance = \sigma^2 = \frac{1}{n}\sum_{i=1}^{n}(x_i - \mu)^2\)
Cons:
- Highly sensitive to outliers, as squaring amplifies the weight of extreme data points.
- Less intuitive to understand, as the units are square of original units.
The square root of the variance, measures average distance of data points from the mean.
- Low standard deviation indicates that the data points are clustered around the mean, whereas
high standard deviation means that the data points are spread out over a wide range.
\(s = sample ~ standard ~ deviation \)
\(\sigma = population ~ standard ~ deviation \)
- Standard Deviation\((1,2,3,4,5) = \sqrt{\frac{1}{n}\sum_{i=1}^{n}(x_i - \bar{x})^2} \) \[ = \sqrt{\frac{1}{5}((1-3)^2 + (2-3)^2 + (3-3)^2 + (4-3)^2 + (5-3)^2)} \\ = \sqrt{\frac{1}{5}(4+1+0+1+4)} \\ = \sqrt{\frac{10}{5}} = \sqrt{2} = 1.414 \]
It is the average of absolute deviation or distance of all data points from mean.
\( mad = \frac{1}{n}\sum_{i=1}^{n}|x_i - \bar{x}| \)
Pros:
- Less sensitive to outliers as compared to standard deviation..
- More intuitive and simpler to understand.
- Mean Absolute Deviation\((1,2,3,4,5) = \\ \frac{1}{5}\left(\left|1-3\right| + \left|2-3\right| + \left|3-3\right| + \left|4-3\right| + \left|5-3\right|\right) = \frac{1}{5}\left(2+1+0+1+2\right) = \frac{6}{5} = 1.2\)
It measures the asymmetry of a data distribution.
Tells us whether the data is concentrated on one side of mean and is there a long tail stretching on the other side.
Positive Skew:
- Tail is longer on the right side of the mean.
- Bulk of data is on the left side of the mean, but there are a few very high values pulling the mean towards the right.
- Mean > Median > Mode.
Negative Skew:
- Tail is longer on the left side of the mean.
- Bulk of data is on the right side of the mean, but there are a few very high values pulling the mean towards the left.
- Mean < Median < Mode.
Zero Skew:
- Perfectly symmetrical like a normal distribution.
- Mean = Median = Mode.
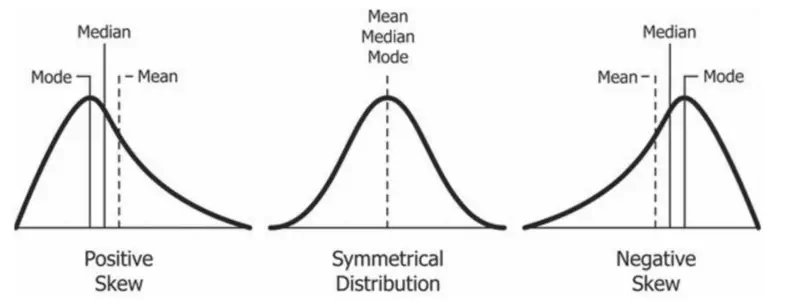
- Consider the salary of employees in a company. Most employees earn a very modest salary, but a few executives earn
extremely high salaries. This dataset will be positively skewed with the mean salary > median salary.
Median salary would be a better representation of the typical salary of employees.
It measures the “tailedness” of a data distribution.
It describes how much the data is concentrated in tails (fat or thin) versus the center.
- It can tell us about the frequency of outliers in the data.
- Thick tails => More outliers.
Excess Kurtosis:
Excess kurtosis is calculated by subtracting 3 from standard kurtosis in order to compare with normal distribution.
Normal distribution has kurtosis = 3.
Mesokurtic:
- Excess kurtosis = 0 i.e normal kurtosis.
- Tails are neither too thick nor too thin.
Leptokurtic:
- High kurtosis, i.e, excess kurtosis > 0 (+ve).
- Heavy or thick tails => High probability of outliers.
- Sharp peak => High concentration of data around mean.
- E.g: Student’s t-distribution, Laplace distribution, etc.
- High risk stock portfolios.
Platykurtic:
- Low kurtosis, i.e, excess kurtosis < 0 (-ve).
- Thin tails => Low probability of outliers.
- Low peak => more uniform distribution of values.
- E.g: Uniform distribution, Bernoulli(P=0.5) distribution, etc.
- Investment in fixed deposits.
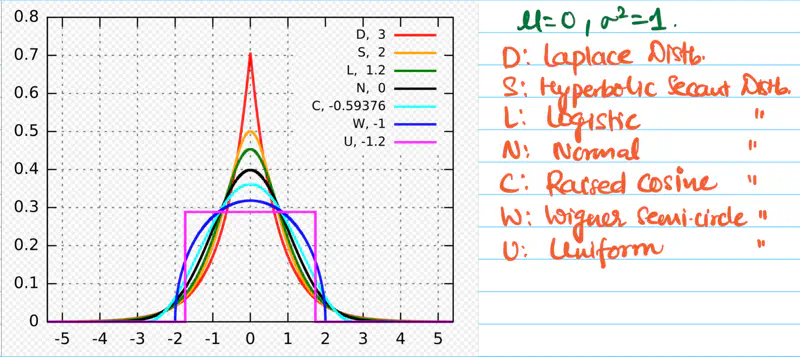
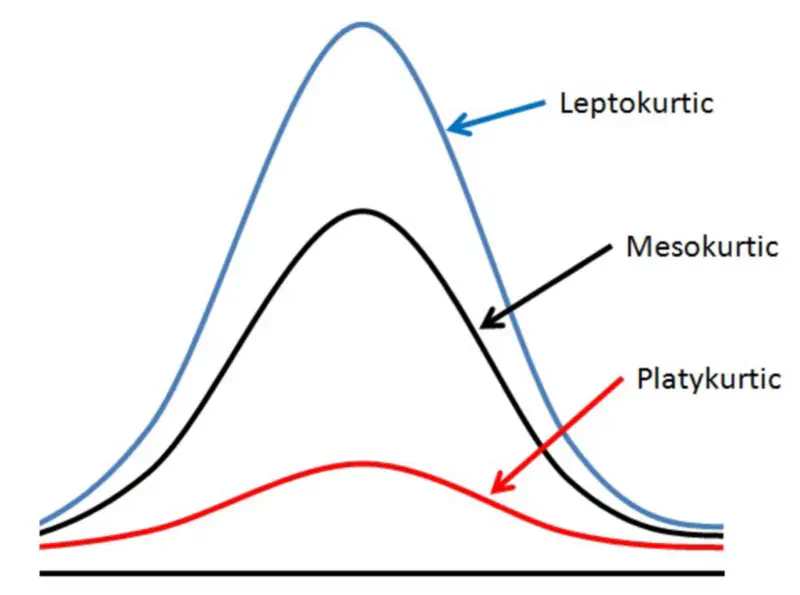
E.g: Percentile, Quartile, Inter Quartile Range(IQR), etc.
It indicates the percentage of scores in a dataset that are equal to or below a specific value.
Here, the complete dataset is divided into 100 equal parts.
- \(k^{th}\) percentile => at least \(k\) percent of the data points are equal to or below the value.
- It is a relative comparison, i.e, compares a score with the entire group’s performance.
- Quartiles are basis for box plots.
- 90th percentile => score is higher than 90% of of all other test takers.
They are special percentiles that divide the complete dataset into 4 equal parts.
Q1 => 25th percentile, value below which 25% of the data falls.
Q2 => 50th percentile, value below which 50% of the data falls; median.
Q3 => 75th percentile, value below which 75% of the data falls.
- Data = \(\{1,2,3,4,5,6,7,8,9,10,100\}\) \[ Q1 = (11+1) * 1/4 = 12*1/4 = 3 \\ Q2 = (11+1) * 1/2 = 12*1/2 = 6 \\ Q3 = (11+1) * 3/4 = 12*3/4 = 9 \]
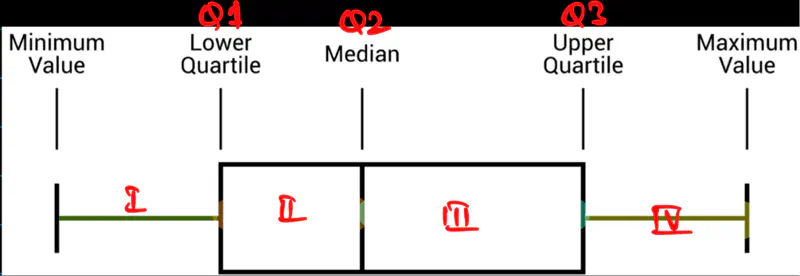
It is the single number that measures the spread of middle 50% of the data, i.e Q1-Q3.
- More robust measure of spread than range as is NOT impacted by outliers.
IQR = Q3 - Q1
- Data = \(\{1,2,3,4,5,6,7,8,9,10,100\}\) \[ Q1 = (11+1) * 1/4 = 12*1/4 = 3 \\ Q2 = (11+1) * 1/2 = 12*1/2 = 6 \\ Q3 = (11+1) * 3/4 = 12*3/4 = 9 \]
Therefore, IQR = Q3-Q1 = 9-3 = 6
IQR is a standard tool for detecting outliers.
Values that fall outside the ‘fences’ can be considered as potential outliers.
Lower fence = Q1 - 1.5 * IQR
Upper fence = Q3 + 1.5 * IQR
- Data = \(\{1,2,3,4,5,6,7,8,9,10,100\}\) \[ Q1 = (11+1) * 1/4 = 12*1/4 = 3 \\ Q2 = (11+1) * 1/2 = 12*1/2 = 6 \\ Q3 = (11+1) * 3/4 = 12*3/4 = 9 \]
IQR = Q3-Q1 = 9-3 = 6
Lower fence = Q1 - 1.5 * IQR = 3 - 9 = -6
Upper fence = Q3 + 1.5 * IQR = 9 + 9 = 18
So, any data point that is less than -6 or greater than 18 is considered as a potential outlier.
As in this example, 100 can be considered as an outlier.
Even though the above metrics give us a good idea of the data distribution,
but still we should always plot the data and visually inspect the data distribution.
As these metrics may not provide the complete picture.
A mathematician called Francis John Anscombe has illustrated this point beautifully in his Anscombe’s Quartet.
Anscombe’s quartet:
It comprises four datasets that have nearly identical simple descriptive statistics,
yet have very different distributions and appear very different when plotted.
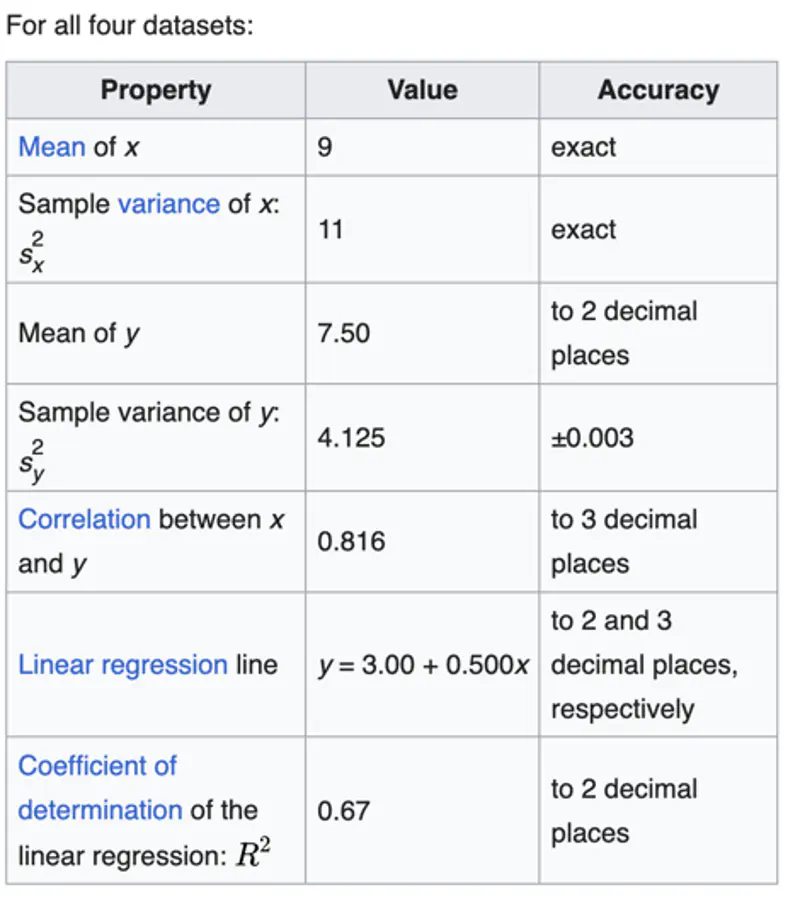
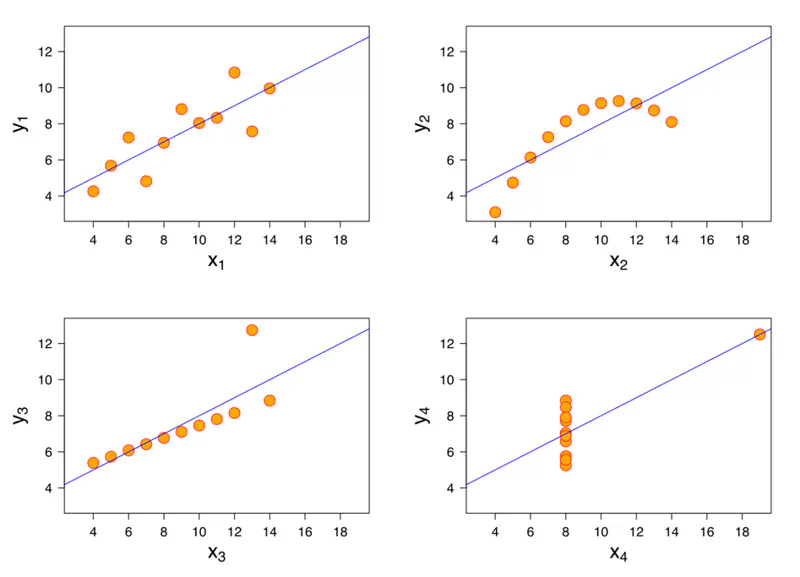
End of Section
2.2.2 - Correlation
It measures the direction of linear relationship between two variables \(X\) and \(Y\).
\(N\) = size of population
\(\mu_{x}\) = population mean of \(X\)
\(\mu_{y}\) = population mean of \(Y\)
\(n\) = size of sample
\(\bar{x}\) = sample mean of \(X\)
\(\bar{y}\) = sample mean of \(Y\)
Note: We have a term (n-1) instead of n in the denominator to make it an unbiased estimate, called Bessel’s Correction.
If both \((x_i - \bar{x})\) and \((y_i - \bar{y})\) have the same sign, then the product is positive(+ve).
If both \((x_i - \bar{x})\) and \((y_i - \bar{y})\) have opposite signs, then the product is negative(-ve).
The final value of covariance depends on the sum of the above individual products.
\( \begin{aligned} \text{Cov}(X, Y) &> 0 &&\Rightarrow \text{ } X \text{ and } Y \text{ increase or decrease together} \\ \text{Cov}(X, Y) &= 0 &&\Rightarrow \text{ } \text{No linear relationship} \\ \text{Cov}(X, Y) &< 0 &&\Rightarrow \text{ } \text{If } X \text{ increases, } Y \text{ decreases (and vice versa)} \end{aligned} \)
Limitation:
Covariance is scale-dependent, i.e, units of X and Y impact its magnitude.
This makes it hard to make comparisons of covariance across different datasets.
E.g: Covariance between age and height will NOT be same as the covariance between years of experience and salary.
Note:It only measures the direction of the relationship, but does NOT give any information about the strength of the relationship.
- \(X = [1, 2, 3] \) and \(Y = [2, 4, 6] \)
Let’s calculate the covariance:
\(\text{Cov}(X, Y) = \frac{1}{n-1}\sum_{i=1}^{n}(x_i - \bar{x})(y_i - \bar{y})\)
\(\bar{x} = 2\) and \(\bar{y} = 4\)
\(\text{Cov}(X, Y) = \frac{1}{3-1}[(1-2)(2-4) + (2-2)(4-4) + (3-2)(6-4)]= 0\)
\( = \frac{1}{2}[2+0+2]= 2\)
=> Cov(X,Y) > 0 i.e if X increases, Y increases and vice versa.
It measures both the strength and direction of the linear relationship between two variables \(X\) and \(Y\).
It is a standardized version of covariance that gives a dimensionless measure of linear relationship.
There are 2 popular ways to calculate correlation coefficient:
- Pearson Correlation Coefficient (r)
- Spearman Rank Correlation Coefficient (\(\rho\))
It is a standardized version of covariance and most widely used measure of correlation.
Assumption: Data is normally distributed.
\(\sigma_{x}\) and \(\sigma_{y}\) are the standard deviations of \(X\) and \(Y\).
Range of \(r\) is between -1 and 1.
\(r = 1\) => perfect +ve linear relationship between X and Y
\(r = -1\) => perfect -ve linear relationship between X and Y
\(r = 0\) => NO linear relationship between X and Y.
Note: A correlation coefficient of 0.9 means that there is a strong linear relationship between X and Y,
irrespective of their units.
- \(X = [1, 2, 3] \) and \(Y = [2, 4, 6] \)
Let’s calculate the covariance:
\(\text{Cov}(X, Y) = \frac{1}{n-1}\sum_{i=1}^{n}(x_i - \bar{x})(y_i - \bar{y})\)
\(\bar{x} = 2\) and \(\bar{y} = 4\)
\(\text{Cov}(X, Y) = \frac{1}{3-1}[(1-2)(2-4) + (2-2)(4-4) + (3-2)(6-4)]= 0\)
\( => \text{Cov}(X, Y) = \frac{1}{2}[2+0+2]= 2\)
Let’s calculate the standard deviation of \(X\) and \(Y\):
\(\sigma_{x} = \sqrt{\frac{1}{n-1}\sum_{i=1}^{n}(x_i - \bar{x})^2} \)
\(= \sqrt{\frac{1}{3-1}[(1-2)^2 + (2-2)^2 + (3-2)^2]}\)
\(= \sqrt{\frac{1+0+1}{2}} =\sqrt{\frac{2}{2}} = 1 \)
Similarly, we can calculate the standard deviation of \(Y\):
\(\sigma_{y} = \sqrt{\frac{1}{n-1}\sum_{i=1}^{n}(y_i - \bar{y})^2} \)
\(= \sqrt{\frac{1}{3-1}[(2-4)^2 + (4-4)^2 + (6-4)^2]}\)
\(= \sqrt{\frac{4+0+4}{2}} =\sqrt{\frac{8}{2}} = 2 \)
Now, we can calulate the pearson correlation coefficient (r):
\(r_{xy} = \frac{Cov(X, Y)}{\sigma_{x} \sigma_{y}}\)
=> \(r_{xy} = \frac{2}{1* 2}\)
=> \(r_{xy} = 1\)
Therefore, we can say that there is a strong +ve linear relationship between X and Y.
It is a measure of the strength and direction of the monotonic relationship between two ranked variables \(X\) and \(Y\).
It captures monotonic relationship, meaning the variables move in the same or opposite direction,
but not necessarily a linear relationship.
- It is used when Pearson’s correlation is not suitable, such as, ordinal data, or when the continuous data does not meet the assumptions of linear methods, such as, Pearson’s correlation.
- Non-parametric measure of correlation that uses ranks instead of raw data.
- Quantifies how well the ranks of one variable predict the ranks of the other variable.
- Range of \(\rho\) is between -1 and 1.
- Compute the correlation of ranks awarded to a group of 5 students by 2 different teacherrs.
Student Teacher A Rank Teacher B Rank \(d_i\) \(d_i^2\) S1 1 2 -1 1 S2 2 1 1 1 S3 3 3 0 0 S4 4 5 -1 1 S5 5 4 1 1
\(\sum_{i}d_i^2 = 4 \)
\( n = 5 \)
\(\rho_{xy} = 1 - \frac{6\sum_{i}d_i^2}{n(n^2-1)}\)
=> \(\rho_{xy} = 1 - \frac{6*4}{5(5^2-1)}\)
=> \(\rho_{xy} = 1 - \frac{24}{5*24}\)
=> \(\rho_{xy} = 1 - \frac{1}{5}\)
=> \(\rho_{xy} = 0.8\)
Therefore, we can say that there is a strong +ve correlation between the ranks given by teacher A and teacher B.
- \(X = [1, 2, 3] \) and \(Y = [1, 8, 27] \)
Here, Spearman’s rank correlation coefficient \(\rho\) will be perfect 1 as there is a monotonic relationship i.e as X increases, Y increases and vice versa.
But, the Pearson’s correlation coefficient (r) will be slightly less than 1 i.e r = 0.9662.
Correlation is very useful in feature selection for training machine learning models.
- If 2 features are highly correlated => they provide redundant information.
- One of the features can be removed without significant loss of information.
- Keeping both can cause issues, such as, multicollinearity.
- If a feature is highly correlated with the target variable => this feature is a strong predictor, so keep it.
- A feature with very low or near zero correlation with the target variable may be considered for removal, as they have little predictive power.
Causation means that one variable directly causes the change in another variable, i.e, direct
cause->effect relationship.
Whereas, correlation means that two variables move together.
- Correlation does NOT imply Causation.
- Correlation simply shows an association between two variables that could be coincidental or due to some third, unobserved, factor.
E.g: Election results and stock market - there may be some correlation between the two,
but establishing clear causal links is difficult.
End of Section
2.2.3 - Central Limit Theorem
Before we understand the Central Limit Theorem, let’s understand a few related concepts.
It is the true average of the entire group.
It describe the central tendency of the entire population.
\( \mu = \frac{1}{N}\sum_{i=1}^{N}x_i \)
N: Number of data points
It is the average of a smaller representative subset (a sample) of the entire population.
It provides an estimate of the population mean.
\( \bar{x} = \frac{1}{n}\sum_{i=1}^{n}x_i \)
n: size of sample
the sample mean converges to the true the population mean.
In other words, a long-run average of a repeated random variable approaches the expected value.
This law states that for a sequence of of I.I.D random variables \( X_1, X_2, \dots, X_n \),
with finite mean and variance, the distribution of the sample mean \( \bar{X} \) approaches a normal distribution
as \( n \rightarrow \infty \), regardless of its original population distribution.
The distribution of the sample mean is : \( \bar{X} \sim N(\mu, \sigma^2/n)\)
Let, \( X_1, X_2, \dots, X_n \) are I.I.D random variables.
- Population mean = \(E[X_i] = \mu < \infty\)
- Population Variance = \(Var[X_i] = \sigma^2 < \infty \)
- Sample mean = \( \bar{X_n} = \frac{1}{n}\sum_{i=1}^{n}X_i = \frac{1}{n}(X_1 + X_2+ \dots +X_n) \)
- Variance of sample means = \( Var[\bar{X_n}] = Var[\frac{1}{n}(X_1+ X_2+ \dots+ X_n)]\)
Now, let’s calculate the variance of sample means.
We know that:
- \(Var[X+Y] = Var[X] + Var[Y] \), for independent random variables X and Y.
- \(Var[cX] = c^2Var[X] \), for constant ‘c’.
Let’s apply above 2 rules on the variance of sample means equation above:
\[ \begin{aligned} Var[\bar{X_n}] &= Var[\frac{1}{n}(X_1+ X_2+ \dots+ X_n)] \\ &= \frac{1}{n^2}[Var[X_1+ X_2+ \dots+ X_n]] \\ &= \frac{1}{n^2}[Var[X_1] + Var[X_2] + \dots + Var[X_n]] \\ \text{We know that: } Var[X_i] = \sigma^2 \\ &= \frac{1}{n^2}[\sigma^2 + \sigma^2 + \dots + \sigma^2] \\ &= \frac{n\sigma^2}{n^2} \\ => Var[\bar{X_n}] &= \frac{\sigma^2}{n} \end{aligned} \]Since, standard deviation = \(\sigma = \sqrt{Variance}\)
Therefore, Standard Deviation\([\bar{X_n}] = \sqrt{\frac{\sigma^2}{n}} = \frac{\sigma}{\sqrt{n}}\)
The standard deviation of the sample means is also known as “Standard Error”.
Note: We can also standardize the sample mean, i.e, mean centering and variance scaling.
Standardisation helps us to use the Z-tables of normal distribution.
We know that, a standardized random variable \(Y_i = \frac{X_i - \mu}{\sigma}\)
Similarly, standardized sample mean:
Note: For practical purposes, \(n \ge 30\) is considered as a sufficient sample size for the CLT to hold.
- Let’s collect the data for height of people in a city to find the average height of people in the city.
- Sample size (n) = 100
- And then repeat this data collection process 1000 times.
- For each of these 1000 (k) samples, calculate the sample mean \(X_1, X_2, \dots, X_{1000(k)} \)
- Now, when we plot these 1000(k) sample means, the resulting distribution will be very close to a normal/Gaussian distribution.
\(\bar{X_n} \sim N(\mu, \sigma^2/n)\), for large n, typically \(n \ge 30\).
Note:
- ‘k’ = a large number of repetitions allows us to observe the distribution of sample means after plotting.
- ’n’ = number of samples in each repetition is fixed for any given calculation of sample mean \(\bar{X_n}\).
The variance must be finite, else, the sample mean will NOT converge to a normal distribution.
If a distribution has a heavy tail, then the expected value calculation diverges.
e.g:
- Cauchy distribution has infinite mean and infinite variance.
- Pareto distribution (with low alpha) has infinite variance, such as distribution of wealth.
End of Section
2.2.4 - Confidence Interval
It is a range of values that is likely to contain the true population mean, based on a sample.
Instead of giving a point estimate, it gives a range of values with confidence level.
For normal distribution, confidence interval :
\(\bar{X}\): Sample mean
\(Z\): Z-score corresponding to confidence level
\(n\): Sample size
\( \sigma \): Population Standard Deviation
Applications:
- A/B testing, i.e., compare 2 or more versions of a product.
- ML model performance evaluation, i.e, instead of giving a single performance score of say 85%,
it is better to provide a 95% confidence interval, such as, [82.5%, 87.8%].
95% confidence interval does NOT mean there is a 95% chance that the true mean lies in the specific calculated interval.
- It just means that if we repeat the sampling process many times, then 95% of of those calculated intervals will capture or contain the true population mean \(\mu\).
- Also, we cannot say there is 95% probability that the true mean is within that specific range because true population mean is a fixed constant, NOT a random variable.
Let’s suppose we want to measure the average weight of a certain species of dog.
We want to estimate the true population
mean \(\mu\) using confidence interval.
Note: True average weight = 30 kg, but this is NOT known to us.
| Sample Number | Sample Mean | 95% Confidence Interval | Did it capture \(\mu\) ? |
|---|---|---|---|
| 1 | 29.8 kg | (28.5, 31.1) | Yes |
| 2 | 30.4 kg | (29.1, 31.7) | Yes |
| 3 | 31.5 kg | (30.2, 32.8) | No |
| 4 | 28.1 kg | (26.7, 29.3) | No |
| - | - | - | - |
| - | - | - | - |
| - | - | - | - |
| 100 | 29.9 kg | (28.6, 31.2) | Yes |
- We generated 100 confidence intervals(CI) each based on different samples.
- 95% CI guarantees that, in long run, 95 out of 100 CIs will include the true average weight, i.e, \(\mu=30kg\), and may be will miss 5 out of 100 times.
Suggest which company is offering a better salary?
Below is the details of the salaries based on a survey of 50 employees.
| Company | Average Salary(INR) | Standard Deviation |
|---|---|---|
| A | 36 lpa | 7 lpa |
| B | 40 lpa | 14 lpa |
For comparison, let’s calculate the 95% confidence interval for the average salaries of both companies A and B.
We know that:
\( CI = \bar{X} \pm Z\frac{\sigma}{\sqrt{n}} \)
Margin of Error(MoE) \( = Z\frac{\sigma}{\sqrt{n}} \)
Z-Score for 95% CI = 1.96
\(MoE_A = 1.96*\frac{7}{\sqrt{50}} \approx 1.94 \)
=> 95% CI for A = \(36 \pm 1.94 \) = [34.06, 37.94]
\(MoE_B = 1.96*\frac{14}{\sqrt{50}} \approx 3.88\)
=> 95% CI for B = \(40 \pm 3.88 \) = [36.12, 43.88]
We can see that initially company B’s salary looked obviously better,
but after calculating the 95% CI, we can see that there is a significant overlap in the salaries of two companies,
i.e [36.12, 37.94].
End of Section
2.2.5 - Hypothesis Testing
Hypothesis Testing is used to determine whether a claim or theory about a population is supported by a sample data,
by assessing whether observed difference or patterns are likely due to chance or represent a true effect.
- It allows companies to test marketing campaigns or new strategies on a small scale before committing to larger investments.
- Based on the results of hypothesis testing, we can make reliable inferences about the whole group based on a representative sample.
- It helps us determine whether an observed result is statistically significant finding, or if it could have just happened by random chance.
It is a statistical inference framework used to make decisions about a population parameter, such as, the mean, variance,
distribution, correlation, etc., based on a sample of data.
It provides a formal method to evaluate competing claims.
Null Hypothesis (\(H_0\)):
Status quo or no-effect or no difference statement; almost always contains a statement of equality.
Alternative Hypothesis (\(H_1 ~or~ H_a\)):
The statement representing an effect, a difference, or a relationship.
It must be true if the null hypothesis is rejected.
- Test of Means:
- 1-Sample Mean Test: Compare sample mean to a known population mean.
- 2-Sample Mean Test: Compare means of 2 populations.
- Paired Mean Test: Compare means when data is paired, e.g., before vs. after test.
- Test of Median:
- Mood’s Median Test
- Sign Test
- Wilcoxon Signed Rank Test (non-parametric)
- Test of Variance:
- Chi-Square Test for a single variance
- F-Test to compare variances of 2 populations
- Test of Distribution(Goodness of Fit):
- Kolmogorov-Smirnov Test
- Shapiro-Wilk Test
- Anderson-Darling Test
- Chi-Square Goodness of Fit Test
- Test of Correlation:
- Pearson’s Correlation Coefficient Test
- Spearman’s Rank Correlation Test
- Kendall’s Tau Correlation Test
- Chi-Square Test of Independence
- Regression Test:
- T-Test: For regression coefficients
- F-Test: For overall regression significance
Step 1: Define the null and alternative hypotheses.
Step 2: Select a relevant statistical test for the task with associated test statistic.
Step 3: Calculate the test statistic under null hypothesis.
Step 4: Select a significance level (\(\alpha\)), i.e, the maximum acceptable false positive rate;
typically - 5% or 1%.
Step 5: Compute the p-value from the observed value of test-statistic.
Step 6: Make a decision to either accept or reject the null hypothesis, based on the significance level (\(\alpha\)).
Say, the data, D: <patient_id, med_1/med_2, recovery_time(in days)>
We need some metric to compare the recovery times of 2 medicines.
We can use the mean recovery time as the metric, because we know that we can use following techniques for comparison:
- 2-Sample T-Test; if \(n < 30\) and population standard deviation \(\sigma\) is unknown.
- 2-Sample Z-Test; if \(n \ge 30\) and population standard deviation \(\sigma\) is known.
Note: Let’s assume the sample size \(n < 30\), because medical tests usually have small sample sizes.
=> We will use the 2-Sample T-Test; we will continue using T-Test throughout the discussion.
Step 1: Define the null and alternative hypotheses.
Null Hypothesis \(H_0\): The mean recovery time of 2 medicines is the same i.e \(Mean_{m1} = Mean_{m2}\) or \(m_{m1} = m_{m2}\).
Alternate Hypothesis \(H_a\): \(m_{m1} < m_{m2}\) (1-Sided T-Test) or \(m_{m1} ⍯ m_{m2}\) (2-Sided T-Test).
Step 2: Select a relevant statistical test for the task with associated test statistic.
Let’s do a 2 sample T-Test, i.e, \(m_{m1} < m_{m2}\)
Step 3: Calculate the test statistic under null hypothesis.
Test Statistic:
For 2 sample T-Test:
s: Standard Deviation
n: Sample Size
Note: If the 2 means are very close then \(t_{obs} \approx 0\).
Step 4: Suppose significance level (\(\alpha\)) = 5% or 0.05.
Step 5: Compute the p-value from the observed value of test-statistic.
P-Value:
p-value = area under curve = probability of observing test statistic \( \ge t_{obs} \) if the null hypothesis is true.
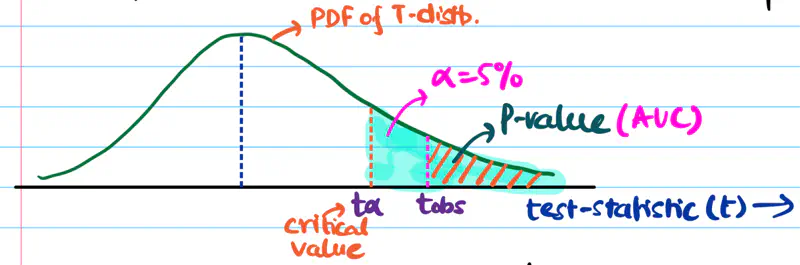
Step 6: Accept or reject the null hypothesis, based on the significance level (\(\alpha\)).
If \(p_{value} < \alpha\), we reject the null hypothesis and accept the alternative hypothesis and vice versa.
Note: In the above example \(p_{value} < \alpha\), so we reject the null hypothesis.
We need to do a left or right sided test, or a 2-sided test, this depends upon our alternate hypothesis and test statistic.
Let’s continue our 2 sample mean T-test to understand the concept:
Left Sided/Tailed Test:
\(H_a\): Mean recovery time of medicine 1 < medicine 2, i.e, \(m_{m_1} < m_{m_2}\)
=> \(m_{m_1} - m_{m_2} < 0\)
Since, the denominator in above equation is always positive.
=> \(t_{obs} < 0\)
Therefore, we need to do a left sided/tailed test.
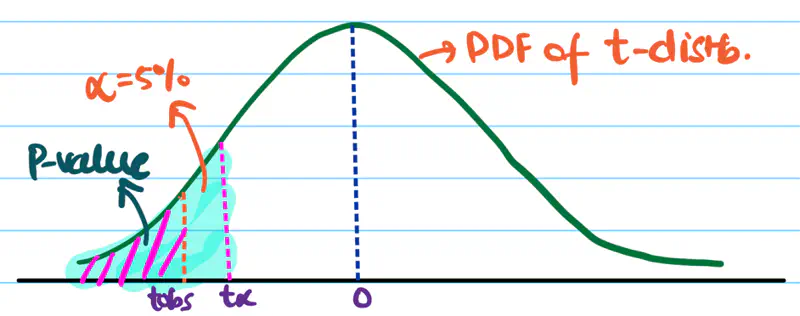
So, we want \(t_{obs}\) to be very negative to confidently conclude that alternate hypothesis is true.
Right Sided/Tailed Test:
\(H_a\): Mean recovery time of medicine 1 > medicine 2, i.e, \(m_{m_1} > m_{m_2}\)
=> \(m_{m_1} - m_{m_2} > 0\)
Similarly, here we need to do a right sided/tailed test.
2 Sided/Tailed Test:
\(H_a\): Mean recovery time of medicine 1 ⍯ medicine 2, i.e, \(m_{m_1} ⍯ ~ m_{m_2}\)
=> \(m_{m_1} - m_{m_2} < 0\) or \(m_{m_1} - m_{m_2} > 0\)
If \(H_a\) is true then \(t_{obs}\) is a large -ve value or a large +ve value.
Since, t-distribution is symmetric, we can divide the significance level \(\alpha\) into 2 equal parts.
i.e \(\alpha = 2.5\%\) on each side.
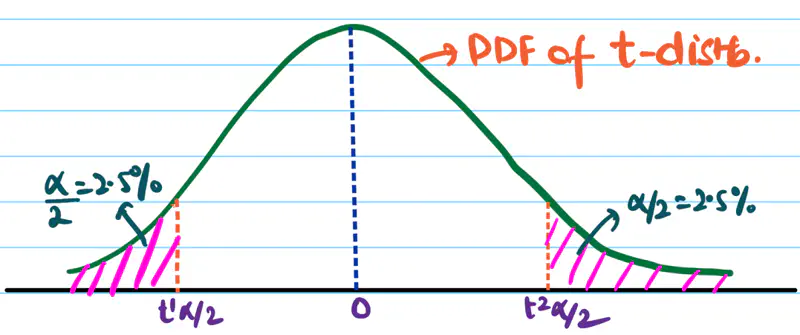
So, we want \(t_{obs}\) to be very negative or very positive to confidently conclude that the alternate hypothesis is true. We accept \(H_a\) if \(t_{obs} < t^1_{\alpha/2}\) or \(t_{obs} > t^2_{\alpha/2}\).
Note: For critical applications ‘\(\alpha\)’ can be very small i.e. 0.1% or 0.01%, e.g medicine.
It is the probability of wrongly rejecting a true null hypothesis, known as a Type I error or false +ve rate.
- Tolerance level of wrongly accepting alternate hypothesis.
- If the p-value < \(\alpha\), we reject the null hypothesis and conclude that the finding is statistically NOT so significant..
It is a specific point on the test-statistic distribution that defines the boundaries of the null hypothesis
acceptance/rejection region.
It tells us that at what value (\(t_{\alpha}\)) of test statistic will the area under curve be equal to the significance level \(\alpha\).
For a right tailed/sided test:
- if \(t_{obs} > t_{\alpha} => p_{value} < \alpha\); therefore, reject null hypothesis.
- if \(t_{obs} < t_{\alpha} => p_{value} \ge \alpha\); therefore, failed to reject null hypothesis.
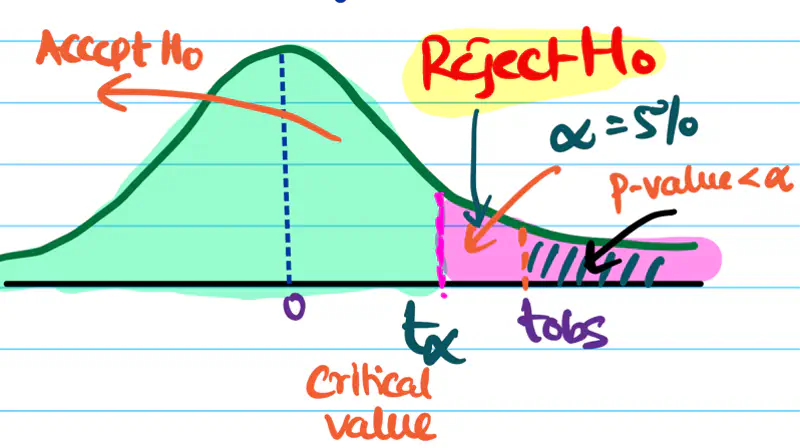
It is the probability that a hypothesis test will correctly reject a false null hypothesis (\(H_{0}\)) when the alternative hypothesis (\(H_{a}\)) is true.
Power of test = \(1 - \beta\)
Probability of correctly accepting alternate hypothesis (\(H_{a}\))
\(\alpha\): Probability of wrongly accepting alternate hypothesis \(H_{a}\)
\(\beta\): Probability of wrongly rejecting alternate hypothesis \(H_{a}\)
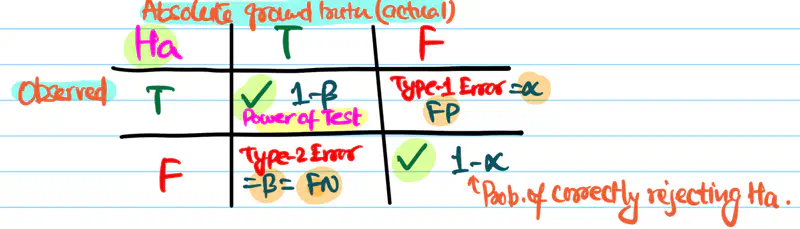

Yes, having a large sample size makes a hypothesis test more powerful.
- As n increases, sample mean \(\bar{x}\) approaches the population mean \(\mu\).
- Also, as n increases, t-distribution approaches normal distribution.
It is a standardized objective measure that complements p-value by clarifying whether a statistically significant
finding has any real world relevance.
It quantifies the magnitude of relationship between two variables.
- Larger effect size => more impactful effect.
- Standardized (mean centered + variance scaled) measure allows us to compare the imporance of effect across various studies or groups, even with different sample sizes.
Effect size is measured using Cohen’s d formula:
\(\bar{X}\): Sample mean
\(s_p\): Pooled Standard deviation
\(n\): Sample size
\(s\): Standard deviation
Note: Theoretically, Cohen’s d value can range from negative infinity to positive infinity.
but for practical purposes, we use the following value:
small effect (\(d=0.2\)), medium effect (\(d=0.5\)), and large effect (\(d\ge 0.8\)).
More overlap => less effect i.e low Cohen’s d value.
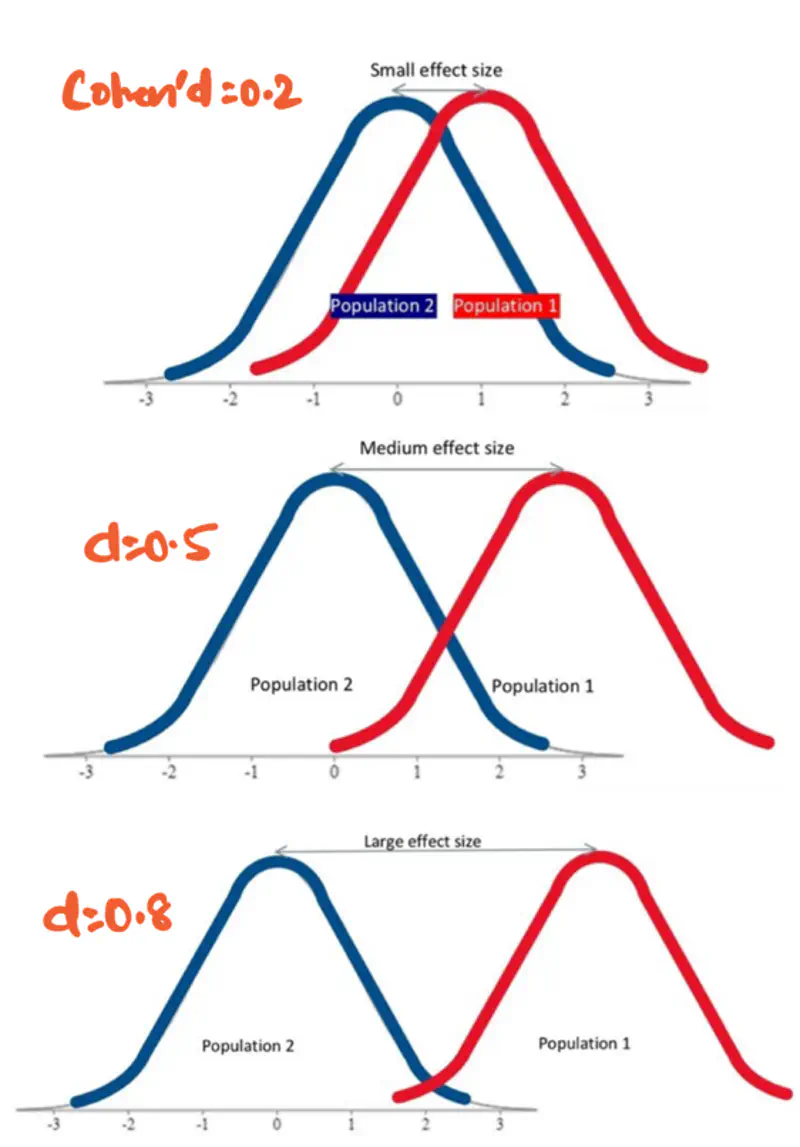
- A study on drug trials finds that patients taking a new drug had statistically significant
improvement (p-value<0.05), compared to a placebo group.
- Small effect size: Cohen’s d = 0.1 => drug had minimal effect.
- Large effect size: Cohen’s d = 0.8 => drug produced substantial improvement.
End of Section
2.2.6 - T-Test
It is a statistical test that is used to determine whether the sample mean is equal to a hypothesized value or
is there a significant difference between the sample means of 2 groups.
- It is a parametric test, since it assumes data to be approximately normally distributed.
- Appropriate when:
- sample size n < 30.
- population standard deviation \(\sigma\) is unknown.
- It is based on Student’s t-distribution.
It is a continuous probability distribution that is a symmetrical, bell-shaped curve similar
to the normal distribution but with heavier tails.
Shape of the curve or mass in tail is controlled by degrees of freedom.
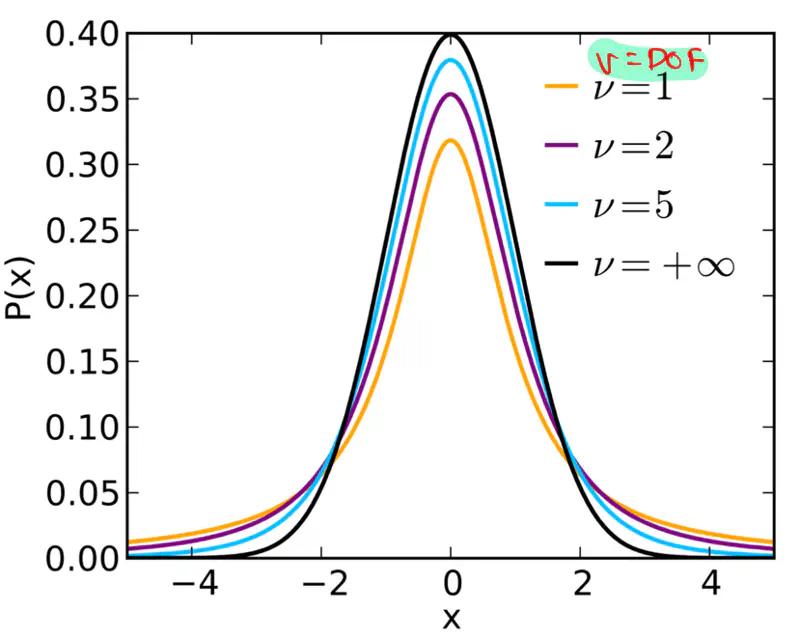
There are 3 types of T-Test:
- 1-Sample T-Test: Test if sample mean differs from hypothesized value.
- 2-Sample T-Test: Test whether there is a significant difference between the means of two independent groups.
- Paired T-Test: Test whether 2 related samples differ, e.g., before and after.
Generally speaking, it represents the number of independent values that are free to vary in a dataset when estimating a parameter.
e.g.: If we have k observations and their sum = 50.
The sum of (k-1) terms can be anything, but the kth term is fixed at 50 - (sum of other (k-1) terms).
So, we have only (k-1) terms that can change independently, therefore, the DOF(\(\nu\)) = k-1.
It is used to test whether the sample mean is equal to a known/hypothesized value.
Test statistic (t):
where,
\(\bar{x}\): sample mean
\(\mu\): hypothesized value
\(s\): sample standard deviation
\(n\): sample size
\(\nu = n-1 \): degrees of freedom
Tester ran the test 20 times and found the average API repsonse time to be 115 ms, with a standard deviation of 25 ms.
Is the developer’s claim valid?
Let’s verify developer’s claim using the tester’s test results using 1 sample t-test.
Null hypothesis: \(H_0\) = The average API response time is 100 ms, i.e, \(\bar{x} = \mu\).
Alternative hypothesis: \(H_a\) = The average API response time > 100 ms, i.e, \(\bar{x} > \mu\) => right tailed test.
Hypothesized mean \(\mu\) = 100 ms
Sample mean \(\bar{x}\) = 115 ms
Sample standard deviation \(s\) = 25 ms
Sample size \(n\) = 20
Degrees of freedom \(\nu\) = 19
\( t_{obs} = \frac{\bar{x} - \mu}{s/\sqrt{n}}\) = \(\frac{115 - 100}{25/\sqrt{20}}\)
= \(\frac{15\sqrt{20}}{25} = \frac{3\sqrt{20}}{5} \approx 2.68\)
Let significance level \(\alpha\) = 5% =0.05.
Critical value \(t_{0.05}\) = 1.729
Important: Find the value of \(t_{\alpha}\) in T-table
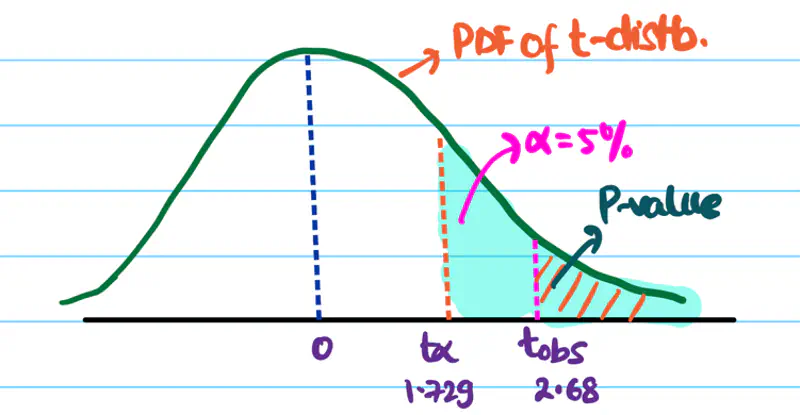
Since \(t_{obs}\) > \(t_{0.05}\), we reject the null hypothesis.
And, accept the alternative hypothesis that the API response time is significantly > 100 ms.
Hence, the developer’s claim is NOT valid.
It is used to determine whether there is a significant difference between the means of two independent groups.
There are 2 types of 2-sample t-test:
- Unequal Variance
- Equal Variance
Unequal Variance:
In this case, the variance of 2 independent groups is not equal.
Also called, Welch’s t-test.
Test statistic (t):
Equal Variance:
In this case, both samples come from equal or approximately equal variance.
Test statistic (t):
Here, degrees of freedom (for equal variance) \(\nu\) = \(n_1 + n_2 - 2\).
\(\bar{x}\): sample mean
\(s\): sample standard deviation
\(n\): sample size
\(\nu\): degrees of freedom
The AI team wants to validate whether the new ML model accuracy is better than the existing model’s accuracy.
Below is the data for the existing model and the new model.
| New Model (A) | Existing Model (B) | |
|---|---|---|
| Sample size (n) | 24 | 18 |
| Sample mean (\(\bar{x}\)) | 91% | 88% |
| Sample std. dev. (s) | 4% | 3% |
Given that the variance of accuracy scores of new and existing models are almost same.
Now, let’s follow our hypothesis testing framework.
Null hypothesis: \(H_0\): The accuracy of new model is same as the accuracy of existing model.
Alternative hypothesis: \(H_a\): The new model’s accuracy is better/greater than the existing model’s accuracy => right tailed test
Let’s solve this using 2 sample T-Test, since the sample size n < 30.
Since the variance of 2 sample are almost equal then we can use the pooled variance method.
Next let’s compute the test statistic, under null hypothesis.
DOF \(\nu\) = \(24+18-2\) = 42 - 2 = 40
Let significance level \(\alpha\) = 5% =0.05.
Critical value \(t_{0.05}\) = 1.684
Important: Find the value of \(t_{\alpha}\) in T-table
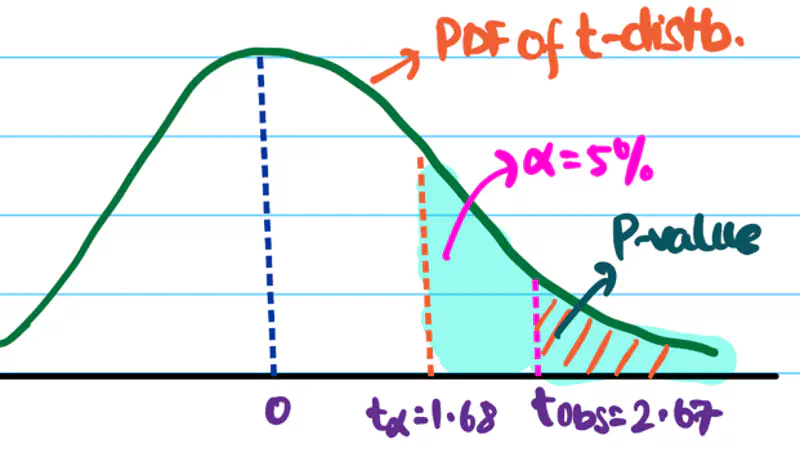
Since \(t_{obs}\) > \(t_{0.05}\), we reject the null hypothesis.
And, accept the alternative hypothesis that the new model has better accuracy than the existing model.
End of Section
2.2.7 - Z-Test
It is a statistical test used to determine whether there is a significant difference between mean of 2 groups or sample and population mean.
- It is a parametric test, since it assumes data to be normally distributed.
- Appropriate when:
- sample size \(n \ge 30\).
- population standard deviation \(\sigma \) is known.
- It is based on Gaussian/normal distribution.
- It compares the difference between means relative to standard error, i.e, standard deviation of sampling distribution of sample mean.
There are 2 types of Z-Test:
- 1-Sample Z-Test: Used to compare the mean of a sample mean with a population mean.
- 2-Sample Z-Test: Used to compare the sample means of 2 independent samples.
It is a standardized score that measures how many standard deviations a particular data point is away from the population mean \(\mu\).
- Transform a normal distribution \(\mathcal{N}(\mu, \sigma^2)\) to a standard normal distribution \(Z \sim \mathcal{N}(0, 1)\).
- Standardized score helps us compare values from different normal distributions.
Z-score is calculated as:
\[Z = \frac{x - \mu}{\sigma}\]x: data point
\(\mu\): population mean
\(\sigma\): population standard deviation
e.g:
- Z-score of 1.5 => data point is 1.5 standard deviations above the mean.
- Z-score of -2.0 => data point is 2.0 standard deviations below the mean.
Z-score helps to define probability areas:
- 68% of the data points fall within \(\pm 1 \sigma\).
- 95% of the data points fall within \(\pm 2 \sigma\).
- 99.7% of the data points fall within \(\pm 3 \sigma\).
Note:
- Z-Test applies the concept of Z-score to sample mean rather than a single data point.
It is used to test whether the sample mean \(\bar{x}\) is significantly different from a known population mean \(\mu\).
Test Statistic:
\(\bar{x}\): sample mean
\(\mu\): hypothesized population mean
\(\sigma\): population standard deviation
\(n\): sample size
\(\sigma / \sqrt{n}\): standard error of mean
Read more about Standard Error
Note: Test statistic Z follows a standard normal distribution \(Z \sim \mathcal{N}(0, 1)\).
It is used to test whether the sample means \(\bar{x_1}\) and \(\bar{x_2}\) of 2 independent samples are significantly different from each other.
Test Statistic:
\(\bar{x_1}\): sample mean of first sample
\(\bar{x_2}\): sample mean of second sample
\(\sigma_1\): population standard deviation of first sample
\(\sigma_2\): population standard deviation of second sample
\(n_1\): sample size of first sample
\(n_2\): sample size of second sample
Note: Test statistic Z follows a standard normal distribution \(Z \sim \mathcal{N}(0, 1)\).
After applying a new optimisation, and n=100 runs, yields a sample mean of 117 seconds.
Does the optimisation significantly reduce the runtime of the model?
Consider the significance level of 5%.
Null Hypothesis: \(\mu = 120\) seconds, i.e, no change.
Alternative Hypothesis: \(\mu < 120\) seconds => left tailed test.
We will use 1-sample Z-Test to test the hypothesis.
Test Statistic:
Since, significance level \(\alpha\) = 5% =0.05.
Critical value \(Z_{0.05}\) = -1.645
Important: Find the value of \(Z_{\alpha}\) in Z-Score Table
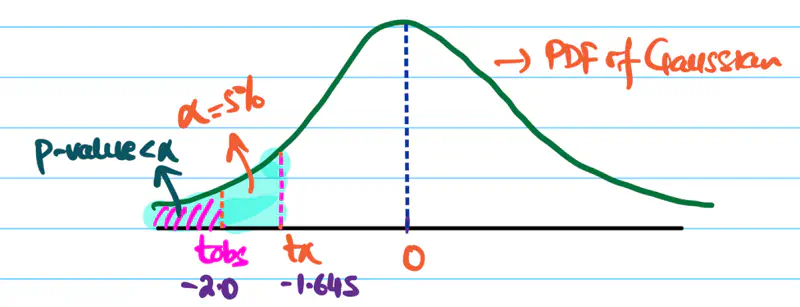
Our \(t_{obs}\) is much more extreme than the the critical value \(Z_{0.05}\), => p-value < 5%.
Hence, we reject the null hypothesis.
Therefore, there is a statistically significant evidence that the new optimisation reduces the runtime of the model.
It is a statistical hypothesis test used to determine if there is a significant difference between the proportion
of a characteristic in two independent samples or to compare a sample proportion to a known population value
- It is used when dealing with categorical data, such as, success/failure, male/female, yes/no etc.
It is of 2 types:
- 1-Sample Z-Test of Proportion: Used to test whether the observed proportion in a sample differs from hypothesized proportion.
- 2-Sample Z-Test of Proportion: Used to compare whether the 2 independent samples differ in their proportions.
The categorical data,i.e success/failure, is discrete that can be modeled as Bernoulli distribution.
Let’s understand how this Bernoulli random variable can be approximated as a Gaussian distribution for a very large sample size,
using Central Limit Theorem.
Read more about Central Limit Theorem
Note:We will not prove the complete thing, but we will understand the concept in enough depth for clarity.
\(Y \sim Bernoulli(p)\)
\(X \sim Binomial(n,p)\)
E[X] = mean = np
Var[X] = variance = np(1-p)
X = total number of successes
p = true probability of success
n = number of trials
Proportion of Success in sample = Sample Proportion = \(\hat{p} = \frac{X}{n}\)
e.g.: If n=100 people were surveyed, and 40 said yes, then \(\hat{p} = \frac{40}{100} = 0.4\)
By Central Limit Theorem, we can state that for very large ’n’ Binomial distribution’s mean and variance can be used as an
approximation for Gaussian/Normal distribution:
Since, \(\hat{p} = \frac{X}{n}\)
We can say that:
Mean = \(\mu_{\hat{p}} = p\) = True proportion of success in the entire population
Standard Error = \(SE_{\hat{p}} = \sqrt{Var[\frac{X}{n}]} = \sqrt{\frac{p(1-p)}{n}}\) = Standard Deviation of the sample proportion
Note: Large Sample Condition - Approximation is only valid when the expected number of successes and failures are both > 10 (sometimes 5).
\(np \ge 10 ~and~ n(1-p) \ge 10\)
It is used to test whether the observed proportion in a sample differs from hypothesized proportion.
\(\hat{p} = \frac{X}{n}\): Proportion of success observed in a sample
\(p_0\): Specific propotion value under the null hypothesis
\(SE_0\): Standard error of sample proportion under the null hypothesis
Z-Statistic: Measures how many standard errors is the observed sample proportion \(\hat{p}\) away from \(p_0\)
Test Statistic:
It is used to compare whether the 2 independent samples differ in their proportions.
- Standard test used in A/B testing.
A company wants to compare its 2 different website designs A & B.
Below is the table that shows the data:
| Design | # of visitors(n) | # of signups(x) | conversion rate(\(\hat{p} = \frac{x}{n}\)) |
|---|---|---|---|
| A | 1000 | 80 | 0.08 |
| B | 1200 | 114 | 0.095 |
Is the design B better, i.e, design B increases conversion rate or proportion of visitors who sign up?
Consider the significance level of 5%.
Null Hypothesis: \(\hat{p_A} = \hat{p_B}\), i.e, no difference in conversion rates of 2 designs A & B.
Alternative Hypothesis: \(\hat{p_B} > \hat{p_A}\) i.e conversion rate of B > A => right tailed test.
Check large sample condition for both samples A & B.
\(n\hat{p_A} = 80 > 10 ~and~ n(1-\hat{p_A}) = 920 > 10\)
Similarly, we can show for B too.
Pooled proportion:
\[ \bar{p} = \frac{x_A+x_B}{n_A+n_B} = \frac{80+114}{1000+1200} = \frac{194}{2200} \\[10pt] => \bar{p}\approx 0.0882 \]Standard Error(Pooled):
\[ SE=\sqrt{\bar{p}(1-\bar{p})(\frac{1}{n_1} +\frac{1}{n_2})} \\[10pt] = \sqrt{0.0882(1-0.0882)(\frac{1}{1000} +\frac{1}{1200})} \\[10pt] => SE \approx 0.0123 \]Test Statistic(Z):
\[ t_{obs} = \frac{\hat{p_B}-\hat{p_A}}{SE_{\hat{p_A}-\hat{p_B}}} \\[10pt] = \frac{0.095-0.0882}{0.0123} \\[10pt] => t_{obs} \approx 1.22 \]Significance level \(\alpha\) = 5% =0.05.
Critical value \(Z_{0.05}\) = 1.645
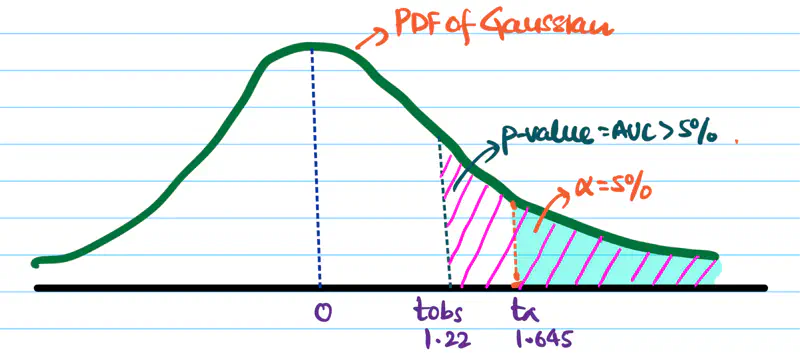
Since, \(t_{obs} < Z_{0.05}\) => p-value > 5%.
Hence, we fail to reject the null hypothesis.
Therefore, the observed conversion rate of design B is due to random chance; thus, B is not a better design.
End of Section
2.2.8 - Chi-Square Test
A random variable Q is said to follow a chi-square distribution with ’n’ degrees of freedom,i.e \(\chi^2(n)\),
if it is the sum of squares of ’n’ independent random variables that follow a standard normal distribution, i.e, \(N(0,1)\).
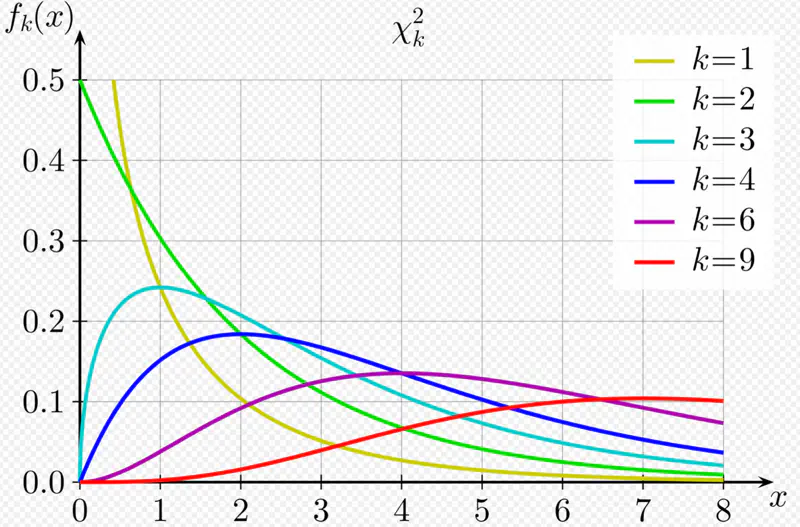
Key Properties:
- Non-negative, since sum of squares.
- Asymmetric, right skewed.
- Shape depends on the degrees of freedom; as \(\nu\) increases, the distribution becomes more symmetric and approaches a normal distribution.
Generally speaking, it represents the number of independent values that are free to vary in a dataset when estimating a parameter.
e.g.: If we have k observations and their sum = 50.
The sum of (k-1) terms can be anything, but the kth term is fixed at 50 - (sum of other (k-1) terms).
So, we have only (k-1) terms that can change independently, therefore, the DOF(\(\nu\)) = k-1.
More broadly, we can also say that sum/count of independent random variables approaches a normal distribution as the sample size increases.
Since, sample mean \(\bar{x} = \frac{sum}{n} \).
Read more about Central Limit Theorem
Note: We are dealing with categorical data, where there is a count associated with each category.
In the context of categorical data, the counts \(O_i\) are governed by multinomial distribution
(a generalisation of binomial distribution).
Multinomial distribution is defined for multiple classes or categories, ‘k’, and multiple trials ’n’.
For \(i^{th}\) category:
Probability of \(i^{th}\) category = \(p_i\)
Mean = Expected count/frequency = \(E_i = np_i \)
Variance = \(Var_i = np_i(1-p_i) \)
By Central Limit Theorem, for very large n, i.e, as \(n \rightarrow \infty\), the multinomial distribution can be approximated as a normal distribution.
The multinomial distribution of count/frequency can be approximated as :
\(O_i \approx N(np_i, np_i(1-p_i))\)
Standardized count (mean centered and variance scaled):
Under Null Hypothesis:
In Pearson’s proof of the chi-square test, the statistic is divided by the expected value (\(E_{i}\)) instead of the variance (\(Var_{i}\)),
because for count data that can be modeled using a Poisson distribution
(or a multinomial distribution where cell counts are approximately Poisson for large samples),
the variance is equal to the expected value (mean).
Therefore, \(Z_i \approx (O_{i}-E_{i})/\sqrt{E_{i}}\)
Note that the denominator is \(\sqrt{E_{i}}\) NOT \(\sqrt{Var_{i}}\).
\(O_{i}\): Observed count for \(i^{th}\) category
\(E_{i}\): Expected count for \(i^{th}\) category
Important: \(E_{i}\): Expected count should be large i.e >= 5 (typically) for a good enough approximation.
It is formed by squaring the approximately standard normal counts above, and summing them up.
For \(k\) categories, the test statistic is:
Note: For very large ’n’, the Pearson’s chi-square (\(\chi^2\)) test statistic follows a chi-square (\(\chi^2\)) distribution.
It is a non-parametric test for categorical data, i.e, does NOT make any assumption about the underlying distribution of the data, such as, normally distributed with known mean and variance; only uses observed and expected count/frequencies.
Note: Requires a large sample size.
It is used to compare the observed frequency distribution of a single categorical variable to a hypothesized or expected
probability distribution.
It can be used to determine whether a sample taken from a population follows a particular distribution,
e.g., uniform, normal, etc.
Test Statistic:
\(O_{i}\): Observed count for \(i^{th}\) category
\(E_{i}\): Expected count for \(i^{th}\) category, under null hypothesis \(H_0\)
\(k\): Number of categories
\(\nu\): Degrees of freedom = k - 1- m
\(m\): Number of parameters estimated from sample data to determine the expected probability
Note: Typical m=0, since, NO parameters are estimated.
- Kolmogorov-Smirnov (KS) Test: Compares empirical CDF with theoretical CDF of distribution.
- Anderson-Darling (AD) Test: Refinement of KS Test.
- Shapiro-Wilk (SW) Test: Specialised for normal distribution; good for small samples.
Find whether it is a fair coin (discrete uniform distribution test)?
Significance level = 5%
We need to find whether the coin is fair i.e we need to do a goodness of fit test for discrete uniform distribution.
Null Hypothesis \(H_0\): Coin is fair.
Alternative Hypothesis \(H_a\): Coin is biased towards head.
\(O_{H}\): Observed count head = 62
\(O_{T}\): Observed count head = 38
\(E_{i}\): Expected count for \(i^{th}\) category, under null hypothesis \(H_0\) = 50 i.e fair coin
\(k\): Number of categories = 2
\(\nu\): Degrees of freedom = k - 1- m = 2 - 1 - 0 = 1
Test Statistic:
Since, significance level = 5% = 0.05
Critical value = \(\chi^2(0.05,1)\) = 3.84
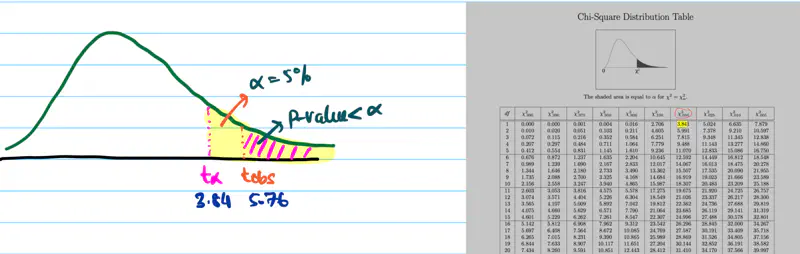
Since, \(t_{obs}\) = 5.76 > 3.84 (critical value), we reject the null hypothesis \(H_0\).
Therefore, the coin is biased towards head.
It is used to determine whether an association exists between two categorical variables,
using a contingency(dependency) table.
It is a non-parametric test, i.e, does NOT make any assumption about the underlying distribution of the data.
Test Statistic:
\(O_{ij}\): Observed count for \(cell_{i,j}\)
\(E_{ij}\): Expected count for \(cell_{i,j}\), under null hypothesis \(H_0\)
\(R\): Number of rows
\(C\): Number of columns
\(\nu\): Degrees of freedom = (R-1)*(C-1)
Let’s understand the above test statistic in more detail.
We know that, if 2 random variables A & B are independent, then,
\(P(A \cap B) = P(A, B) = P(A)*P(B)\)
i.e Joint Probability = Product of marginal probabilities.
Null Hypothesis \(H_0\): \(A\) and \(B\) are independent.
Alternative Hypothesis \(H_a\): \(A\) and \(B\) are dependent or associated.
N = Sample size
\(P(A_i) \approx \frac{Row ~~ Total_i}{N}\)
\(P(B_j) \approx \frac{Col ~~ Total_j}{N}\)
\(E_{ij}\) : Expected count for \(cell_{i,j}\) = \( N*P(A_i)*P(B_j)\)
=> \(E_{ij}\) = \(N*\frac{Row ~~ Total_i}{N}*\frac{Col ~~ Total_j}{N}\)
=> \(E_{ij}\) = \(\frac{Row ~~ Total_i * Col ~~ Total_j}{N}\)
\(O_{ij}\): Observed count for \(cell_{i,j}\)
A survey of 100 students was conducted to understand whether there is any relation between gender and beverage preference.
Below is the table that shows the number of students who prefer each beverage.
| Gender | Tea | Coffee | |
|---|---|---|---|
| Male | 20 | 30 | 50 |
| Female | 10 | 40 | 50 |
| 30 | 70 |
Significance level = 5%
Null Hypothesis \(H_0\): Gender and beverage preference are independent.
Alternative Hypothesis \(H_a\): Gender and beverage preference are dependent.
We know that Expected count for cell(i,j) = \(E_{ij}\) = \(\frac{Row ~~ Total_i * Col ~~ Total_j}{N}\)
\(E_{11} = \frac{50*30}{100} = 15\)
\(E_{12} = \frac{50*70}{100} = 35\)
\(E_{21} = \frac{50*30}{100} = 15\)
\(E_{22} = \frac{50*70}{100} = 35\)
Test Statistic:
Degrees of freedom = (R-1)(C-1) = (2-1)(2-1) = 1
Since, significance level = 5% = 0.05
Critical value = \(\chi^2(0.05,1)\) = 3.84
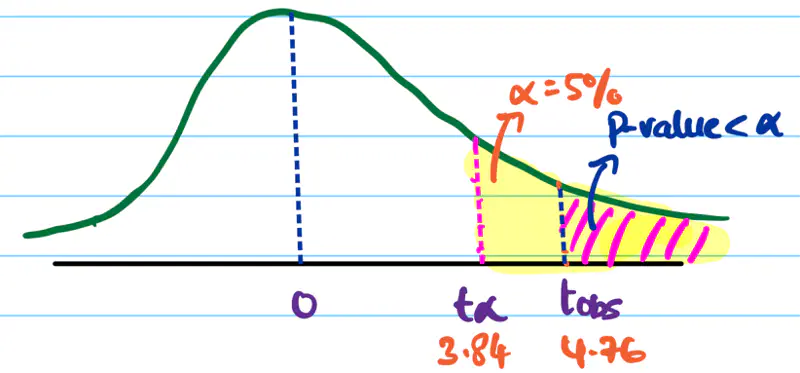
Since, \(t_{obs}\) = 4.76 > 3.84 (critical value), we reject the null hypothesis \(H_0\).
Therefore, the gender and beverage preference are dependent.
End of Section
2.2.9 - Performance Metrics
E.g.: For regression models, we have - MSE, RMSE, MAE, R^2 metric, etc.
Here, we will discuss various performance metrics for classification models.
It is a table that summarizes model’s predictions against the actual class labels, detailing where the model
succeeded and where it failed.
It is used for binary or multi-class classification problems.
| Predicted Positive | Predicted Negative | |
|---|---|---|
| Actual Positive | True Positive (TP) | False Negative (FN) |
| Actual Negative | False Positive (FP) | True Negative (TN) |
Type-1 Error:
It is the number of false positives.
e.g.: Model predicted that a patient has diabetes, but the patient actually does NOT have diabetes; “false alarm”.
Type-2 Error:
It is the number of false negatives.
e.g.: Model predicted that a patient does NOT have diabetes, but the patient actually has diabetes; “a miss”.
Many metrics are derived from the confusion matrix.
Precision:
It answers the question: “Of all the instances that the model predicted as positive, how many were actually positive?”
It measures exactness or quality of the positive predictions.
Recall:
It answers the question: “Of all the actual positive instances, how many did the model correctly identify?”
It measures completeness or coverage of the positive predictions.
F1 Score:
It is the harmonic mean of precision and recall.
It is used when we need a balance between precision and recall; also helpful when we have imbalanced data.
Harmonic mean penalizes extreme values more heavily, encouraging both metrics to be high.
| Precision | Recall | F1 Score | Mean |
|---|---|---|---|
| 0.5 | 0.5 | 0.50 | 0.5 |
| 0.7 | 0.3 | 0.42 | 0.5 |
| 0.9 | 0.1 | 0.18 | 0.5 |
Trade-Off:
Precision Focus:: Critical when cost of false positives is high.
e.g: Identify a potential terrorist.
A false positive, i.e, wrongly flagging an innocent person as a potential terrorist is very harmful.
Recall Focus:: Critical when cost of false negatives is high.
e.g.: Medical diagnosis of a serious disease.
A false negative, i.e, falsely missing a serious disease can cost someone’s life.
Analyze the performance of an access control system. Below is the data for 1000 access attempts.
| Predicted Authorised Access | Predicted Unauthorised Access | |
|---|---|---|
| Actual Authorised Access | 90 (TP) | 10 (FN) |
| Actual Unauthorised Access | 1 (FP) | 899 (TN) |
When the system allows access, it is correct 98.9% of the time.
The system caught 90% of all authorized accesses.
It is a graphical plot that shows the discriminating ability of a binary classifier system, as its
discrimination threshold is varied.
Y-axis: True Positive Rate (TPR), Recall, Sensitivity
\(TPR = \frac{TP}{TP + FN}\)
X-axis: False Positive Rate (FPR); (1 - Specificity)
\(FPR = \frac{FP}{FP + TN}\)
Note: A binary classifier model outputs a probability score between 0 and 1.
and a threshold (default=0.5) is applied to the probability score to get the final class label.
\(p \ge 0.5\) => Positive Class
\(p < 0.5\) => Negative Class
Algorithm:
- Sort the data by the probability score in descending order.
- Set each probability score as the threshold for classification and calculate the TPR and FPR for each threshold.
- Plot each pair of (TPR, FPR) for all ’n’ data points to get the final ROC curve.
e.g.:
| Patient_Id | True Label \(y_i\) | Predicted Probability Score \(\hat{y_i}\) |
|---|---|---|
| 1 | 1 | 0.95 |
| 2 | 0 | 0.85 |
| 3 | 1 | 0.72 |
| 4 | 1 | 0.63 |
| 5 | 0 | 0.59 |
| 6 | 1 | 0.45 |
| 7 | 1 | 0.37 |
| 8 | 0 | 0.20 |
| 9 | 0 | 0.12 |
| 10 | 0 | 0.05 |
Set the threshold \(\tau_1\) = 0.95, calculate \({TPR}_1, {FPR}_1\)
Set the threshold \(\tau_2\) = 0.85, calculate \({TPR}_2, {FPR}_2\)
Set the threshold \(\tau_3\) = 0.72, calculate \({TPR}_3, {FPR}_3\)
…
…
Set the threshold \(\tau_n\) = 0.05, calculate \({TPR}_n, {FPR}_n\)
Now, we have ’n’ pairs of (TPR, FPR) for all ’n’ data points.
Plot the points on a graph to get the final ROC curve.
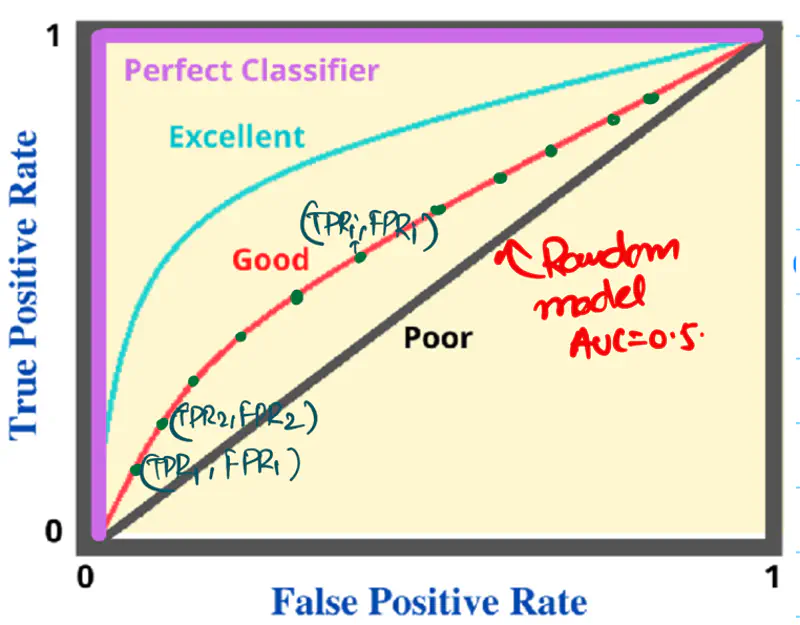
AU ROC = AUC = Area under the ROC curve = Area under the curve
Note:
- If AUC < 0.5, then invert the labels of the classes.
- ROC does NOT perform well on imbalanced data.
- Either balance the data or
- Use Precision-Recall curve.
Since, labels are randomly generated as 0/1 for binary classification, so 50% labels from each class.
Because random number generators generate numbers uniformly in the given range.
Let’s understand this with the below fraud detection example.
Below is a dataset from a fraud detection system for N = 10,000 transactions.
Fraud = 100, NOT fraud = 9900
| Predicted Fraud | Predicted NOT Fraud | |
|---|---|---|
| Actual Fraud | 80 (TP) | 20 (FN) |
| Actual NOT Fraud | 220 (FP) | 9680 (TN) |
If we check the location of above (TPR, FPR) pair on the ROC curve, then we can see that it is
very close to the top-left corner.
This means that the model is very good at detecting fraudulent transactions, but that is NOT the case.
This is happening because of the imbalanced data, i.e, count of NOT fraud transactions is 99 times
of fraudulent transactions.
Let’s look at the Precision value:
We can see that the model has poor precision,i.e, only 26.7% of flagged transactions are actual frauds.
Unacceptable precision for a good fraud detection system.
It is used to evaluate the performance of a binary classifier model across various thresholds.
It is similar to the ROC curve, but it uses Precision instead of TPR on the Y-axis.
Plots Precision (Y-axis) against Recall (X-axis) for different classification thresholds.
Note: It is useful when the data is imbalanced.
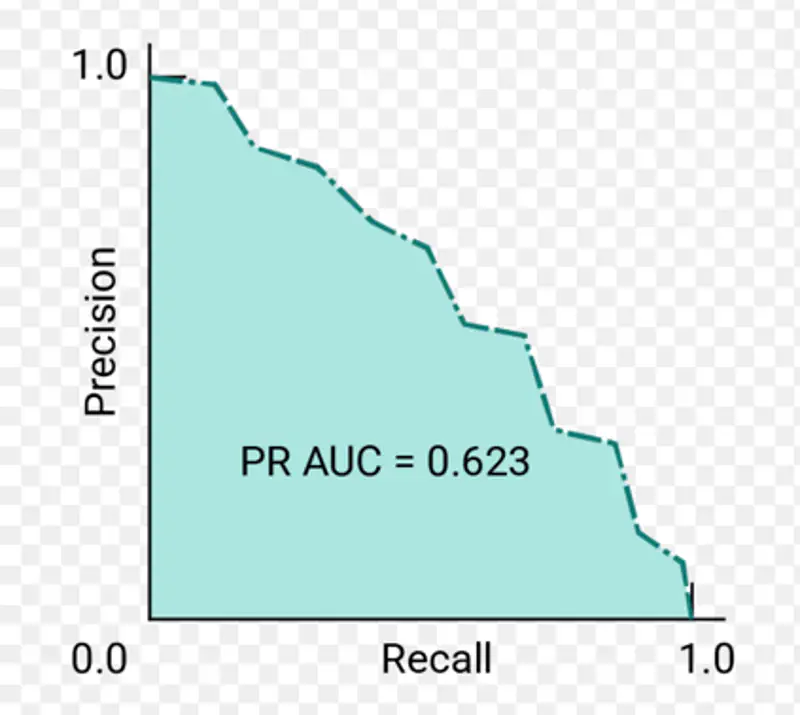
AU PRC = PR AUC = Area under Precision-Recall curve
Let’s revisit the fraud detection example discussed above to understand the utility of PR curve.
| Predicted Fraud | Predicted NOT Fraud | |
|---|---|---|
| Actual Fraud | 80 (TP) | 20 (FN) |
| Actual NOT Fraud | 220 (FP) | 9680 (TN) |
If we check the location of above (Precision, Recall) point on PRC curve, we will find that it is located near the
bottom right corner, i.e, the model performance is poor.
End of Section
2.3.1 - Vector Fundamentals
It is used to describe the surrounding space, e.g, lines, planes, 3D space, etc.
And, In machine learning, vectors are used to represent data, both the input features and the output of a model.
It is a collection of scalars(numbers) that has both magnitude and direction.
Geometrically, it is a line segment in space characterized by its length(magnitude) and direction.
By convention, we represent vectors as column vectors.
e.g.:
\(\vec{x} = \mathbf{x} = \begin{bmatrix} x_1 \\ x_2 \\ \vdots \\ x_n \end{bmatrix}_{\text{n×1}}\) i.e ’n’ rows and 1 column.
Important: In machine learning, we will use bold notation \(\mathbf{v}\) to represent vectors, instead
of arrow notation \(\mathbf{v}\).
Swap the rows and columns, i.e, a column vector becomes a row vector after transpose.
e.g:
\(\mathbf{v} = \begin{bmatrix} v_1 \\ v_2 \\ \vdots \\ v_d \end{bmatrix}_{\text{d×1}}\)
\(\mathbf{v}^\mathrm{T} = \begin{bmatrix} v_1 & v_2 & \cdots & v_d \end{bmatrix}_{\text{1×d}}\)
The length (or magnitude or norm) of a vector \(\mathbf{v}\) is the distance from the origin to the point represented
by \(\mathbf{v}\) in n-dimensional space.
\(\mathbf{v} = \begin{bmatrix} v_1 \\ v_2 \\ \vdots \\ v_d \end{bmatrix}_{\text{d×1}}\)
Length of vector:
Note: The length of a zero vector is 0.
The direction of a vector tells us where the vector points in space, independent of its length.
Direction of vector:
\[\mathbf{v} = \frac{\vec{v}} {\|\mathbf{v}\|} \]It is a collection of vectors that can be added together and scaled by numbers (scalars), such that,
the results are still in the same space.
Vector space or linear space is a non-empty set of vectors equipped with 2 operations:
- Vector addition: for any 2 vectors \(a, b\), \(a + b\) is also in the same vector space.
- Scalar multiplication: for a vector \(\mathbf{v}\), \(\alpha\mathbf{v}\) is also in the same vector space;
where \(\alpha\) is a scalar.
Note: These operations must satisfy certain rules (called axioms), such as, associativity, commutativity, distributivity, existence of a zero vector, and additive inverses.
e.g.: Set of all points in 2D is a vector space.
Addition:
We can only add vectors of the same dimension.
- Commutative: \(a + b = b + a\)
- Associative: \(a + (b + c) = (a + b) + c\)
e.g: lets add 2 real d-dimensional vectors, \(\mathbf{u} , \mathbf{v} \in \mathbb{R}^{d \times 1}\):
\(\mathbf{u} = \begin{bmatrix} u_1 \\ u_2 \\ \vdots \\ u_d \end{bmatrix}_{\text{d×1}}\),
\(\mathbf{v} = \begin{bmatrix} v_1 \\ v_2 \\ \vdots \\ v_d \end{bmatrix}_{\text{d×1}}\)
\(\mathbf{u} + \mathbf{v} = \begin{bmatrix} u_1+v_1 \\ u_2+v_2 \\ \vdots \\ u_d+v_d \end{bmatrix}_{\text{d×1}}\)
Multiplication:
1. Multiplication with Scalar:
All elements of the vector are multiplied with the scalar.
- \(c(\mathbf{u} + \mathbf{v}) = c(\mathbf{u}) + c(\mathbf{v})\)
- \((c+d)\mathbf{v} = c\mathbf{v} + d\mathbf{v}\)
- \((cd)\mathbf{v} = c(d\mathbf{v})\)
e.g:
\(\alpha\mathbf{v} = \begin{bmatrix} \alpha v_1 \\ \alpha v_2 \\ \vdots \\ \alpha v_d \end{bmatrix}_{\text{d×1}}\)
2.Inner (Dot) Product:
Inner(dot) product \(\mathbf{u} \cdot \mathbf{v}\) of 2 vectors gives a scalar output.
The two vectors must be of the same dimensions.
- \(\mathbf{u} \cdot \mathbf{v} = u_1v_1 + u_2v_2 + \cdots + u_dv_d\)
Dot product:
\(\mathbf{u} \cdot \mathbf{v} = \mathbf{u}^\mathrm{T} \mathbf{v}\)
= \(\begin{bmatrix} u_1 & u_2 & \cdots & u_d \end{bmatrix}_{\text{1×d}}
\cdot
\begin{bmatrix} v_1 \\ v_2 \\ \vdots \\ v_d
\end{bmatrix}_{\text{d×1}} = u_1v_1 + u_2v_2 + \cdots + u_dv_d\)
Geometrically, \(\mathbf{u} \cdot \mathbf{v}\) = \(|u||v|cos\theta\)
where \(\theta\) is the angle between \(\mathbf{u}\) and \(\mathbf{v}\).
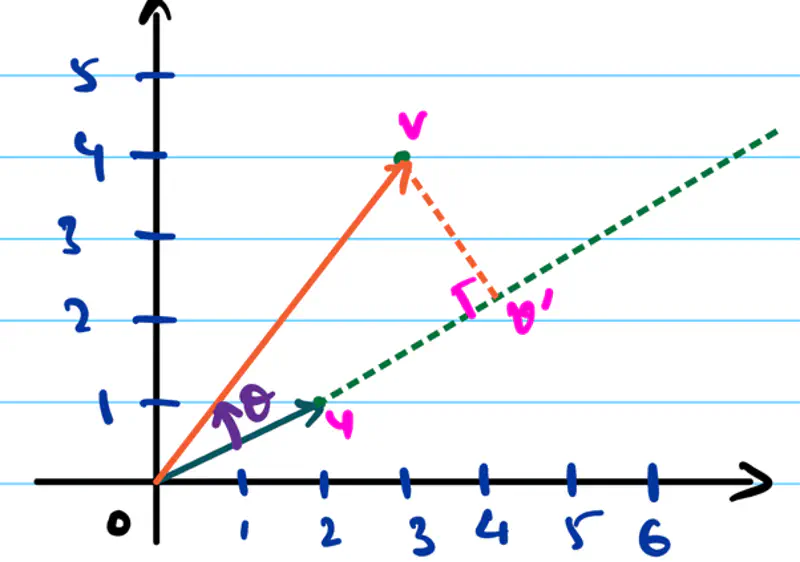
\(\mathbf{u} = \begin{bmatrix} 1 \\ \\ 2 \\ \end{bmatrix}_{\text{2×1}}\), \(\mathbf{v} = \begin{bmatrix} 3 \\ \\ 4 \\ \end{bmatrix}_{\text{2×1}}\)
\(\mathbf{u} \cdot \mathbf{v}\) = \(|u||v|cos\theta = 1 \times 3 + 2 \times 4 = 11\)
2.Outer (Tensor) Product:
Outer (tensor) product \(\mathbf{u} \otimes \mathbf{v}\) of 2 vectors gives a matrix output.
The two vectors must be of the same dimensions.
Tensor product:
\(\mathbf{u} \otimes \mathbf{v} = \mathbf{u} \mathbf{v}^\mathrm{T} \)
= \(\begin{bmatrix} u_1 \\ u_2 \\ \vdots \\ u_d \end{bmatrix}_{\text{d×1}}
\otimes
\begin{bmatrix} v_1 & v_2 & \cdots & v_d \end{bmatrix}_{\text{1×d}}
= \begin{bmatrix}
u_1v_1 & u_1v_2 & \cdots & u_1v_d \\
u_2v_1 & u_2v_2 & \cdots & u_2v_n \\
\vdots & \vdots & \ddots & \vdots \\
u_dv_1 & u_dv_2 & \cdots & u_dv_d
\end{bmatrix}
\in \mathbb{R}^{d \times d}\)
e.g:
\(\mathbf{u} = \begin{bmatrix} 1 \\ \\ 2 \\ \end{bmatrix}_{\text{2×1}}\),
\(\mathbf{v} = \begin{bmatrix} 3 \\ \\ 4 \\ \end{bmatrix}_{\text{2×1}}\)
\(\mathbf{u} \otimes \mathbf{v} = \mathbf{u} \mathbf{v}^\mathrm{T} \) = \(\begin{bmatrix} 1 \\ \\ 2 \\ \end{bmatrix}_{\text{2×1}} \otimes \begin{bmatrix} 3 & 4 \end{bmatrix}_{\text{1×2}} = \begin{bmatrix} 1 \times 3 & 1 \times 4 \\ \\ 2 \times 3 & 2 \times 4 \\ \end{bmatrix} _{\text{2×2}} = \begin{bmatrix} 3 & 4 \\ \\ 6 & 8 \\ \end{bmatrix} _{\text{2×2}}\)
Note: We will NOT discuss about cross product \(\mathbf{u} \times \mathbf{v}\); product perpendicular to both vectors.
\(\mathbf{v} = \alpha_1 \mathbf{u}_1 + \alpha_2 \mathbf{u}_2 + \cdots + \alpha_k \mathbf{u}_k\)
where \(\alpha_1, \alpha_2, \cdots, \alpha_k\) are scalars.
A set of vectors are linearly independent if NO vector in the set can be expressed as a linear combination of
the other vectors in the set.
The only solution for :
\(\alpha_1 \mathbf{u}_1 + \alpha_2 \mathbf{u}_2 + \cdots + \alpha_k \mathbf{u}_k\) = 0
is \(\alpha_1 = \alpha_2, \cdots, = \alpha_k = 0\).
e.g.:
The below 3 vectors are linearly independent.
\(\mathbf{u} = \begin{bmatrix} 1 \\ 1 \\ 1 \\ \end{bmatrix}_{\text{3×1}}\), \(\mathbf{v} = \begin{bmatrix} 1 \\ 2 \\ 3 \\ \end{bmatrix}_{\text{3×1}}\), \(\mathbf{w} = \begin{bmatrix} 1 \\ 3 \\ 6 \\ \end{bmatrix}_{\text{3×1}}\)The below 3 vectors are linearly dependent.
\(\mathbf{u} = \begin{bmatrix} 1 \\ 1 \\ 1 \\ \end{bmatrix}_{\text{3×1}}\), \(\mathbf{v} = \begin{bmatrix} 1 \\ 2 \\ 3 \\ \end{bmatrix}_{\text{3×1}}\), \(\mathbf{w} = \begin{bmatrix} 2 \\ 4 \\ 6 \\ \end{bmatrix}_{\text{3×1}}\)
because, \(\mathbf{w} = 2\mathbf{v}\), and we have a non-zero solution for the below equation:
\(\alpha_1 \mathbf{u} + \alpha_2 \mathbf{v} + \alpha_3 \mathbf{w} = 0\);
\(\alpha_1 = 0, \alpha_2 = -2, \alpha_3 = 1\) is a valid non-zero solution.
Note:
- A common method to check linear independence is to arrange the column vectors in a matrix form and calculate its determinant, if determinant ≠ 0, then the vectors are linearly independent.
- If number of vectors > number of dimensions, then the vectors are linearly dependent.
Since, the \((n+1)^{th}\) vector can be expressed as a linear combination of the other ’n’ vectors in n-dimensional space. - In machine learning, if a feature can be expressed in terms of other features, then it is linearly dependent,
and the feature is NOT bringing any new information.
e.g.: In 2 dimensions, 3 vectors \(\mathbf{x_1}, \mathbf{x_2}, \mathbf{x_3} \) are linearly dependent.
Span of a set of vectors is the geometric shape by all possible linear combinations of those vectors, such as,
line, plane, or higher dimensional volume.
e.g.:
- Span of a single vector (1,0) is the entire X-axis.
- Span of 2 vectors (1,0) and (0,1) is the entire X-Y (2D) plane, as any vector in 2D plane can be expressed as a linear combination of the 2 vectors - (1,0) and (0,1).
It is the minimal set of linearly independent vectors that spans or defines the entire vector space,
providing a unique co-ordinate system for every vector within the space.
Every vector in the vector space can be represented as a unique linear combination of the basis vectors.
e.g.:
- X-axis(1,0) and Y-axis(0,1) are the basis vectors of 2D space or form the co-ordinate system.
- \(\mathbf{u} = (3,1)\) and \(\mathbf{v} = (-1, 2) \) are the basis of skewed or parallelogram co-ordinate system.
Note: Basis = Dimensions
Two vectors are orthogonal if their dot product is 0.
A set of vectors \(\mathbf{x_1}, \mathbf{x_2}, \cdots ,\mathbf{x_n} \) are said to be orthogonal if:
\(\mathbf{x_i} \cdot \mathbf{x_j} = 0 \forall i⍯j\), for every pair, i.e, every pair must be orthogonal.
e.g.:
- \(\mathbf{u} = (1,0)\) and \(\mathbf{v} = (0,1) \) are orthogonal vectors.
- \(\mathbf{u} = (1,0)\) and \(\mathbf{v} = (1,1) \) are NOT orthogonal vectors.
Note:
Orthogonal vectors are linearly independent, but the inverse may NOT be true,
i.e, linear independence does NOT imply that vectors are orthogonal. e.g.:
Vectors \(\mathbf{u} = (2,0)\) and \(\mathbf{v} = (1,3) \) are linearly independent but NOT orthogonal.
Since, \(\mathbf{u} \cdot \mathbf{v} = 2*1 + 3*0 = 2 ⍯ 0\).Orthogonality is the most extreme case of linear independence, i.e \(90^{\degree}\) apart or perpendicular.
Orthonormal vectors are vectors that are orthogonal and have unit length.
A set of vectors \(\mathbf{x_1}, \mathbf{x_2}, \cdots ,\mathbf{x_n} \) are said to be orthonormal if:
\(\mathbf{x_i} \cdot \mathbf{x_j} = 0 \forall i⍯j\) and \(\|\mathbf{x_i}\| = 1\), i.e, unit vector.
e.g.:
- \(\mathbf{u} = (1,0)\) and \(\mathbf{v} = (0,1) \) are orthonormal vectors.
- \(\mathbf{u} = (1,0)\) and \(\mathbf{v} = (0,2) \) are NOT orthonormal vectors.
It is a set of vectors that functions as a basis for a space while also being orthonormal,
meaning each vector is a unit vector (has a length of 1) and all vectors are mutually perpendicular (orthogonal) to each other.
A set of vectors \(\mathbf{x_1}, \mathbf{x_2}, \cdots ,\mathbf{x_n} \) are said to be orthonormal basis of a vector space
\(\mathbb{R}^{n \times 1}\), if every vector:
e.g.:
- \(\mathbf{u} = (1,0)\) and \(\mathbf{v} = (0,1) \) form an orthonormal basis for 2-D space.
- \(\mathbf{u} = (\frac{1}{\sqrt{2}},\frac{1}{\sqrt{2}})\) and \(\mathbf{v} = (-\frac{1}{\sqrt{2}},-\frac{1}{\sqrt{2}}) \)
also form an orthonormal basis for 2-D space.
Note: In a n-dimensional space, there are only \(n\) possible orthonormal bases.
End of Section
2.3.2 - Matrix Operations
Matrices let us store, manipulate, and transform data efficiently.
e.g:
- Represent a system of linear equations AX=B.
- Data representation, such as images, that are stored as a matrix of pixels.
- When multiplied, matrix, linearly transforms a vector, i.e, its direction and magnitude, making it useful in image rotation, scaling, etc.
e.g:
\( \mathbf{A} = \begin{bmatrix} a_{11} & a_{12} & \cdots & a_{1n} \\ a_{21} & a_{22} & \cdots & a_{2n} \\ \vdots & \vdots & \ddots & \vdots \\ a_{m1} & a_{m2} & \cdots & a_{mn} \end{bmatrix} _{\text{m x n}} \)
\( \mathbf{A}^T = \begin{bmatrix} a_{11} & a_{21} & \cdots & a_{m1} \\ a_{12} & a_{22} & \cdots & a_{m2} \\ \vdots & \vdots & \ddots & \vdots \\ a_{1n} & a_{2n} & \cdots & a_{mn} \end{bmatrix} _{\text{n x m}} \)
Important: \( (AB)^T = B^TA^T \)
Addition:
We add two matrices by adding the corresponding elements.
They must have same dimensions.
\( \mathbf{A} + \mathbf{B} =
\begin{bmatrix}
a_{11} + b_{11} & a_{12} + b_{12} & \cdots & a_{1n} + b_{1n} \\
a_{21} + b_{21} & a_{22} + b_{22} & \cdots & a_{2n} + b_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{m1} + b_{m1} & a_{m2} + b_{m2} & \cdots & a_{mn} + b_{mn}
\end{bmatrix}
_{\text{m x n}}
\)
Multiplication:
We can multiply two matrices only if their inner dimensions are equal.
\( \mathbf{C}_{m x n} = \mathbf{A}_{m x d} ~ \mathbf{B}_{d x n} \)
\( c_{ij} \) = Dot product of \(i^{th}\) row of A and \(j^{th}\) row of B.
\( c_{ij} = \sum_{k=1}^{d} A_{ik} * B_{kj} \)
=> \( c_{11} = a_{11} * b_{11} + a_{12} * b_{21} + \cdots + a_{1d} * b_{d1} \)
e.g:
\(
\mathbf{A} =
\begin{bmatrix}
1 & 2 \\
\\
3 & 4
\end{bmatrix}
_{\text{2 x 2}},
\mathbf{B} =
\begin{bmatrix}
5 & 6 \\
\\
7 & 8
\end{bmatrix}
_{\text{2 x 2}}
\)
\(
\mathbf{C} = \mathbf{A} \times \mathbf{B} =
\begin{bmatrix}
1 * 5 + 2 * 7 & 1 * 6 + 2 * 8 \\
\\
3 * 5 + 4 * 7 & 3 * 6 + 4 * 8
\end{bmatrix}
_{\text{2 x 2}}
= \begin{bmatrix}
19 & 22 \\
\\
43 & 50
\end{bmatrix}
_{\text{2 x 2}}
\)
Key Properties:
- AB ≠ BA ; NOT commutative
- (AB)C = A(BC) ; Associative
- A(B+C) = AB+AC ; Distributive
Square Matrix:
It is a matrix with same number of rows and columns (m=n).
\(
\mathbf{A} =
\begin{bmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{n1} & a_{n2} & \cdots & a_{nn}
\end{bmatrix}
_{\text{n x n}}
\)
Diagonal Matrix:
It is a square matrix with all non-diagonal elements equal to zero.
e.g.:
\( \mathbf{D} =
\begin{bmatrix}
d_{11} & 0 & \cdots & 0 \\
0 & d_{22} & \cdots & 0 \\
\vdots & \vdots & \ddots & \vdots \\
0 & 0 & \cdots & d_{nn}
\end{bmatrix}
_{\text{n x n}}
\)
Note: Product of 2 diagonal matrices is a diagonal matrix.
Lower Triangular Matrix:
It is a square matrix with all elements above the diagonal equal to zero.
e.g.:
\( \mathbf{L} =
\begin{bmatrix}
l_{11} & 0 & \cdots & 0 \\
l_{21} & l_{22} & \cdots & 0 \\
\vdots & \vdots & \ddots & \vdots \\
l_{n1} & l_{n2} & \cdots & l_{nn}
\end{bmatrix}
_{\text{n x n}}
\)
Note: Product of 2 lower triangular matrices is an lower triangular matrix.
Upper Triangular Matrix:
It is a square matrix with all elements below the diagonal equal to zero.
e.g.:
\( \mathbf{U} =
\begin{bmatrix}
u_{11} & u_{12} & \cdots & u_{1n} \\
0 & u_{22} & \cdots & u_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
0 & 0 & \cdots & u_{nn}
\end{bmatrix}
_{\text{n x n}}
\)
Note: Product of 2 upper triangular matrices is an upper triangular matrix.
Symmetric Matrix:
It is a square matrix that is equal to its own transpose, i.e, flip the matrix along its diagonal,
it remains unchanged.
\( \mathbf{A} = \mathbf{A}^T \)
e.g.:
\( \mathbf{S} =
\begin{bmatrix}
s_{11} & s_{12} & \cdots & s_{1n} \\
s_{12} & s_{22} & \cdots & s_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
s_{1n} & s_{2n} & \cdots & s_{nn}
\end{bmatrix}
_{\text{n x n}}
\)
Note: Diagonal matrix is a symmetric matrix.
Skew Symmetric Matrix:
It is a square symmetric matrix where the elements across the diagonal have opposite signs.
Also called Anti Symmetric Matrix.
\( \mathbf{A} = -\mathbf{A}^T \)
e.g.:
\( \mathbf{S} =
\begin{bmatrix}
0 & s_{12} & \cdots & s_{1n} \\
-s_{12} & 0 & \cdots & s_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
-s_{1n} & -s_{2n} & \cdots & 0
\end{bmatrix}
_{\text{n x n}}
\)
Note: Diagonal elements of a skew symmetric matrix are zero, since the number should be equal to its negative.
Identity Matrix:
It is a square matrix with all the diagonal values equal to 1, rest of the elements are equal to zero.
e.g.:
\( \mathbf{I} =
\begin{bmatrix}
1 & 0 & \cdots & 0 \\
0 & 1 & \cdots & 0 \\
\vdots & \vdots & \ddots & \vdots \\
0 & 0 & \cdots & 1
\end{bmatrix}
_{\text{n x n}}
\)
Important:
\( \mathbf{I} \times \mathbf{A} = \mathbf{A} \times \mathbf{I} = \mathbf{A} \)
Trace:
It is the sum of the elements on the main diagonal of a square matrix.
Note: Main diagonal is NOT defined for a rectangular matrix.
\( \text{trace}(A) = \sum_{i=1}^{n} a_{ii} = a_{11} + a_{22} + \cdots + a_{nn}\)
If \( \mathbf{A} \in \mathbb{R}^{m \times n} \), and \( \mathbf{B} \in \mathbb{R}^{n \times m} \), then
\( \text{trace}(AB)_{m \times m} = \text{trace}(BA)_{n \times n} \)
Determinant:
It is a scalar value that reveals crucial properties about the matrix and its linear transformation.
- If determinant = 0 => singular matrix, i.e linearly dependent rows or columns.
- Determinant is also equal to the scaling factor of the linear transformation.
\(a_{1j} \) = element in the first row and j-th column
\(M_{1j} \) = submatrix of the matrix excluding the first row and j-th column
If n = 2, then,
\( |A| = a_{11} \, a_{22} - a_{12} \, a_{21} \)
Singular Matrix:
A square matrix with linearly dependent rows or columns, i.e, determinant = 0.
A singular matrix collapses space, say, a 3D space, into a 2D space or a higher dimensional space
to a lower dimensional space, making the transformation impossible to reverse.
Hence, a singular matrix is NOT invertible; i.e, inverse does NOT exist.
Singular matrix is also NOT invertible, because the inverse has division by determinant, and its determinant is zero.
Also called rank deficient matrix, because the rank < number of dimensions of the matrix,
due to the presence of linearly dependent rows or columns.
e.g: Below is a linearly dependent 2x2 matrix.
\( \mathbf{A} =
\begin{bmatrix}
a_{11} & a_{12} \\
\\
\beta a_{11} & \beta a_{12}
\end{bmatrix}
_{\text{2 x 2}},
det(\mathbf{A}) =
a_{11}\cdot \beta a_{12} - a_{12} \cdot \beta a_{11} = 0
\)
Note:
- \( det(A) = det(A^T) \)
- \( det(\beta A) = \beta^n det(A) \), where n is the number of dimensions of A and \(\beta\) is a scalar.
Inverse:
It is a square matrix that when multiplied by the original matrix, gives the identity matrix.
\( \mathbf{A} \mathbf{A}^{-1} = \mathbf{A}^{-1} \mathbf{A} = \mathbf{I} \)
Steps to compute inverse of a matrix:
- Calculate the determinant of the matrix. \( |A| = \det(A) \)
- For each element \(a_{ij}\), compute its minor \(M_{ij}\).
\(M_{ij}\) is the determinant of the submatrix of the matrix excluding the i-th row and j-th column. - Form the co-factor matrix \(C_{ij}\), using the minor.
\( C_{ij} = (-1)^{\,i+j}\,M_{ij} \) - Take the transpose of the cofactor matrix to get the adjugate matrix.
\( \mathrm{adj}(A) = \mathrm{C}^\mathrm{T} \) - Compute the inverse:
\( A^{-1} = \frac{1}{|A|}\,\mathrm{adj}(A) = \frac{1}{|A|}\,\mathrm{C}^\mathrm{T}\)
e.g:
\(
\mathbf{A} =
\begin{bmatrix}
1 & 0 & 3\\
2 & 1 & 1\\
1 & 1 & 1 \\
\end{bmatrix}
_{\text{3 x 3}}
\),
Co-factor matrix:
\(
\mathbf{C} =
\begin{bmatrix}
0 & -1 & 1\\
3 & -2 & -1\\
-3 & 5 & 1 \\
\end{bmatrix}
_{\text{3 x 3}}
\)
Adjugate matrix, adj(A) =
\(
\mathbf{C}^\mathrm{T} =
\begin{bmatrix}
0 & 3 & -3\\
-1 & -2 & 5\\
1 & -1 & 1 \\
\end{bmatrix}
_{\text{3 x 3}}
\)
determinant of \( \mathbf{A} \) =
\(
\begin{vmatrix}
1 & 0 & 3\\
2 & 1 & 1\\
1 & 1 & 1 \\
\end{vmatrix}
= 0 -\cancel 1 + \cancel 1 + \cancel3 - 2 -\cancel1 -\cancel 3 + 5 + \cancel1 = 3
\)
\(
\mathbf{A}^{-1} = \frac{1}{|A|}\,\mathrm{adj}(A)
= \frac{1}{3}\,
\begin{bmatrix}
0 & 3 & -3\\
-1 & -2 & 5\\
1 & -1 & 1 \\
\end{bmatrix}
_{\text{3 x 3}}
\)
=> \(
\mathbf{A}^{-1} =
\begin{bmatrix}
0 & 1 & -1\\
-1/3 & -2/3 & 5/3\\
1/3 & -1/3 & 1/3 \\
\end{bmatrix}
_{\text{3 x 3}}
\)
Note:
- Inverse of an Identity matrix is the Identity matrix itself.
- Inverse of \( \beta \mathbf{A} = \frac{1}{\beta}\mathbf{A}^{-1} \).
- \( (\mathbf{A}^\mathrm{T})^{-1} = (\mathbf{A}^{-1})^\mathrm{T} \).
- Inverse of a diagonal matrix is a diagonal matrix with reciprocal of the diagonal elements.
- Inverse of a upper triangular matrix is the upper triangular matrix.
- Inverse of a lower triangular matrix is the lower triangular matrix.
- Symmetric matrix, if invertible, then \((\mathbf{A}^{-1})^\mathrm{T} = \mathbf{A}^{-1}\), since, \(\mathbf{A} = \mathbf{A}^\mathrm{T}\)
It is a square matrix that whose rows and columns are orthonormal vectors, i.e, they are perpendicular to each other
and have a unit length.
\( \mathbf{A} \mathbf{A}^\mathrm{T} = \mathbf{A}^\mathrm{T} \mathbf{A} = \mathbf{A} \mathbf{A}^{-1} = \mathbf{I} \)
=> \( \mathbf{A}^\mathrm{T} = \mathbf{A}^{-1} \)
Note:
- Orthogonal matrix preserves the length and angles of vectors, acting as rotation or reflection in geometry.
- The determinant of an orthogonal matrix is always +1 or -1, because \( \mathbf{A} \mathbf{A}^\mathrm{T} = \mathbf{I} \)
Let’s check out why the rows and columns of an orthogonal matrix are orthonormal.
\( \mathbf{A} =
\begin{bmatrix}
a_{11} & a_{12} \\
\\
a_{21} & a_{22}
\end{bmatrix}
_{\text{2 x 2}}
\),
=> \( \mathbf{A}^\mathrm{T} =
\begin{bmatrix}
a_{11} & a_{21} \\
\\
a_{12} & a_{22}
\end{bmatrix}
_{\text{2 x 2}}
\)
Since, \( \mathbf{A} \mathbf{A}^\mathrm{T} = \mathbf{I} \)
\(
\begin{bmatrix}
a_{11} & a_{12} \\
\\
a_{21} & a_{22}
\end{bmatrix}
\begin{bmatrix}
a_{11} & a_{21} \\
\\
a_{12} & a_{22}
\end{bmatrix}
= \begin{bmatrix}
a_{11}^2 + a_{12}^2 & a_{11}a_{21} + a_{12}a_{22} \\
\\
a_{21}a_{11} + a_{21}a_{12} & a_{21}^2 + a_{22}^2
\end{bmatrix}
= \begin{bmatrix}
1 & 0 \\
\\
0 & 1
\end{bmatrix}
\)
Equating the terms, we get:
\( a_{11}^2 + a_{12}^2 = 1 \) => row 1 is a unit vector
\( a_{21}^2 + a_{22}^2 = 1 \) => row 2 is a unit vector
\( a_{11}a_{21} + a_{12}a_{22} = 0 \) => row 1 and row 2 are orthogonal to each other, since dot product = 0
\( a_{21}a_{11} + a_{21}a_{12} = 0 \) => row 1 and row 2 are orthogonal to each other, since dot product = 0
Therefore, the rows and columns of an orthogonal matrix are orthonormal.
Let, \(\mathbf{Q}\) is an orthogonal matrix, and \(\mathbf{v}\) is a vector.
Let’s calculate the length of \( \mathbf{Q} \mathbf{v} \)
Therefore, linear transformation of a vector by an orthogonal matrix does NOT change its length.
\( 2x + y = 5 \)
\( x + 2y = 4 \)
Lets solve the system of equations using matrix:
\(\begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix}
\)
\(
\begin{bmatrix}
x \\
\\
y
\end{bmatrix}
= \begin{bmatrix}
5 \\
\\
4
\end{bmatrix}
\)
The above equation can be written as:
\(\mathbf{AX} = \mathbf{B} \)
=> \( \mathbf{X} = \mathbf{A}^{-1} \mathbf{B} \)
\( \mathbf{A} = \begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix},
\mathbf{B} = \begin{bmatrix}
5 \\
\\
4
\end{bmatrix},
\mathbf{X} = \begin{bmatrix}
x \\
\\
y
\end{bmatrix}
\)
Lets compute the inverse of A matrix:
\( \mathbf{A}^{-1} = \frac{1}{3}\begin{bmatrix}
2 & -1 \\
\\
-1 & 2
\end{bmatrix}
\)
Since, \( \mathbf{X} = \mathbf{A}^{-1} \mathbf{B} \)
=> \( \mathbf{X} = \begin{bmatrix}
x \\
\\
y
\end{bmatrix}
= \frac{1}{3}\begin{bmatrix}
2 & -1 \\
\\
-1 & 2
\end{bmatrix}
\begin{bmatrix}
5 \\
\\
4
\end{bmatrix}
= \frac{1}{3}\begin{bmatrix}
6 \\
\\
3
\end{bmatrix}
= = \begin{bmatrix}
2 \\
\\
1
\end{bmatrix}
\)
Therefore, \( x = 2 \) and \( y = 1 \).
End of Section
2.3.3 - Eigen Value Decomposition
It tells us about the characteristic properties of a matrix.
Multiplying a vector by a matrix can change the direction or magnitude or both of the vector.
\( \mathbf{A} = \begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix}
\),
\(\mathbf{u} = \begin{bmatrix} 0 \\ \\ 1 \\ \end{bmatrix}\),
\(\mathbf{v} = \begin{bmatrix} 1 \\ \\ 1 \\ \end{bmatrix}\)
\(\mathbf{Au} = \begin{bmatrix} 1 \\ \\ 2 \\ \end{bmatrix}\) , \(\quad\)
\(\mathbf{Av} = \begin{bmatrix} 3 \\ \\ 3 \\ \end{bmatrix}\)
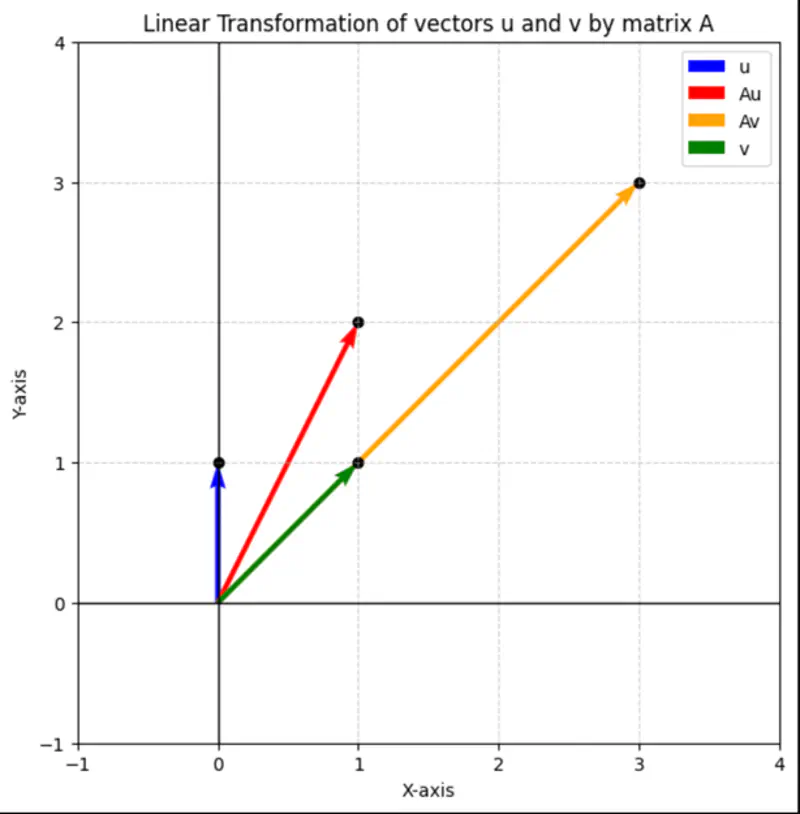
It might get scaled up or down but does not change its direction.
Result of linear transformation, i.e, multiplying the vector by a matrix, is just a scalar multiple of the original vector.
\(|\lambda| > 1 \): Vector stretched
\(0 < |\lambda| < 1 \): Vector shrunk
\(|\lambda| = 1 \): Same size
\(\lambda < 0 \): Vector’s direction is reversed
Since, for an eigen vector, result of linear transformation, i.e, multiplying the vector by a matrix, is just a scalar multiple of the original vector, =>
\[ \mathbf{A} \mathbf{v} = \lambda \mathbf{v} \\ \mathbf{A} \mathbf{v} - \lambda \mathbf{v} = 0 \\ \mathbf{v}(\mathbf{A} - \lambda \mathbf{I}) = 0 \\ \]For a non-zero eigen vector, \((\mathbf{A} - \lambda \mathbf{I})\) must be singular, i.e,
\(det(\mathbf{A} - \lambda \mathbf{I}) = 0 \)
If, \( \mathbf{A} =
\begin{bmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{n1} & a_{n2} & \cdots & a_{nn}
\end{bmatrix}
_{\text{n x n}}
\),
then, \((\mathbf{A} - \lambda \mathbf{I}) =
\begin{bmatrix}
a_{11}-\lambda & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22}-\lambda & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{n1} & a_{n2} & \cdots & a_{nn}-\lambda
\end{bmatrix}
_{\text{n x n}}
\)
\(det(\mathbf{A} - \lambda \mathbf{I})) = 0 \), will give us a polynomial equation in \(\lambda\) of degree \(n\),
we need to solve this polynomial equation to get the \(n\) eigen values.
e.g.:
\( \mathbf{A} = \begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix}
\),
\( \quad det(\mathbf{A} - \lambda \mathbf{I})) = 0 \quad \) =>
\( det \begin{bmatrix}
2-\lambda & 1 \\
\\
1 & 2-\lambda
\end{bmatrix} = 0
\)
\( (2-\lambda)^2 - 1 = 0\)
\( => |\lambda - 2| = \pm 1 \)
\( => \lambda_1 = 3\) and \( \lambda_2 = 1\)
Therefore, eigen vectors corresponding to eigen values \(\lambda_1 = 3\) and \( \lambda_2 = 1\) are:
\((\mathbf{A} - \lambda \mathbf{I}) \mathbf{v} = 0 \)
\(\lambda_1 = 3\)
=>
\(
\begin{bmatrix}
2-3 & 1 \\
\\
1 & 2-3
\end{bmatrix} \begin{bmatrix} v_1 \\ \\ v_2 \\ \end{bmatrix} =
\begin{bmatrix}
-1 & 1 \\
\\
1 & -1
\end{bmatrix} \begin{bmatrix} v_1 \\ \\ v_2 \\ \end{bmatrix} = 0
\)
=> Both the equations will be \(v_1 - v_2 = 0 \), i.e, \(v_1 = v_2\)
So, we can choose any vector, where x-axis and y-axis components are same, i.e, \(v_1 = v_2\)
=> Eigen vector: \(v_1 = \begin{bmatrix} 1 \\ \\ 1 \\ \end{bmatrix}\)
Similarly, for \(\lambda_2 = 1\)
we will get, eigen vector: \(v_2 = \begin{bmatrix} 1 \\ \\ -1 \\ \end{bmatrix}\)
\(\mathbf{Iv} = \lambda \mathbf{v}\)
Therefore, identity matrix has only one eigen value \(\lambda = 1\), and all non-zero vectors can be eigen vectors.
\(\mathbf{A} = \begin{bmatrix} 0 & 1 \\ \\ -1 & 0 \end{bmatrix} \), \( \quad det(\mathbf{A} - \lambda \mathbf{I}) = 0 \quad \) => det \(\begin{bmatrix} 0-\lambda & 1 \\ \\ -1 & 0-\lambda \end{bmatrix} = 0 \)
=> \(\lambda^2 + 1 = 0\) => \(\lambda = \pm i\)
So, eigen values are complex.
e.g.:
\( \mathbf{D} = \begin{bmatrix} d_{11} & 0 & \cdots & 0 \\ 0 & d_{22} & \cdots & 0 \\ \vdots & \vdots & \ddots & \vdots \\ 0 & 0 & \cdots & d_{nn} \end{bmatrix} _{\text{n x n}} \), \( \quad det(\mathbf{D} - \lambda \mathbf{I}) = 0 \quad \) => det \( \begin{bmatrix} d_{11}-\lambda & 0 & \cdots & 0 \\ 0 & d_{22}-\lambda & \cdots & 0 \\ \vdots & \vdots & \ddots & \vdots \\ 0 & 0 & \cdots & d_{nn}-\lambda \end{bmatrix} _{\text{n x n}} \)
=> \((d_{11}-\lambda)(d_{22}-\lambda) \cdots (d_{nn}-\lambda) = 0\)
=> \(\lambda = d_{11}, d_{22}, \cdots, d_{nn}\)
So, eigen values are the diagonal elements of the matrix.
- For a \(n \times n\) matrix, there are \(n\) eigen values.
- Eigen values need NOT be unique, e.g, identity matrix has only one eigen value \(\lambda = 1\).
- Sum of eigen values = trace of matrix = sum of diagonal elements, i.e,
\(tr(\mathbf{A}) = \lambda_1 + \lambda_2 + \cdots + \lambda_n\). - Product of eigen values = determinant of matrix, i.e,
\(det(\mathbf{A}) = |\mathbf{A}|= \lambda_1 \lambda_2 \cdots \lambda_n\). e.g: For the example matrix given above,
\( \mathbf{A} = \begin{bmatrix} 2 & 1 \\ \\ 1 & 2 \end{bmatrix} \), \(\quad \lambda_1 = 3\) and \( \lambda_2 = 1\)
\(tr(\mathbf{A}) = 2 + 2 = 3 + 1 = 4\)
\(det(\mathbf{A}) = 2 \times 2 - 1 \times 1 = 3 \times 1 = 3 \)
- Eigen vectors corresponding to distinct eigen values of a real symmetric matrix are orthogonal to each other.
Proof:
Let, \(\mathbf{v_1}, \mathbf{v_2}\) be eigen vectors corresponding to distinct eigen value \(\lambda_1,\lambda_2\) of a real symmetric matrix \(\mathbf{A} = \mathbf{A}^T\).
We know that for eigen vectors -
\[ \mathbf{Av_1} = \lambda_1 \mathbf{v_1} ~and~ \mathbf{Av_2} = \lambda_2 \mathbf{v_2} \\[10pt] \text{ let's calculate the dot product: } \\[10pt] \mathbf{Av_1} \cdot \mathbf{v_2} = (\mathbf{Av_1})^T\mathbf{v_2} = \mathbf{v_1^TA^Tv_2} = \mathbf{v_1^T} ~ \mathbf{Av_2}, \quad \text {since } \mathbf{A} = \mathbf{A}^T\\[10pt] => (\mathbf{Av_1}) \cdot \mathbf{v_2} = \mathbf{v_1} \cdot (\mathbf{Av_2}) \\[10pt] => (\lambda_1 \mathbf{v_1}) \cdot \mathbf{v_2} = \mathbf{v_1} \cdot (\lambda_2\mathbf{v_2}) \\[10pt] => \lambda_1 (\mathbf{v_1 \cdot v_2}) = \lambda_2 (\mathbf{v_1 \cdot v_2}) \\[10pt] => (\lambda_1 - \lambda_2) (\mathbf{v_1} \cdot \mathbf{v_2}) = 0 \\[10pt] \text{ since, eigen values are distinct,} => \lambda_1 ≠ \lambda_2 \\[10pt] \therefore \mathbf{v_1} \cdot \mathbf{v_2} = 0 \\[10pt] => \text{ eigen vectors are orthogonal to each other,} => \mathbf{v_1} \perp \mathbf{v_2} \\[10pt] \]
Let’ calculate the 2nd power of a square matrix.
e.g.:
\(\mathbf{A} = \begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix}
\),
\(\quad \mathbf{A}^2 = \begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix} \begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix} = \begin{bmatrix}
5 & 4 \\
\\
4 & 5
\end{bmatrix}
\)
So, we need to find an easier way to calculate the power of a matrix.
Let’s calculate the 2nd power of a diagonal matrix.
e.g.:
\(\mathbf{A} = \begin{bmatrix}
3 & 0 \\
\\
0 & 2
\end{bmatrix}
\),
\(\quad \mathbf{A}^2 = \begin{bmatrix}
3 & 0 \\
\\
0 & 2
\end{bmatrix} \begin{bmatrix}
3 & 0 \\
\\
0 & 2
\end{bmatrix} = \begin{bmatrix}
9 & 0 \\
\\
0 & 4
\end{bmatrix}
\)
Note that when we square the diagonal matrix, then all the diagonal elements got squared.
Similarly, if we want to calculate the kth power of a diagonal matrix, then all we need to do is to just compute the kth
powers of all diagonal elements, instead of complex matrix multiplications.
\(\quad \mathbf{A}^k = \begin{bmatrix}
3^k & 0 \\
\\
0 & 2^k
\end{bmatrix}
\)
Therefore, if we diagonalize a square matrix then the computation of power of the matrix will become very easy.
Next, let’s see how to diagonalize a matrix.
\(\mathbf{V}\): Matrix of all eigen vectors(as columns) of matrix \(\mathbf{A}\)
\( \mathbf{V} = \begin{bmatrix}
\mathbf{v}_1 & \mathbf{v}_2 & \cdots & \mathbf{v}_n \\
\vdots & \vdots & \ddots & \vdots \\
\vdots & \vdots & \vdots & \vdots\\
\end{bmatrix}
\),
where, each column is an eigen vector corresponding to an eigen value \(\lambda_i\).
If \( \mathbf{v}_1 \mathbf{v}_2 \cdots \mathbf{v}_n \) are linearly independent, then,
\(det(\mathbf{V}) ~⍯ ~ 0\).
=> \(\mathbf{V}\) is NOT singular and \(\mathbf{V}^{-1}\) exists.
\(\Lambda\): Diagonal matrix of all eigen values of matrix \(\mathbf{A}\)
\( \Lambda = \begin{bmatrix}
\lambda_1 & 0 & \cdots & 0 \\
\\
0 & \lambda_2 & \cdots & 0 \\
\vdots & \vdots & \ddots & \vdots \\
0 & 0 & \cdots & \lambda_n
\end{bmatrix}
\),
where, each diagonal element is an eigen value corresponding to an eigen vector \(\mathbf{v}_i\).
Let’s recall the characteristic equation of a matrix for calculating eigen values:
\(\mathbf{A} \mathbf{v} = \lambda \mathbf{v}\)
Let’s use the consolidated matrix for eigen values and eigen vectors described above:
i.e, \(\mathbf{A} \mathbf{V} = \mathbf{\Lambda} \mathbf{V}\)
For diagonalisation:
=> \(\mathbf{\Lambda} = \mathbf{V}^{-1} \mathbf{A} \mathbf{V}\)
We can see that, using the above equation, we can represent the matrix \(\mathbf{A}\) as a
diagonal matrix \(\mathbf{\Lambda}\) using the matrix of eigen vectors \(\mathbf{V}\).
Just reshuffling the above equation will give us the Eigen Value Decomposition of the matrix \(\mathbf{A}\):
\(\mathbf{A} = \mathbf{V} \mathbf{\Lambda} \mathbf{V}^{-1}\)
Important: Given that \(\mathbf{V}^{-1}\) exists, i.e, all the eigen vectors are linearly independent.
Let’s revisit the example given above:
\(\mathbf{A} = \begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix}
\),
\(\quad \lambda_1 = 3\) and \( \lambda_2 = 1\),
\(\mathbf{v_1} = \begin{bmatrix} 1 \\ \\ 1 \\ \end{bmatrix}\),
\(\mathbf{v_2} = \begin{bmatrix} 1 \\ \\ -1 \\ \end{bmatrix}\)
=> \( \mathbf{V} = \begin{bmatrix}
1 & 1 \\
\\
1 & -1 \\
\end{bmatrix}
\),
\(\quad \mathbf{\Lambda} = \begin{bmatrix}
3 & 0 \\
\\
0 & 1
\end{bmatrix}
\)
\( \because \mathbf{A} = \mathbf{V} \mathbf{\Lambda} \mathbf{V}^{-1}\)
We know, \( \mathbf{V} ~and~ \mathbf{\Lambda} \), we need to calculate \(\mathbf{V}^{-1}\).
\(\mathbf{V}^{-1} = \frac{1}{2}\begin{bmatrix} 1 & 1 \\ \\ 1 & -1 \\ \end{bmatrix} \)
\( \therefore \mathbf{V} \mathbf{\Lambda} \mathbf{V}^{-1} = \begin{bmatrix}
1 & 1 \\
\\
1 & -1 \\
\end{bmatrix} \begin{bmatrix}
3 & 0 \\
\\
0 & 1
\end{bmatrix} \frac{1}{2}\begin{bmatrix}
1 & 1 \\
\\
1 & -1 \\
\end{bmatrix}
\)
\( = \frac{1}{2}
\begin{bmatrix}
3 & 1 \\
\\
3 & -1 \\
\end{bmatrix} \begin{bmatrix}
1 & 1 \\
\\
1 & -1 \\
\end{bmatrix} = \frac{1}{2}\begin{bmatrix}
4 & 2 \\
\\
2 & 4 \\
\end{bmatrix}
\)
\( = \begin{bmatrix}
2 & 1 \\
\\
1 & 2
\end{bmatrix} = \mathbf{A}
\)
Spectral decomposition is a specific type of eigendecomposition that applies to a symmetric matrix,
requiring its eigenvectors to be orthogonal.
In contrast, a general eigendecomposition applies to any diagonalizable matrix and does not require the eigenvectors
to be orthogonal.
The eigen vectors corresponding to distinct eigen values are orthogonal.
However, the matrix \(\mathbf{V}\) formed by the eigen vectors as columns is NOT orthogonal,
because the rows/columns are NOT orthonormal i.e unit length.
So, in order to make the matrix \(\mathbf{V}\) orthogonal, we need to normalize the rows/columns of the matrix
\(\mathbf{V}\), i.e, make, each eigen vector(column) unit length, by dividing the vector by its magnitude.
After normalisation we get orthogonal matrix \(\mathbf{Q}\) that is composed of unit length and orthogonal eigen vectors
or orthonormal eigen vectors.
Since, matrix \(\mathbf{Q}\) is orthogonal, => \(\mathbf{Q}^T = \mathbf{Q}^{-1}\)
The eigen value decomposition of a square matrix is:
\(\mathbf{A} = \mathbf{V} \mathbf{\Lambda} \mathbf{V}^{-1}\)
And, the spectral decomposition of a real symmetric matrix is:
\(\mathbf{A} = \mathbf{Q} \mathbf{\Lambda} \mathbf{Q}^T\)
Important: Note that, we are discussing only the special case of real symmetric matrix here,
because a real symmetric matrix is guaranteed to have all real eigenvalues.
- Principal Component Analysis (PCA): For dimensionality reduction.
- Page Rank Algorithm: For finding the importance of a web page.
- Structural Engineering: By calculating the eigen values of a bridge’s structural model, we can identify its natural frequencies to ensure that the bridge won’t resonate and be damaged by external forces, such as wind, and seismic waves.
End of Section
2.3.4 - Principal Component Analysis
In the diagram below, if we need to reduce the dimensionality of the data to 1, which feature should be dropped?

Since, information = variance, we should drop the feature that brings least information, i.e, has least variance.
Therefore, drop the feature 1.
What if the variance in both directions is same ?
What should be done in this case? Check the diagram below.
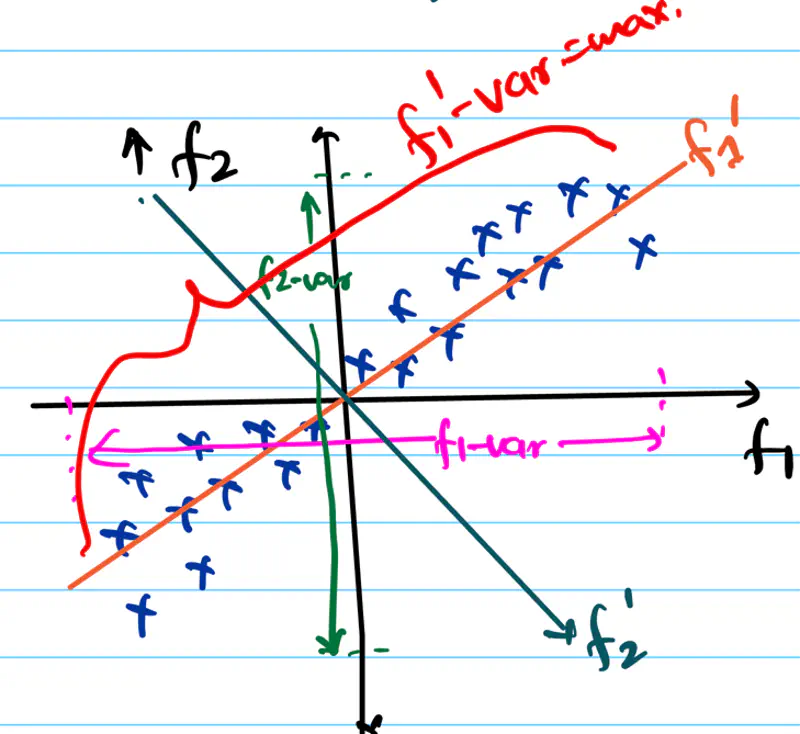
we will rotate the f1-axis in the direction of maximum spread/variance of data, i.e, f1’-axis and then we can drop f2’-axis, which is perpendicular to f1’-axis.
It is a dimensionality reduction technique that finds the direction of maximum variance in the data.
Note: Some loss of information will always be there in dimensionality reduction, because there will be some variability
in data along the direction that is dropped, and that will be lost.
Goal:
Fundamental goal of PCA is to find the new set of orthogonal axes, called the principal components, onto which
the data can be projected, such that, the variance of the projected data is maximum.
Say, we have data, \(D:X \in \mathbb{R}^{n \times d}\),
n is the number of samples
d is the number of features or dimensions of each data point.
In order to find the directions of maximum variance in data, we will use the covariance matrix of data.
Covariance matrix (C) summarizes the spread and relationship of the data in the original d-dimensional space.
\(C_{d \times d} = \frac{1}{n-1}X^TX \), where \(X\) is the data matrix.
Note: (n-1) in the denominator is for unbiased estimation(Bessel’s correction) of covariance matrix.
\(C_{ii}\) is the variance of the \(i^{th}\) feature.
\(C_{ij}\) is the co-variance between feature \(i\) and feature \(j\).
Trace(C) = Sum of diagonal elements of C = Total variance of data.
Algorithm:
Data is first mean centered, i.e, make mean = 0, i.e, subtract mean from each data point.
\(X = X - \mu\)Compute the covariance matrix with mean centered data.
\(C = \frac{1}{n-1}X^TX \), \( \quad \Sigma = \begin{bmatrix} var(f_1) & cov(f_1f_2) & \cdots & cov(f_1f_d) \\ cov(f_2f_1) & var(f_2) & \cdots & cov(f_2f_d) \\ \vdots & \vdots & \ddots & \vdots \\ cov(f_df_1) & cov(f_df_2) & \cdots & var(f_d) \end{bmatrix} _{\text{d x d}} \)Perform the eigen value decomposition of covariance matrix.
\( C = Q \Lambda Q^T \)
\(C\): Orthogonal matrix of eigen vectors of covariance matrix.
New rotated axes or prinicipal components of the data.
\(\Lambda\): Diagonal matrix of eigen values of covariance matrix.
Scaling of variance along new eigen basis.
Note: Eigen values are sorted in descending order, i.e \( \lambda_1 \geq \lambda_2 \geq \cdots \geq \lambda_d \).Project the data onto the new axes or principal components/directions.
Note: k < d = reduced dimensionality.
\(X_{new} = Z = XQ_k\)
\(X_{new}\): Projected data or principal component score
Variance of projected data is given by the eigen value of the co-variance matrix.
Covariance of projected data = \((XQ)^TXQ \)
Therefore, the diagonal matrix \( \Lambda \) captures the variance along every principal component direction.
End of Section
2.3.5 - Singular Value Decomposition
It decomposes any matrix into a rotation, a scaling (based on singular values), and another rotation.
It is a generalization of the eigen value decomposition(for square matrix) to rectangular matrices.
Any rectangular matrix can be decomposed into a product of three matrices using SVD, as follows:
\(\mathbf{U}\): Set of orthonormal eigen vectors of \(\mathbf{AA^T}_{m \times m} \)
\(\mathbf{V}\): Set of orthonormal eigen vectors of \(\mathbf{A^TA}_{n \times n} \)
\(\mathbf{\Sigma}\): Rectangular diagonal matrix, whose diagonal values are called singular values;
square root of non-zero eigen values of \(\mathbf{AA^T}\).
Note: The number of non-zero diagonal entries in \(\mathbf{\Sigma}\) = rank of matrix \(\mathbf{A}\).
Rank (r): Number of linearly independent rows or columns of a matrix.
\( \Sigma = \begin{bmatrix}
\sigma_{11} & 0 & \cdots & 0 \\
\\
0 & \sigma_{22} & \cdots & 0 \\
\vdots & \vdots & \ddots & \vdots \\
\cdots & \cdots & \sigma_{rr} & \vdots \\
0 & 0 & \cdots & 0
\end{bmatrix}
\),
such that, \(\sigma_{11} \geq \sigma_{22} \geq \cdots \geq \sigma_{rr}\), r = rank of \(\mathbf{A}\).
Singular value decomposition, thus refers to the set of scale factors \(\mathbf{\Sigma}\) that are fundamentally
linked to the matrix’s singularity and rank.
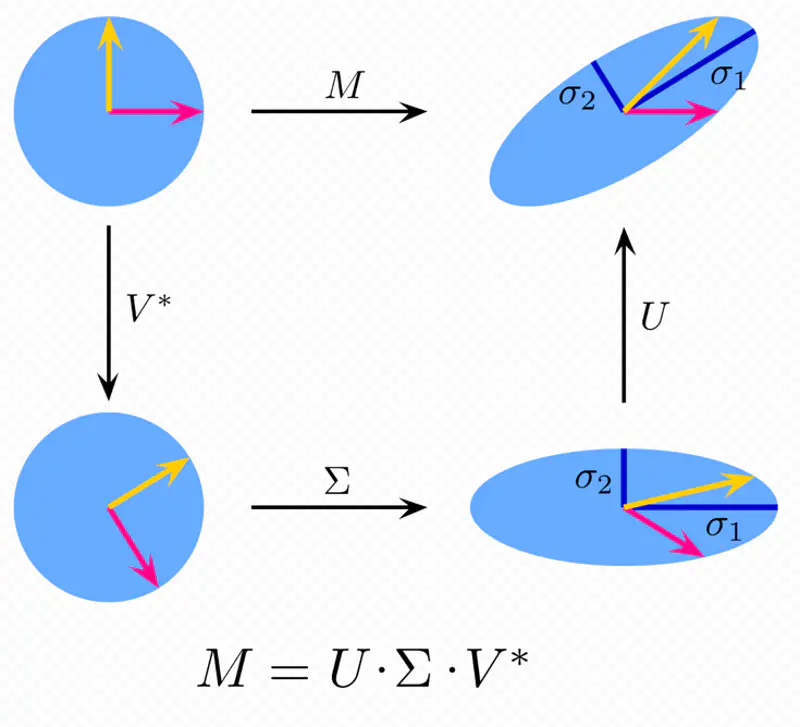
- All singular values are non-negative.
- Square roots of the eigen values of the matrix \(\mathbf{AA^T}\) or \(\mathbf{A^TA}\).
- Arranged in non-decreasing order.
\(\sigma_{11} \geq \sigma_{22} \geq \cdots \geq \sigma_{rr} \ge 0\)
Note: If rank of matrix < dimensions, then 1 or more of the singular values are zero, i.e, dimension collapse.
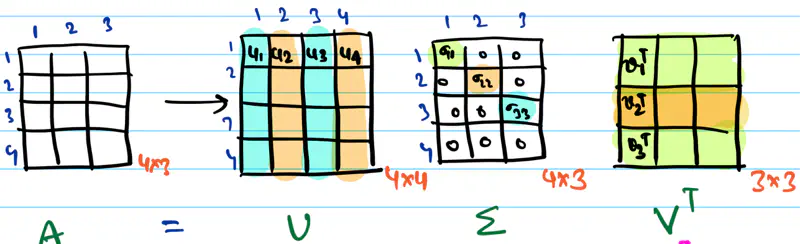
How can we compress the image size to save satellite bandwidth?
We can perform singular value decomposition on the image matrix and find the minimum number of top ranks required to
to successfully reconstruct the original image back.
Say we performed the SVD on the image matrix and found that top 20 rank singular values, out of 1000, are sufficient to
tell what the picture is about.
\(\mathbf{A} = \mathbf{U} \mathbf{\Sigma} \mathbf{V}^T \)
\( = u_1 \sigma_1 v_1^T + u_2 \sigma_2 v_2^T + \cdots + u_r \sigma_r v_r^T \), where \(r = 20\)
A = sum of ‘r=20’ matrices of rank=1.
Now, we need to send only the \(u_i, \sigma_i , v_i\) values for i=20, i.e, top 20 ranks to earth
and then do the calculation to reconstruct the approximation of original image.
\(\mathbf{u} = \begin{bmatrix} u_1 \\ u_2 \\ \vdots \\ u_{1000} \end{bmatrix}\)
\(\mathbf{v} = \begin{bmatrix} v_1 \\ v_2 \\ \vdots \\ v_{1000} \end{bmatrix}\)
So, we need to send data corresponding to 20 (rank=1) matrices, i.e, \(u_i, v_i ~and~ \sigma_i\) = (1000 + 1000 + 1)20 = 200020 pixels (approx)
Therefore, compression rate = (2000*20)/(10^6) = 1/25
- Image compression.
- Low Rank Approximation: Compress data by keeping top rank singular values.
- Noise Reduction: Capture main structure, ignore small singular values.
- Recommendation Systems: Decompose user-item rating matrix to discover underlying user preferences and make recommendations.
The process of approximating any matrix by a matrix of a lower rank, using singular value decomposition.
It is used for data compression.
Any matrix A of rank ‘r’ can be written as sum of rank=1 outer products:
\[ \mathbf{A} = \mathbf{U} \mathbf{\Sigma} \mathbf{V}^T = \sum_{i=1}^{r} \sigma_i \mathbf{u}_i \mathbf{v}_i^T \]\( \mathbf{u}_i : i^{th}\) column vector of \(\mathbf{U}\)
\( \mathbf{v}_i^T : i^{th}\) column vector of \(\mathbf{V}^T\)
\( \sigma_i : i^{th}\) singular value, i.e, diagonal entry of \(\mathbf{\Sigma}\)
Since, the singular values are arranged from largest to smallest, i.e, \(\sigma_1 \geq \sigma_2 \geq \cdots \geq \sigma_r\),
The top ranks capture the vast majority of the information or variance in the matrix.
So, in order to get the best rank-k approximation of the matrix,
we simply truncate the summation after the k’th term.
The approximation \(\mathbf{A_k}\) is called the low rank approximation of \(\mathbf{A}\),
which is achieved by keeping only the largest singular values and corresponding vectors.
Applications:
- Image compression
- Data compression, such as, LoRA (Low Rank Adaptation)
End of Section
2.3.6 - Vector & Matrix Calculus
Let \(\mathbf{x} = \begin{bmatrix} x_1 \\ x_2 \\ \vdots \\ x_n \end{bmatrix}_{\text{n×1}}\) be a vector, i.e, \(\mathbf{x} \in \mathbb{R}^n\).
Let \(f(x)\) be a function that maps a vector to a scalar, i.e, \(f : \mathbb{R}^n \rightarrow \mathbb{R}\).
The derivative of function \(f(x)\) with respect to \(\mathbf{x}\) is defined as:
Assumption: All the first order partial derivatives exist.
e.g.:
- \(f(x) = a^Tx\), where,
\(\mathbf{x} = \begin{bmatrix} x_1 \\ x_2 \\ \vdots \\ x_n \end{bmatrix}\),
\(\quad a = \begin{bmatrix} a_1 \\ a_2 \\ \vdots \\ a_n \end{bmatrix}\),
\(\quad a^T = \begin{bmatrix} a_1 & a_2 & \cdots & a_n \end{bmatrix}\)
\(=> f(x) = a^Tx = a_1x_1 + a_2x_2 + \cdots + a_nx_n\)
\(=> \frac{\partial f(x)}{\partial x} = \begin{bmatrix} \frac{\partial x_1}{\partial x} \\ \frac{\partial x_2}{\partial x} \\ \vdots \\ \frac{\partial x_n}{\partial x} \end{bmatrix} \) \(=\begin{bmatrix} a_1 \\ a_2 \\ \vdots \\ a_n \end{bmatrix} = a \)
\( => f'(x) = a \)
- Now, let’s find the derivative for a bit complex, but very widely used function.
\(f(x) = \mathbf{x^TAx} \quad\), where, \(\mathbf{x} = \begin{bmatrix} x_1 \\ x_2 \\ \vdots \\ x_n \end{bmatrix}\), so, \(\quad x^T = \begin{bmatrix} x_1 & x_2 & \cdots & x_n \end{bmatrix}\),
A is a square matrix, \( \mathbf{A} = \begin{bmatrix} a_{11} & a_{12} & \cdots & a_{1k} & \cdots a_{1n} \\ a_{21} & a_{22} & \cdots & a_{2k} & \cdots a_{2n} \\ \vdots & \vdots & \ddots & \vdots & \vdots \\ a_{k1} & a_{k2} & \cdots & a_{kk} & \cdots a_{kn} \\ \vdots & \vdots & \ddots & \vdots & \vdots \\ a_{n1} & a_{n2} & \cdots & a_{nk} & \cdots a_{nn} \end{bmatrix} _{\text{n x n}} \)
So, \( \mathbf{A^T} = \begin{bmatrix} a_{11} & a_{21} & \cdots & a_{k1} & \cdots a_{n1} \\ a_{12} & a_{22} & \cdots & a_{k2} & \cdots a_{n2} \\ \vdots & \vdots & \ddots & \vdots & \vdots \\ a_{1k} & a_{2k} & \cdots & a_{kk} & \cdots a_{nk} \\ \vdots & \vdots & \ddots & \vdots & \vdots \\ a_{1n} & a_{2n} & \cdots & a_{kn} & \cdots a_{nn} \end{bmatrix} _{\text{n x n}} \)
Let’s calculate \(\frac{\partial y}{\partial x_k}\), i.e, for the \(k^{th}\) element and sum it over 1 to n.
The \(k^{th}\) element \(x_k\) will appear 2 times in the below summation, i.e, when \(i=k ~and~ j=k\)
Above, we saw the gradient of a scalar valued function, i.e, a function that maps a vector to a scalar, i.e,
\(f : \mathbb{R}^n \rightarrow \mathbb{R}\).
There is another kind of function called vector valued function, i.e, a function that maps a vector to another vector, i.e,
\(f : \mathbb{R}^n \rightarrow \mathbb{R}^m\).
The Jacobian is the matrix of all first-order partial derivatives of a vector-valued function,
while the gradient is a vector representing the partial derivatives of a scalar-valued function.
- Jacobian matrix provides the best linear approximation of a vector valued function near a given point,
similar to how a derivative/gradient is the best linear approximation for a scalar valued function
Note: Gradient is a special case of the Jacobian;
it is the transpose of the Jacobian for a scalar valued function.
Let, \(f(x)\) be a function that maps a vector to another vector, i.e, \(f : \mathbb{R}^n \rightarrow \mathbb{R}^m\)
\(f(x)_{m \times 1} = \mathbf{A_{m \times n}x_{n \times 1}}\), where,
\(\quad
\mathbf{A} =
\begin{bmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{m1} & a_{m2} & \cdots & a_{mn}
\end{bmatrix}
_{\text{m x n}}
\),
\(\mathbf{x} = \begin{bmatrix} x_1 \\ x_2 \\ \vdots \\ x_n \end{bmatrix}_{\text{n x 1}}\),
\(f(x) = \begin{bmatrix} f_1(x) \\ f_2(x) \\ \vdots \\ f_m(x) \end{bmatrix}_{\text{m x 1}}\)
The above matrix is called Jacobian matrix of \(f(x)\).
Assumption: All the first order partial derivatives exist.
Hessian Matrix:
It is a square matrix of second-order partial derivatives of a scalar-valued function.
This is used to characterize the curvature of a function at a give point.
Let, \(f(x)\) be a function that maps a vector to a scalar value, i.e, \(f : \mathbb{R}^n \rightarrow \mathbb{R}\)
The Hessian matrix is defined as:
Note: Most functions in Machine Learning where second-order partial derivatives are continuous, the Hessian is symmetrix.
\( H_{i,j} = H_{j,i} = \frac{\partial f^2(x)}{\partial x_i \partial x_j } = \frac{\partial f^2(x)}{\partial x_j \partial x_i } \)
e.g:
\(f(x) = \mathbf{x^TAx}\), where, A is a symmetric matrix, and f(x) is a scalar valued function.
\(Gradient = \frac{\partial }{\partial x}(\mathbf{x^TAx}) = 2\mathbf{Ax}\), since A is symmetric.
\(Hessian = \frac{\partial^2 }{\partial x^2}(\mathbf{x^TAx}) = \frac{\partial }{\partial x}2\mathbf{Ax} = 2\mathbf{A}\)\(f(x,y) = x^2 + y^2\)
\(Gradient = \nabla f = \begin{bmatrix} \frac{\partial f}{\partial x}\\ \\ \frac{\partial f}{\partial x} \end{bmatrix} = \begin{bmatrix} 2x+0 \\ \\ 0+2y \end{bmatrix} = \begin{bmatrix} 2x \\ \\ 2y \end{bmatrix}\)
\(Hessian = \nabla^2 f = \begin{bmatrix} \frac{\partial^2 f}{\partial x^2} & \frac{\partial^2 f}{\partial x \partial y}\\ \\ \frac{\partial^2 f}{\partial y \partial x} & \frac{\partial^2 f}{\partial y^2} \end{bmatrix} = \begin{bmatrix} 2 & 0 \\ \\ 0 & 2 \end{bmatrix}\)
Let, A is a mxn matrix, i.e \(A \in \mathbb{R}^{m \times n}\)
\(\mathbf{A} =
\begin{bmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{m1} & a_{m2} & \cdots & a_{mn}
\end{bmatrix}
_{\text{m x n}}
\)
Let f(A) be a function that maps a matrix to a scalar value, i.e, \(f : \mathbb{R}^{m \times n} \rightarrow \mathbb{R}\)
, then derivative of function f(A) w.r.t A is defined as:
e.g.:
- Let, A is a square matrix, i.e \(A \in \mathbb{R}^{n \times n}\)
and f(A) = trace(A) = \(a_{11} + a_{22} + \cdots + a_{nn}\)
We know that:
\((\frac{\partial f}{\partial A})_{(i,j)} = \frac{\partial f}{\partial a_{ij}}\)
Since, the trace only contains diagonal elements,
=> \((\frac{\partial f}{\partial A})_{(i,j)}\) = 0 for all \(i ⍯ j\)
similarly, \((\frac{\partial f}{\partial A})_{(i,i)}\) = 1 for all \(i=j\)
=> \( \frac{\partial Tr(A)}{\partial A} = \begin{cases} 1, & \text{if } i=j \\ \\ 0, & \text{if } i⍯j \end{cases} \)
\(\frac{\partial f}{\partial A} = \frac{\partial Tr(A)}{\partial A} \begin{bmatrix} 1 & 0 & \cdots & 0 \\ 0 & 1& \cdots & 0 \\ \vdots & \vdots & \ddots & \vdots \\ 0 & 0 & \cdots & 1 \end{bmatrix} = \mathbf{I} \)
Therefore, derivative of trace(A) w.r.t A is an identity matrix.
End of Section
2.3.7 - Vector Norms
It is a measure of the size of a vector or distance from the origin.
Vector norm is a function that maps a vector to a real number, i.e, \({\| \cdot \|} : \mathbb{R}^n \rightarrow \mathbb{R}\).
Vector Norm should satisfy following 3 properties:
- Non-Negativity:
Norm is always greater than or equal to zero,
\( {\| x \|} \ge 0\), and \( {\| x \|} = 0\), if and only if \(\vec{x} = \vec{0}\). - Homogeneity (or Scaling):
\( {\| \alpha x \|} = |\alpha| {\| x \|} \). - Triangle Inequality:
\( {\| x + y \|} \le {\| x \|} + {\| y \|} \).
P-Norm:
It is a generalised form of most common family of vector norms, also called Minkowski norm.
It is defined as:
We can change the value of \(p\) to get different norms.
L1 Norm:
It is the sum of absolute values of all the elements of a vector; also known as Manhattan distance.
p=1:
L2 Norm:
It is the square root of the sum of squares of all the elements of a vector; also known as Euclidean distance.
p=2:
L-\(\infty\) Norm:
It is the maximum of absolute values of all the elements of a vector; also known as Chebyshev distance.
p=\(\infty\):
- Let, vector \(\mathbf{x} = \begin{bmatrix} 3 \\ \\ -4 \end{bmatrix}\), then
\({\| x \|_1} = |3| + |-4| = 7\)
\({\| x \|_2} = \sqrt{3^2 + (-4)^2} = \sqrt{25} = 5\)
\({\| x \|_\infty} = max(|3|, |-4|) = max(3, 4) = 4\)
It is a function that assigns non-negative size or magnitude to a matrix.
Matrix Norm is a function that maps a matrix to a non-negative real number, i.e, \({\| \cdot \|} : \mathbb{R}^{m \times n} \rightarrow \mathbb{R}\)
It should satisfy following 3 properties:
- Non-Negativity:
Norm is always greater than or equal to zero,
\( {\| x \|} \ge 0\), and \( {\| x \|} = 0\), if and only if \(\vec{x} = \vec{0}\). - Homogeneity (or Scaling):
\( {\| \alpha x \|} = |\alpha| {\| x \|} \). - Triangle Inequality:
\( {\| x + y \|} \le {\| x \|} + {\| y \|} \).
There are 2 types of matrix norms:
- Element wise norms, e.g,, Frobenius norm
- Vector induced norms Frobenius Norm:
Frobenius Norm:
It is equivalent to the Euclidean norm of the matrix if it were flattened into a single vector.
If A is a matrix of size \(m \times n\), then, Frobenius norm is defined as:
\(\sigma_i\) is the \(i\)th singular value of matrix A.
Vector Induced Norm:
It measures the maximum stretching a matrix can apply when multiplied with a vector,
where the vector has a unit length under the chosen vector norm.
Matrix Induced by Vector P-Norm:
P-Norm:
P=1 Norm:
P=\(\infty\) Norm:
P=2 Norm:
Also called Spectral norm, i.e, maximum factor by which the matrix can stretch a unit vector in Euclidean norm.
Let, matrix \(\mathbf{A} = \begin{bmatrix} a_{11} & a_{12} \\ \\ a_{21} & a_{22} \end{bmatrix}\), then find Frobenius norm.
\({\| A \|_F} = \sqrt{a_{11}^2 + a_{12}^2 + a_{21}^2 + a_{22}^2}\)
\(\mathbf{A}^T = \begin{bmatrix} a_{11} & a_{22} \\ \\ a_{12} & a_{22} \end{bmatrix}\)
=> \(\mathbf{A^TA} = \begin{bmatrix} a_{11} & a_{22} \\ \\ a_{12} & a_{22} \end{bmatrix} \begin{bmatrix} a_{11} & a_{12} \\ \\ a_{21} & a_{22} \end{bmatrix} = \begin{bmatrix} a_{11}^2 + a_{12}^2 & a_{11}.a_{21} + a_{12}.a_{22} \\ \\ a_{21}.a_{11} + a_{22}.a_{12} & a_{21}^2 + a_{22}^2 \end{bmatrix} \)
Therefore, \(Trace(\mathbf{A^TA}) = a_{11}^2 + a_{12}^2 + a_{21}^2 + a_{22}^2\)
=> \({\| A \|_F} = \sqrt{Trace(A^TA)} = \sqrt{a_{11}^2 + a_{12}^2 + a_{21}^2 + a_{22}^2}\)Let, matrix \(\mathbf{A} = \begin{bmatrix} 1 & -2 & 3 \\ \\ 4 & 5 & -6 \end{bmatrix}\), then
Column 1 absolute value sum = |1|+|4| = 5
Column 2 absolute value sum = |-2|+|5|= 7
Column 3 absolute value sum = |3|+|-6|= 9
Row 1 absolute value sum = |1|+|-2|+|3| = 6
Row 2 absolute value sum = |4|+|5|+|-6| = 15
\({\| A \|_1} = max(5,7,9) = 9\) = max column absolute value sum.
\({\| A \|_\infty} = max(6,15) = 15\) = max row absolute value sum.
- Let, matrix \(\mathbf{A} = \begin{bmatrix} 2 & 1 \\ \\ 1 & 2 \end{bmatrix}\), then find spectral norm.
Spectral norm can be found using the singular value decomposition, in order to get the largest singular value.
\({\| A \|_2} = \sigma_{max}(A) \)
\(\mathbf{A} = \mathbf{U \Sigma V^T} \) , where
\(\mathbf{U} = \mathbf{AA^T} \),
\(\mathbf{V} = \mathbf{A^TA} \)
Let’s find the largest eigen value of \(\mathbf{A^TA} \), square root of which will give the largest singular value of \(\mathbf{A}\).
\( \mathbf{V} = \mathbf{A^TA} =
\begin{bmatrix} 2 & 1 \\ \\ 1 & 2 \end{bmatrix}
\begin{bmatrix} 2 & 1 \\ \\ 1 & 2 \end{bmatrix}
= \begin{bmatrix} 5 & 4 \\ \\ 4 & 5 \end{bmatrix}
\)
Now, lets find the eigen values of the above matrix V:
\(det(V-\lambda I) = 0 \)
=> \(
det\begin{bmatrix} 5 - \lambda & 4 \\ \\ 4 & 5- \lambda \end{bmatrix} = 0
\)
=> \( (5 - \lambda)^2 - 16 = 0 \)
=> \( (5 - \lambda) = \pm 4 \)
=> \( (5 - \lambda) = 4 \) or \( (5 - \lambda) = -4 \)
=> \( \lambda = 1 \) or \( \lambda = 9 \)
=> Largest Singular Value = Square root of largest eigen value = \(\sqrt{9} = 3\)
Therefore, \({\| A \|_2} = \sigma_{max}(A) = 3\)
End of Section
2.3.8 - Hyperplane
Equation of a line is of the form \(y = mx + c\).
To represent a line in 2D space, we need 2 things:
- m = slope or direction of the line
- c = y-intercept or distance from the origin
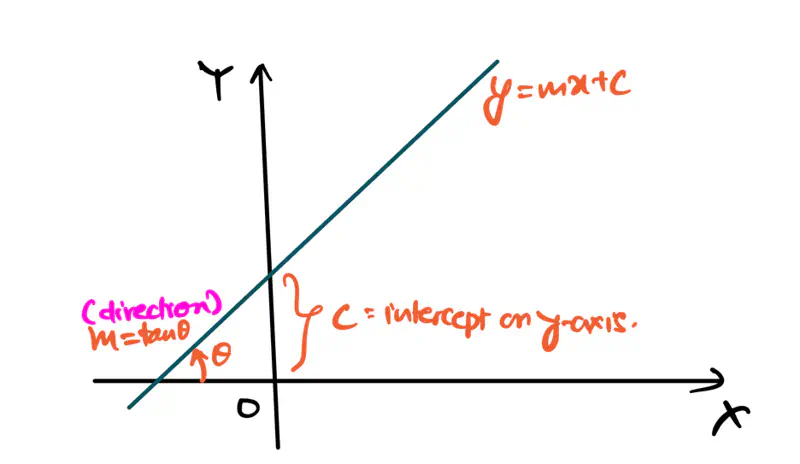
Similarly, to represent a hyperplane in d-dimensions, we need 2 things:
- \(\vec{w}\) = direction of the hyperplane = vector perpendicular to the hyperplane
- \(w_0\) = distance from the origin
Note: There can be only 2 directions of the hyperplane, i.e, direction of a unit vector perpendicular to the hyperplane:
- Towards the origin
- Away from the origin
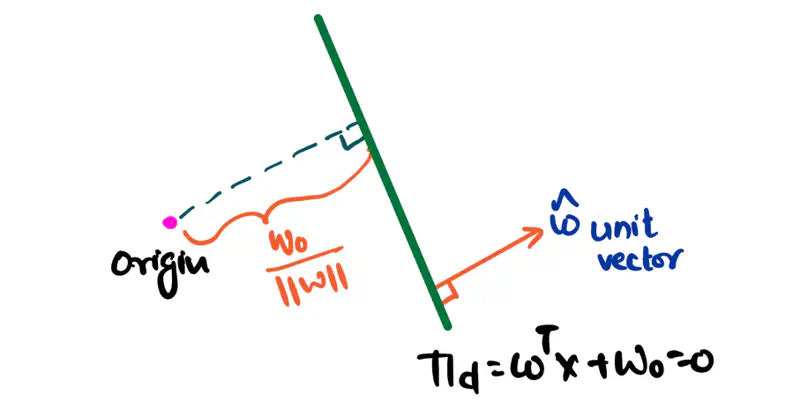
If a point ‘x’ is on the hyperplane, then it satisfies the below equation:
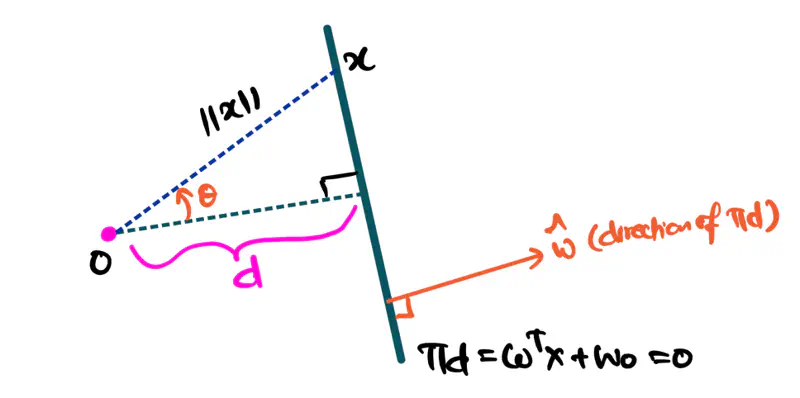
Key Points:
- By convention, the direction of the hyperplane is given by a unit vector perpendicular to the hyperplane , i.e, \({\Vert \mathbf{w} \Vert} = 1\), since the direction is only important.
- \(w_0\) gives the signed perpendicular distance from the origin.
\(w_0 = 0\) => Hyperplane passes through the origin.
\(w_0 < 0\) => Hyperplane is in the same direction of unit vector \(\mathbf{\widehat{w}}\) w.r.t the origin.
\(w_0 > 0\) => Hyperplane is in the opposite direction of unit vector \(\mathbf{\widehat{w}}\) w.r.t the origin.
What is the direction of the hyperplane w.r.t the origin and the direction of the unit vector ?
Equation of a hyperplane is: \(\pi_d = \mathbf{w}^\top \mathbf{x} + w_0 = 0\)
Let, the equation of the line/hyperplane in 2D be:
\(\pi_d = 1.x + 0.y + w_0 = x + w_0 = 0\)
Case 1: \(w_0 < 0\), say \(w_0 = -5\)
Therefore, equation of hyperplane: \( x - 5 = 0 => x = 5\)
Here, the hyperplane(line) is located in the same direction as the unit vector w.r.t the origin,
i.e, towards the +ve x-axis direction.
Case 2: \(w_0 > 0\), say \(w_0 = 5\)
Therefore, equation of hyperplane: \( x + 5 = 0 => x = -5\)
Here, the hyperplane(line) is located in the opposite direction as the unit vector w.r.t the origin,
i.e, towards the -ve x-axis direction.
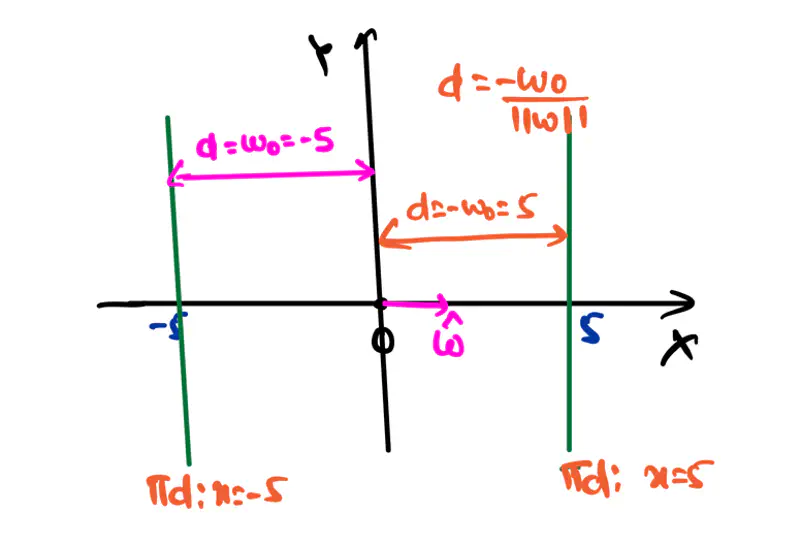
A hyperplane divides a space into 2 distinct parts called half-spaces.
e.g.: A 2D hyperplane divided a 3D space into 2 distinct parts.
Similar example in real world: A wall divides a room into 2 distinct spaces.
Positive Half-Space:
A half space that is in the same direction as the unit vector w.r.t the origin.
Negative Half-Space:
A half space that is in the opposite direction as the unit vector w.r.t the origin.
from equations (1) & (2), we can say that:
\[ \Vert \mathbf{w} \Vert \Vert \mathbf{x_1} \Vert cos{\theta} + w_0 < \Vert \mathbf{w} \Vert \Vert \mathbf{x_1\prime} \Vert cos{\theta} + w_0 \]Everything is same on both the sides except \(\Vert \mathbf{x_1}\Vert\) and \(\Vert \mathbf{x_1\prime\Vert}\), so:
\[ \mathbf{w}^\top \mathbf{x_1} + w_0 < 0 \]i.e, negative half-space, opposite to the direction of unit vector or towards the origin.
Similarly,
i.e, positive half-space, same as the direction of unit vector or away from the origin.
Equation of distance of any point \(x\prime\) from the hyperplane:
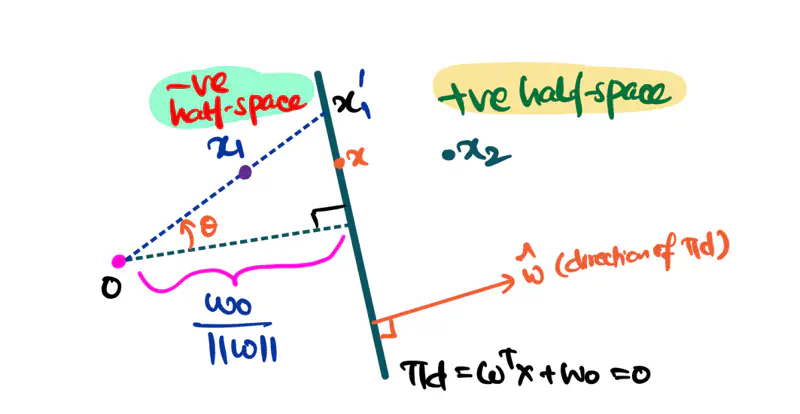
The above concept of equation of hyperplane will be very helpful when we discuss the following topics later:
- Logistic Regression
- Support Vector Machines
End of Section
2.4.1 - Calculus Fundamentals
Integration is a mathematical tool that is used to find the area under a curve.
Let’s understand integration with the help of few simple examples for finding area under a curve:
- Area of a triangle:
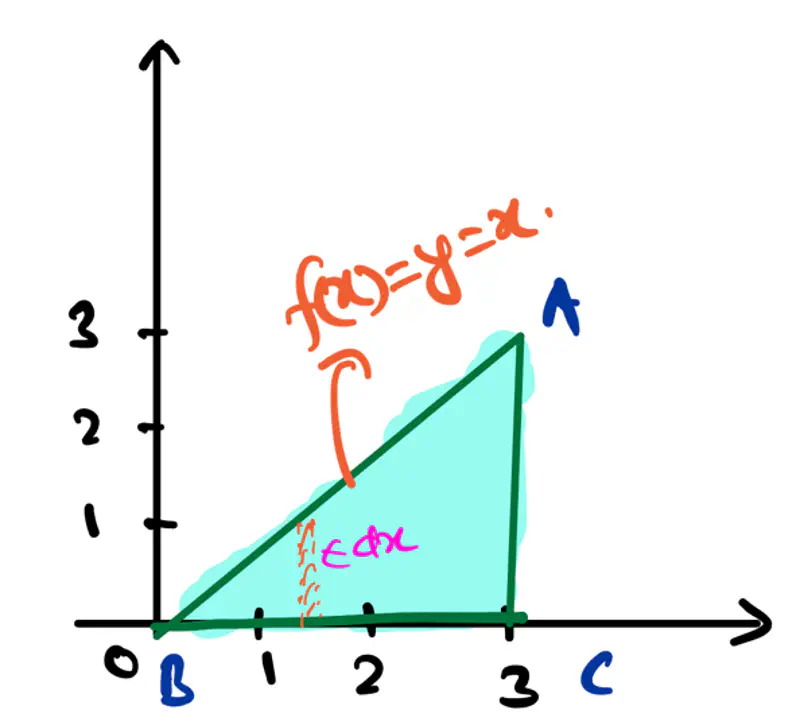
- Area of a rectangle:
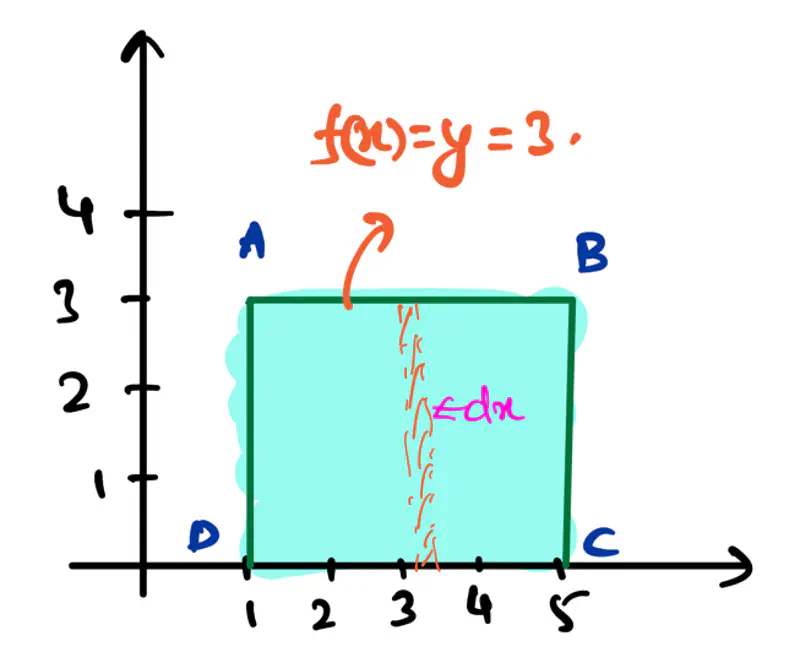
Note: The above examples were standard straight forward, where we know a direct formula for finding area under a curve.
But, what if we have such a shape, for which, we do NOT know a ready-made formula, then how do we calculate
the area under the curve.
Let’s see an example:
3. Area of a part of parabola:
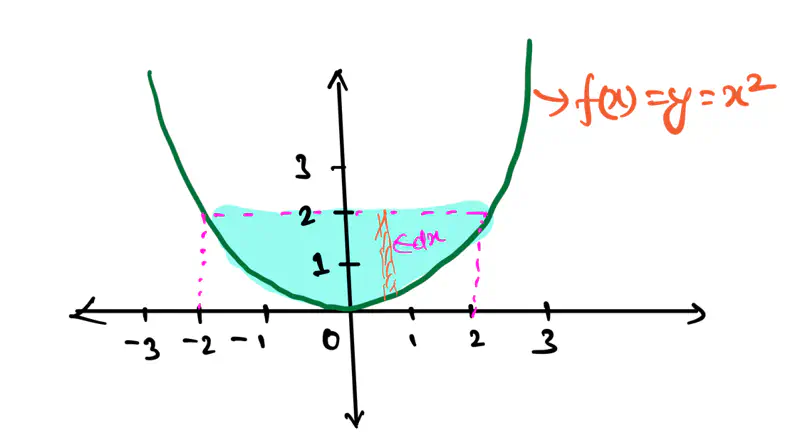
Differentiation is a mathematical tool that is used to find the derivative or rate of change of a function
at a specific point.
- \(\frac{dy}{dx} = f\prime(x) = tan\theta\) = Derivative = Slope = Gradient
- Derivative tells us how fast the function is changing at a specific point in relation to another variable.
Note: For a line, the slope is constant, but for a parabola, the slope is changing at every point.
How do we calculate the slope at a given point?
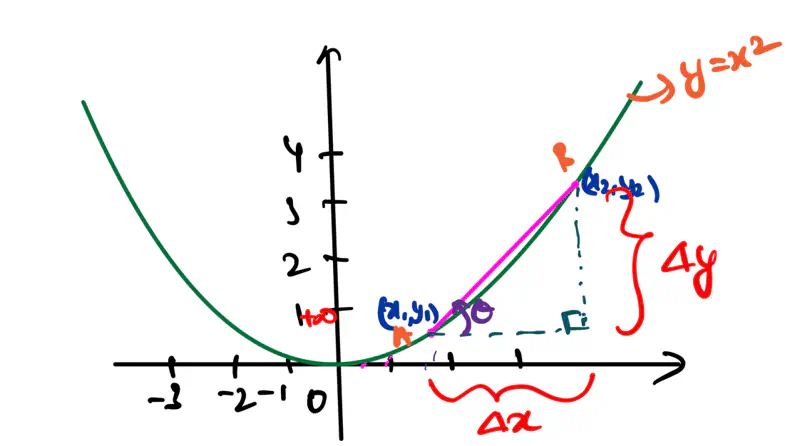
Let AB is a secant on the parabola, i.e, line connecting any 2 points on the curve.
Slope of secant = \(tan\theta = \frac{\Delta y}{\Delta x} = \frac{y_2-y_1}{x_2-x_1}\)
As \(\Delta x \rightarrow 0\), secant will become a tangent to the curve, i.e, the line will touch the curve only
at 1 point.
\(dx = \Delta x \rightarrow 0\)
\(x_2 = x_1 + \Delta x \)
\(y_2 = f(x_2) = f(x_1 + \Delta x) \)
Slope at \(x_1\) =
e.g.:
\( y = f(x) = x^2\), find the derivative of f(x) w.r.t x.
We will understand few important rules of differentiation that are most frequently used in Machine Learning.
Sum Rule:
\[ \frac{d}{dx} (f(x) + g(x)) = \frac{d}{dx} f(x) + \frac{d}{dx} g(x) = f\prime(x) + g\prime(x) \]Product Rule:
\[ \frac{d}{dx} (f(x).g(x)) = \frac{d}{dx} f(x).g(x) + f(x).\frac{d}{dx} g(x) = f\prime(x).g(x) + f(x).g\prime(x) \]e.g.:
\[ h(x) = f(x).g(x) \\[10pt] => h\prime(x) = f\prime(x).g(x) + f(x).g\prime(x) \\[10pt] => h\prime(x) = 2x.sin(x) + x^2.cos(x) \\[10pt] \]
\( h(x) = x^2 sin(x) \), find the derivative of h(x) w.r.t x.
Let, \(f(x) = x^2 , g(x) = sin(x)\).Quotient Rule:
\[ \frac{d}{dx} \frac{f(x)}{g(x)} = \frac{f\prime(x).g(x) - f(x).g\prime(x)}{(g(x))^2} \]e.g.:
\[ h(x) = \frac{f(x)}{g(x)} \\[10pt] => h\prime(x) = \frac{f\prime(x).g(x) - f(x).g\prime(x)}{(g(x))^2} \\[10pt] => h\prime(x) = \frac{cos(x).x - sin(x)}{x^2} \\[10pt] \]
\( h(x) = sin(x)/x \), find the derivative of h(x) w.r.t x.
Let, \(f(x) = sin(x) , g(x) = x\).Chain Rule:
\[ \frac{d}{dx} (f(g(x))) = f\prime(g(x)).g\prime(x) \]e.g.:
\[ h(x) = log(u) \\[10pt] => h\prime(x) = \frac{d h(x)}{du} \cdot \frac{du}{dx} \\[10pt] => h\prime(x) = \frac{1}{u} \cdot 2x = \frac{2x}{x^2} \\[10pt] => h\prime(x) = \frac{2}{x} \]
\( h(x) = log(x^2) \), find the derivative of h(x) w.r.t x.
Let, \( u = x^2 \)
Now, let’s dive deeper and understand the concepts that required for differentiation, such as, limits, continuity, differentiability, etc.
Limit of a function f(x) at any point ‘c’ is the value that f(x) approaches, as x gets very close to ‘c’,
but NOT necessarily equal to ‘c’.
One-sided limit: value of the function, as it approaches a point ‘c’ from only one direction, either left or right.
Two-sided limit: value of the function, as it approaches a point ‘c’ from both directions, left and right, simultaneously.
e.g.:
- \(f(x) = \frac{1}{x}\), find the limit of f(x) at x = 0.
Let’s check for one-sided limit at x=0:
\[ \lim_{x \rightarrow 0^+} \frac{1}{x} = + \infty \\[10pt] \lim_{x \rightarrow 0^-} \frac{1}{x} = - \infty \\[10pt] so, \lim_{x \rightarrow 0^+} \frac{1}{x} ⍯ \lim_{x \rightarrow 0^-} \frac{1}{x} \\[10pt] => \text{ limit does NOT exist at } x = 0. \]
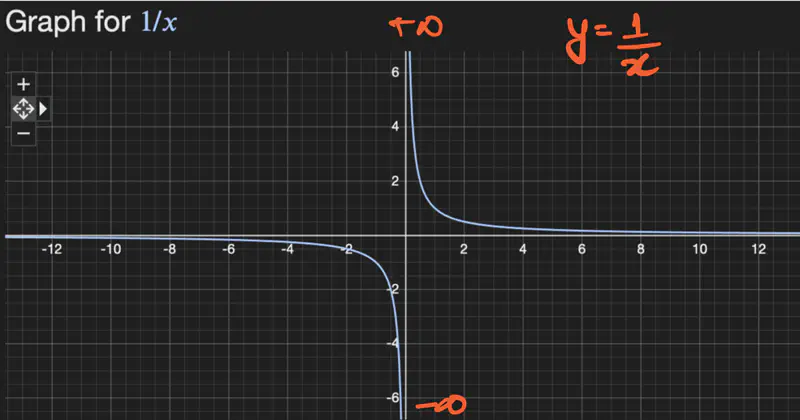
2. \(f(x) = x^2\), find the limit of f(x) at x = 0.
Let’s check for one-sided limit at x=0:
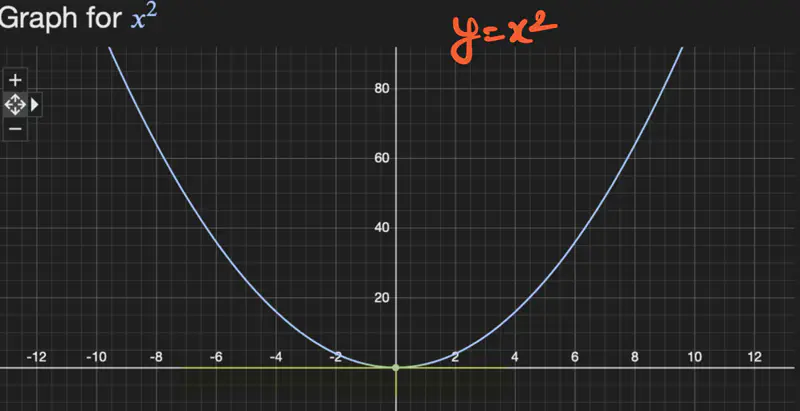
Note: Two-Sided Limit
3. f(x) = |x|, find the limit of f(x) at x = 0.
Let’s check for one-sided limit at x=0:
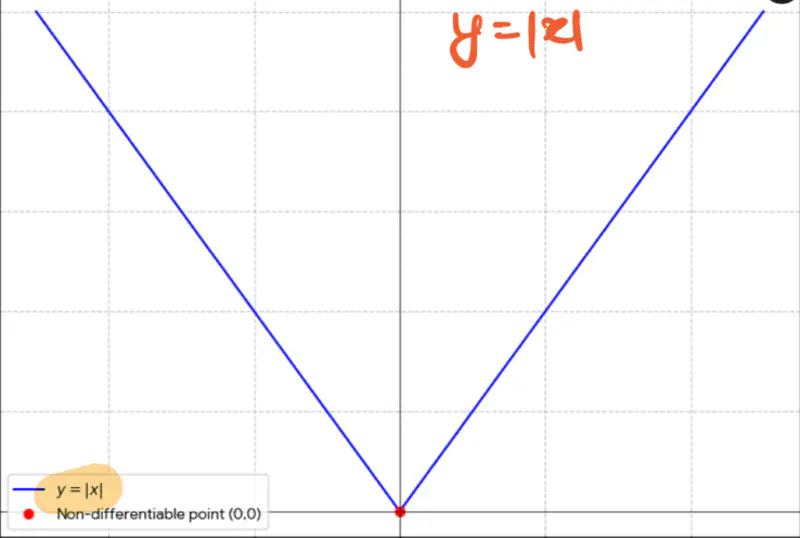
A function f(x) is said to be continuous at a point ‘c’, if its graph can be drawn through that point,
without lifting the pen.
Continuity bridges the gap between the function’s value at the given point and the limit.
Conditions for Continuity:
A function f(x) is continuous at a point ‘c’, if and only if, all the below 3 conditions are met:
- f(x) must be defined at point ‘c’.
- Limit of f(x) must exist at point ‘c’, i.e, left and right limits must be equal. \[ \lim_{x \rightarrow c^+} f(x) = \lim_{x \rightarrow c^-} f(x) \]
- Value of f(x) at ‘c’ must be equal to its limit at ‘c’. \[ \lim_{x \rightarrow c} f(x) = f(c) \]
e.g.:
- \(f(x) = \frac{1}{x}\) is NOT continuous at x = 0, since, f(x) is not defined at x = 0.

- \(f(x) = |x|\) is continuous everywhere.

- \(f(x) = tanx \) is discontinuous at infinite points.
A function is differentiable at a point ‘c’, if derivative of the function exists at that point.
A function must be continuous at the given point ‘c’ to be differentiable at that point.
Note: A function can be continuous at a given point, but NOT differentiable at that point.
e.g.:
We know that \( f(x) = |x| \) is continuous at x=0, but its NOT differentiable at x=0.
Critical Point:
A point of the function where the derivative is either zero or undefined.
These critical points are candidates for local maxima or minima,
which are the highest and lowest points in a function’s immediate neighborhood, respectively.
Maxima:
Highest point w.r.t immediate neighbourhood.
f’(x)/gradient/slope changes from +ve to 0 to -ve, therefore, change in f’(x) is -ve.
=> f’’(x) < 0
Let, \(f(x) = -x^2; \quad f'(x) = -2x; \quad f''(x) = -2 < 0 => maxima\)
| x | f’(x) |
|---|---|
| -1 | 2 |
| 0 | 0 |
| 1 | -2 |
Minima:
Lowest point w.r.t immediate neighbourhood.
f’(x)/gradient/slope changes from -ve to 0 to +ve, therefore, change in f’(x) is +ve.
=> f’’(x) > 0
Let, \(f(x) = x^2; \quad f'(x) = 2x; \quad f''(x) = 2 > 0 => minima\)
| x | f’(x) |
|---|---|
| -1 | -2 |
| 0 | 0 |
| 1 | 2 |

e.g.:
- Let \(f(x) = 2x^3 + 5x^2 + 3 \), find the maxima and minima of f(x).
To find the maxima and minima, lets take the derivative of the function and equate it to zero.
\[ f'(x) = 6x^2 + 10x = 0\\[10pt] => x(6x+10) = 0 \\[10pt] => x = 0 \quad or \quad x = -10/6 = -5/3 \\[10pt] \text{ lets check the second order derivative to find which point is maxima and minima: } \\[10pt] f''(x) = 12x + 10 \\[10pt] => at ~ x = 0, \quad f''(x) = 12*0 + 10 = 10 >0 \quad => minima \\[10pt] => at ~ x = -5/3, \quad f''(x) = 12*(-5/3) + 10 = -20 + 10 = -10<0 \quad => maxima \\[10pt] \]
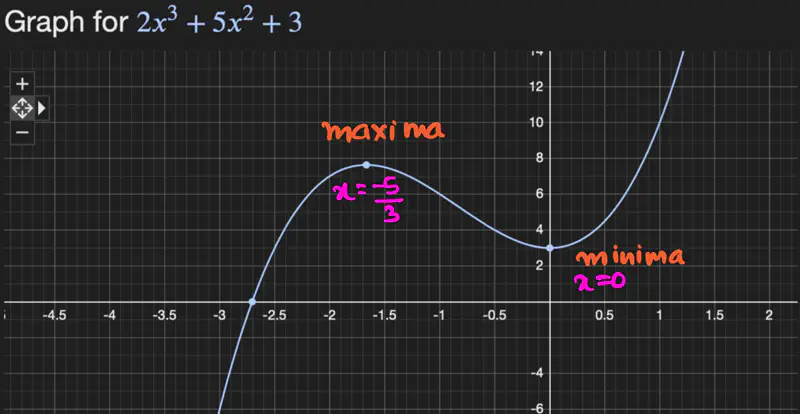
- \(f(x,y) = z = x^2 + y^2\), find the minima of f(x,y).
Since, this is a multi-variable function, we will use vector and matrix for calculation.
\[ Gradient = \nabla f_z = \begin{bmatrix} \frac{\partial f_z}{\partial x} \\ \\ \frac{\partial f_z}{\partial y} \end{bmatrix} = \begin{bmatrix} 2x \\ \\ 2y \end{bmatrix} = \begin{bmatrix} 0 \\ \\ 0 \end{bmatrix} \\[10pt] => x=0, y=0 \text{ is a point of optima for } f(x,y) \]
Partial Derivative:
Partial derivative \( \frac{\partial f(x,y)}{\partial x} ~or~ \frac{\partial f(x,y)}{\partial y} \)is the rate of change
or derivative of a multi-variable function w.r.t one of its variables,
while all the other variables are held constant.
Let’s continue solving the above problem, and calculate the Hessian, i.e, 2nd order derivative of f(x,y):
Since, determinant of Hessian = 4 > 0 and \( \frac{\partial^2 f_z}{\partial x^2} > 0\) => (x=0, y=0) is a point of minima.
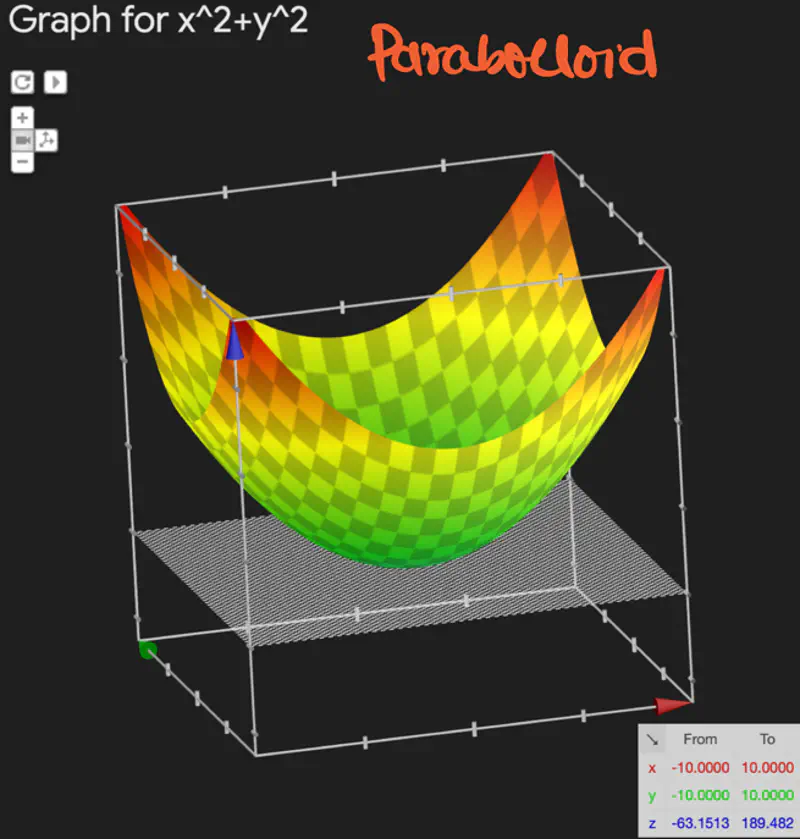
Hessian Interpretation:
- Minima: If det(Hessian) > 0 and \( \frac{\partial^2 f(x,y)}{\partial x^2} > 0\)
- Maxima: If det(Hessian) > 0 and \( \frac{\partial^2 f(x,y)}{\partial x^2} < 0\)
- Saddle Point: If det(Hessian) < 0
- Inconclusive: If det(Hessian) = 0, need to perform other tests.
Saddle Point is a critical point where the function is maximum w.r.t one variable,
and minimum w.r.t to another.
e.g.:
Let, \(f(x,y) = z = x^2 - y^2\), find the point of optima for f(x,y).
Since, determinant of Hessian = -4 < 0 => (x=0, y=0) is a saddle point.
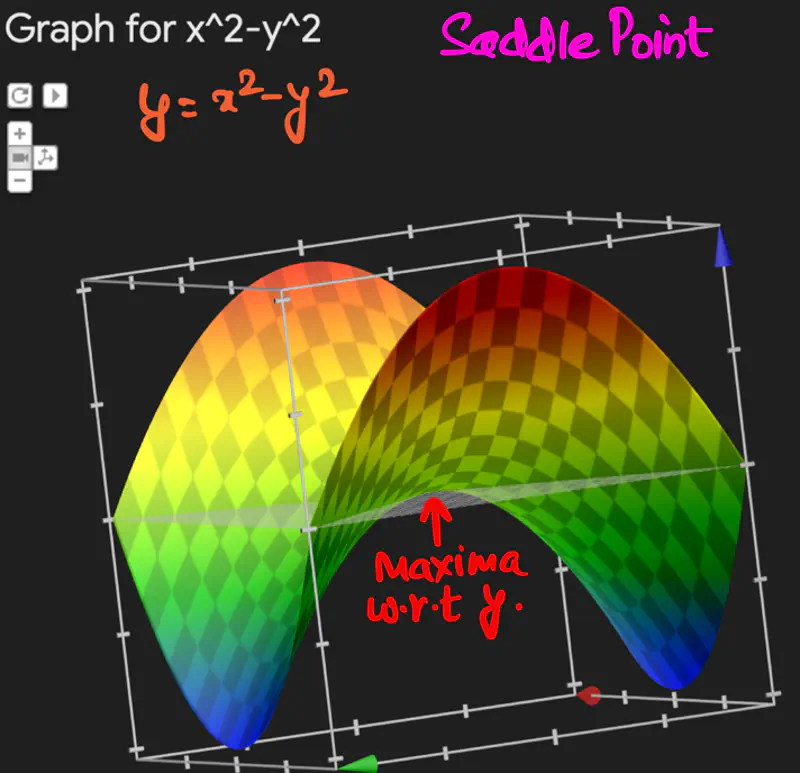
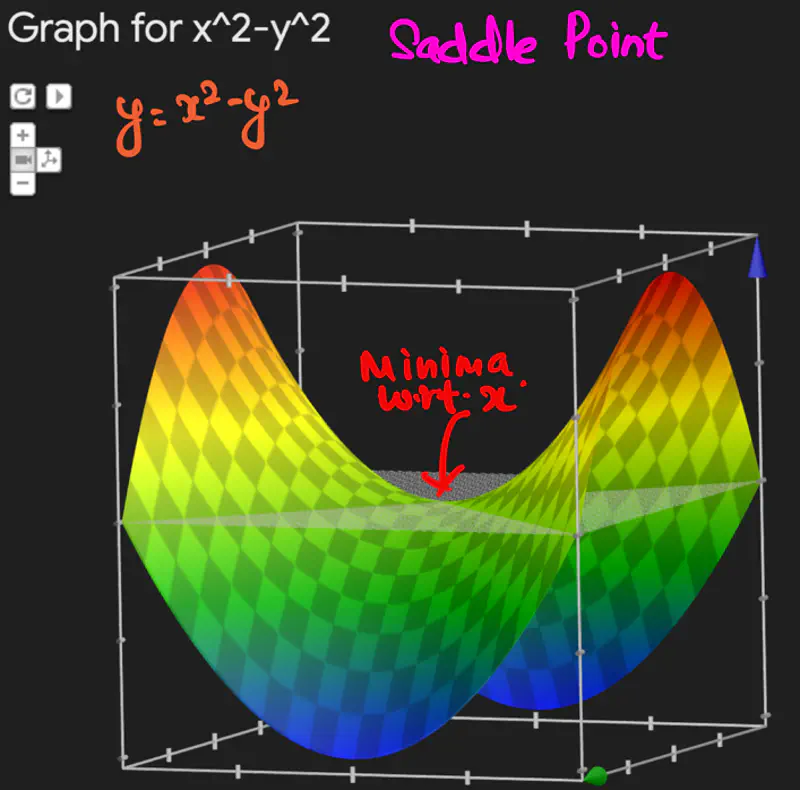
End of Section
2.4.2 - Optimization
How do we measure and minimize these mistakes in predictions made by the model?
minimizing a loss function.
Loss Function:
It quantifies error of a single data point in a dataset.
e.g.: Squared Loss, Hinge Loss, Absolute Loss, etc, for a single data point.
Cost Function:
It is the average of all losses over the entire dataset.
e.g.: Mean Squared Error(MSE), Mean Absolute Error(MAE), etc.
Objective Function:
It is the over-arching objective of an optimization problem, representing the function that is minimized or maximized.
e.g.: Minimize MSE.
Let’s understant this through an example:
Task:
Predict the price of a company’s stock based on its historical data.
Objective Function:
Minimize the difference between actual and predicted price.
Let, \(y\): original or actual price
\(\hat y\): predicted price
Say, the dataset has ’n’ such data points.
Loss Function:
loss = \( y - \hat y \) for a single data point.
We want to minimize the loss for all ’n’ data points.
Cost Function:
We want to minimize the average/total loss over all ’n’ data points.
So, what are the ways ?
- We take a simple sum of all the losses, but this can be misleading, as loss for a
single data point can be +ve or -ve, we can get a net-zero loss even for very large losses, when we sum them all. - We take the sum of absolute value of each loss, i.e, \( |y - \hat y| \), this way the losses will not cancel out each other.
But the absolute value function is NOT differentiable at y=0, and this can cause issues in optimisation algorithms, such as, gradient descent.
Read more about Differentiability - So, we choose squared loss, i.e, \( (y - \hat y)^2 \), this solves the above issues.
Note: In general, we refer to the cost function as the loss function also, the terms are used interchangeably.
Cost = Loss = Mean Squared Error(MSE) \[\frac{1}{n} \sum_{i=1}^{n} (y_i - \hat y_i)^2 \] The task is to minimize the above loss.
Key Points:
- Loss is the bridge between ‘data’ and ‘optimization’.
- Good loss functions are differentiable and convex.
Convexity:
It refers to a property of a function where a line segment connecting any two points on its graph
lies above or on the graph itself.
A convex function is curved upwards.
It is always described by a convex set.
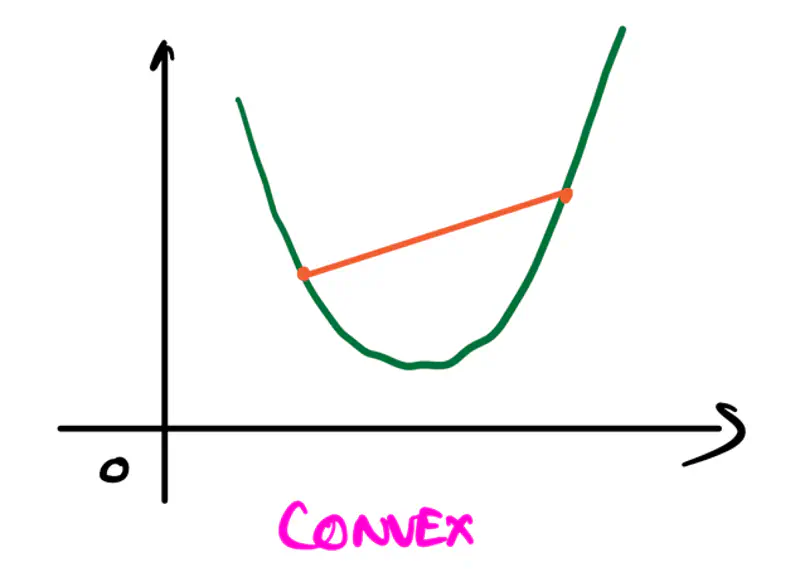
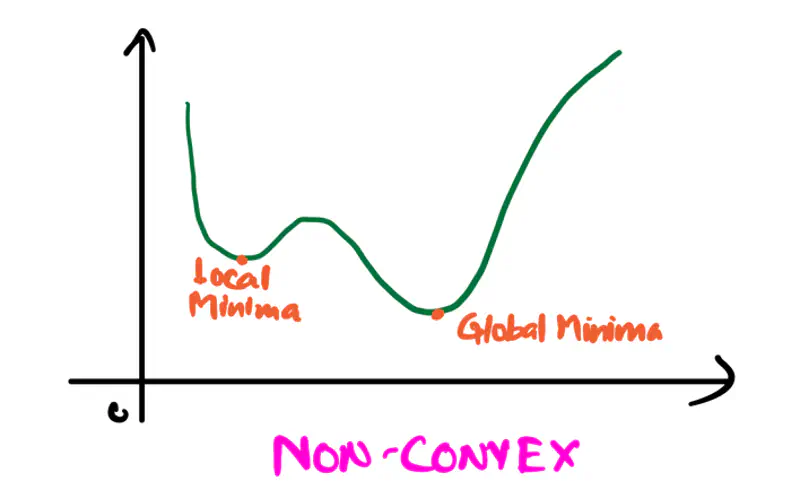
Convex Set:
A convex set is a set of points in which the straight line segment connecting any two points in the set lies
entirely within that set.
A set \(C\) is convex if for any two points \(x\) and \(y\) in \(C\), the convex combination
\(\theta x+(1-\theta )y\) is also in \(C\) for all values of \(\theta \) where \(0\le \theta \le 1\).
A function \(f: \mathbb{R}^n \rightarrow \mathbb{R}\) is convex if for all values of \(x,y\) and \(0\le \theta \le 1\),
\[ f(\theta x + (1-\theta )y) \le \theta f(x) + (1-\theta )f(y) \]Second-Order Test:
If a function is twice differentiable, i.e, 2nd derivative exists, then the function is convex,
if and only if, the Hessian is positive semi-definite for all points in its domain.
Positive Definite:
A symmetric matrix is positive definite if and only if:
- Eigenvalues are all strictly positive, or
- For any non-zero vector \(z\), the quadratic form \(z^THz > 0\)
Note: If the Hessian is positive definite, then the function is convex; has upward curvature in all directions.
Positive Semi-Definite:
A symmetric matrix is positive semi-definite if and only if:
- Eigenvalues are all non-negative (i.e, greater than or equal to zero), or
- For any non-zero vector \(z\), the quadratic form \(z^THz \ge 0\)
Note: If the Hessian is positive definite, then the function is not strictly convex, but flat in some directions.
All machine learning algorithms minimize loss (mostly), so we need to find the optimum parameters for the model that
minimizes the loss.
This is an optimization problem, i.e, find the best solution from a set of alternatives.
Note: Convexity ensures that there is only 1 minima for the loss function.
Optimization:
It is the iterative procedure of finding the optimum parameter \(x^*\) that minimizes the loss function f(x).
Let’s formulate an optimization problem for a model to minimize the MSE loss function discussed above:
Note: To minimize the loss, we want \(y_i, \hat y_i\) to be as close as possible, for that we want to find the optimum weights \(w, w_0\) of the model.
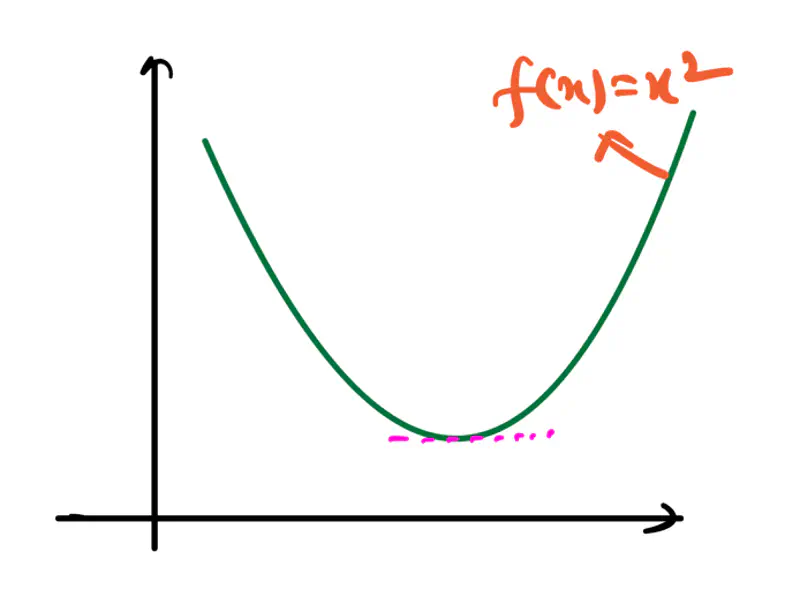
Important:
Deep Learning models have non-convex loss function, so it is challenging to reach the global minima,
so any local minima is also a good enough solution.
Read more about Maxima-Minima
Constrained Optimization:
It is an optimization process to find the best possible solution (min or max), but within a set of limitations or
restrictions called constraints.
Constraints limit the range of acceptable values; they can be equality constraints or inequality constraints.
e.g.:
Minimize f(x) subject to following constraints:
Equality Constraints: \( g_i(x) = c_i \forall ~i \in \{1,2,3, \ldots, n\} \)
Inequality Constraints: \( h_j(x) \le d_j \forall ~j \in \{1,2,3, \ldots, m\}\)
Lagrangian Method:
Lagrangian method converts a constrained optimization problem to an unconstrained optimization problem,
by introducing a new variable called Lagrange multiplier (\(\lambda\)).
Note: Addition of Lagrangian function that incorporates the constraints,
makes the problem solvable using standard calculus.
e.g.:
Let f(x) be the objective function with single equality constraint \(g(x) = c\),
then the Lagrangian function \( \mathcal{L}\) is defined as:
Now, the above constrained optimization problem becomes an unconstrained optimization problem:
By solving the above unconstrained optimization problem,
we get the optimum solution for the original constrained problem.
Objective: To minimize the distance between point (x,y) on the line 2x + 3y = 13 and the origin (0,0).
distance, d = \(\sqrt{(x-0)^2 + (y-0)^2}\)
=> Objective function = minimize distance = \( \underset{x^*, y^*}{\mathrm{argmin}}\ f(x,y) =
\underset{x^*, y^*}{\mathrm{argmin}}\ x^2+y^2\)
Constraint: Point (x,y) must be on the line 2x + 3y = 13.
=> Constraint (equality) function = \(g(x,y) = 2x + 3y - 13 = 0\)
Lagragian function =
To find the optimum solution, we solve the below unconstrained optimization problem.
Take the derivative and equate it to zero.
Since, it is multi-variable function, we take the partial derivatives, w.r.t, x, y and \(\lambda\).
Now, we have 3 variables and 3 equations (1), (2) and (3), lets solve them.
Hence, the point (x=2, y=3) on the line 2x + 3y = 13 that is closest to the origin.
To solve the optimization problem, there are many methods, such as, analytical method, which gives the normal equation for the linear regression,
but we will discuss that method later in detail, when we have understood what is linear regression?
Normal Equation for linear regression:
X: Feature variables
y: Vector of all observed target values
End of Section
2.4.3 - Gradient Descent
Till now, we have understood how to formulate a minimization problem as a mathematical optimization problem.
Now, lets take a step forward and understand how to solve these optimization problems.
We will focus on two important iterative gradient based methods:
- Gradient Descent: First order method, uses only the gradient.
- Newton’s Method: Second order method, used both the gradient and the Hessian.
Gradient Descent:
It is a first order iterative optimization algorithm that is used to find the local minimum of a differentiable function.
It iteratively adjusts the parameters of the model in the direction opposite to the gradient of cost function,
since moving opposite to the direction of gradient leads towards the minima.
Algorithm:
Initialize the weights/parameters with random values.
Calculate the gradient of the cost function at current parameter values.
Update the parameters using the gradient.
\[ w_{new} = w_{old} - \eta \cdot \frac{\partial f}{\partial w_{old}} \\[10pt] \eta: \text{ learning rate or step size to take for each parameter update} \]Repeat steps 2 and 3 iteratively until convergence (to minima).
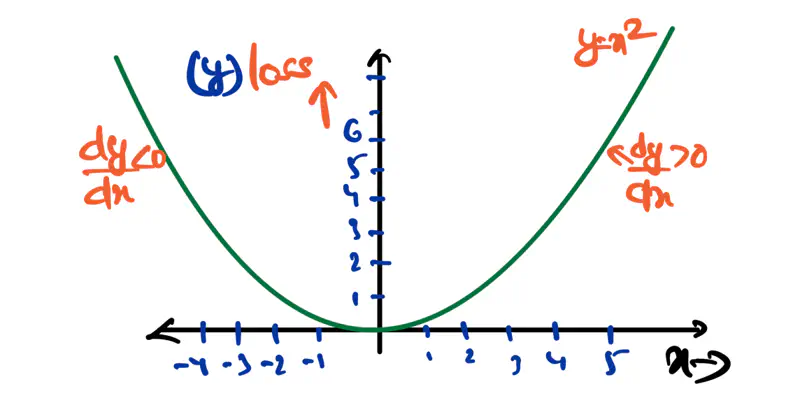
There are 3 types of Gradient Descent:
- Batch Gradient Descent
- Stochastic Gradient Descent
- Mini-Batch Gradient Descent
Batch Gradient Descent (BGD):
Computes the gradient using all the data points in the dataset for parameter update in each iteration.
Say, number of data points in the dataset is \(n\).
Let, the loss function for individual data point be \(l_i(w)\)
Key Points:
- Slow steps towards convergence, i.e, TC = O(n).
- Smooth, direct path towards minima.
- Number of steps/iterations is minimum.
- Not suitable for large datasets; impractical for Deep Learning, as n = millions/billions.
Stochastic Gradient Descent (SGD):
It uses only 1 data point selected randomly from dataset to compute gradient for parameter update in each iteration.
Key Points:
- Computationally fastest per step; TC = O(1).
- Highly noisy, zig-zag path to minima.
- High variance in gradient estimation makes path to minima volatile, requiring a careful decay of learning rate \(\eta\) to ensure convergence to minima.
Mini Batch Gradient Descent (MBGD):
It uses small randomly selected subsets of dataset, called mini-batch, (1<k<n) to compute gradient for
parameter update in each iteration.
Key Points:
- Moderate time consumption per step; TC = O(k<n).
- Less noisy, and more reliable convergence than stochastic gradient descent.
- More efficient and faster than batch gradient descent.
- Standard optimization algorithm for Deep Learning.
Note: Vectorization on GPUs allows for parallel processing of mini-batches; also GPUs are the reason for the mini-batch size to be a power of 2.
End of Section
2.4.4 - Newton's Method
Newton’s Method:
It is a second-order iterative gradient based optimization technique known for its extremely fast convergence.
When close to optimum, it achieves quadratic convergence, better than gradient descent’s linear convergence.
Algorithm:
- Start at a random point \(x_k\).
- Compute the slope at \(x_k, ~i.e,~ f'(x_k)\).
- Compute the curvature at \(x_k, ~i.e,~ f''(x_k)\).
- Draw a parabola at \(x_k\) that locally approximates the function.
- Jump directly to the minimum of that parabola; that’s the next step. Note: So, instead of walking down the slope step by step (gradient descent), we are jumping straight to the point where the curve bends downwards towards the bottom.
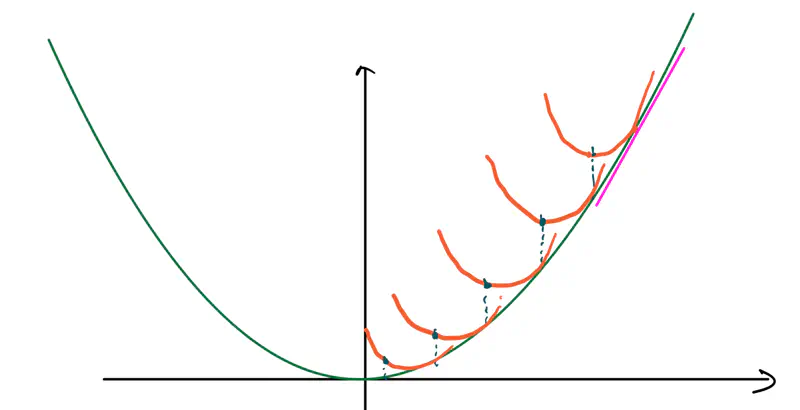
- Find the minima of \(f(x) = x^2 - 4x + 5\)
To find the minima, lets calulate the first derivative and equate to zero.
\(f'(x) = 2x - 4 = 0 \)
\( => x^* = 2 \)
\(f''(x) = 2 >0 \) => minima is at \(x^* = 2\)
Now, we will solve this using Newton’s Method.
Let’s start at x = 0.
Hence, we can see that using Newton’s Method we can get to the minima \(x^* = 2\) in just 1 step.
Full Newton’s Method is rarely used in Machine Learning/Deep Learning optimization,
because of the following limitations:
- \(TC = O(n^2\)) for Hessian calculation, since for a network with \(n\) parameters
requires \(n^2\) derivative calculations. - \(TC = O(n^3\)) for Hessian Inverse calculation.
- If it encounters a Saddle point, then it can go to a maxima rather than minima.
Because of the above limitations, we use Quasi-Newton methods like BFGS and L-BFGS.
The quasi-Newton methods make use of approximations for Hessian calculation, in order to gain the benefits
of curvature without incurring the cost of Hessian calculation.
BFGS: Broyden-Fletcher-Goldfarb-Shanno
L-BFGS: Limited-memory BFGS
End of Section